{
"cells": [
{
"cell_type": "markdown",
"id": "3bbe8002-bdf3-490c-bde0-80dd3713a3d0",
"metadata": {},
"source": [
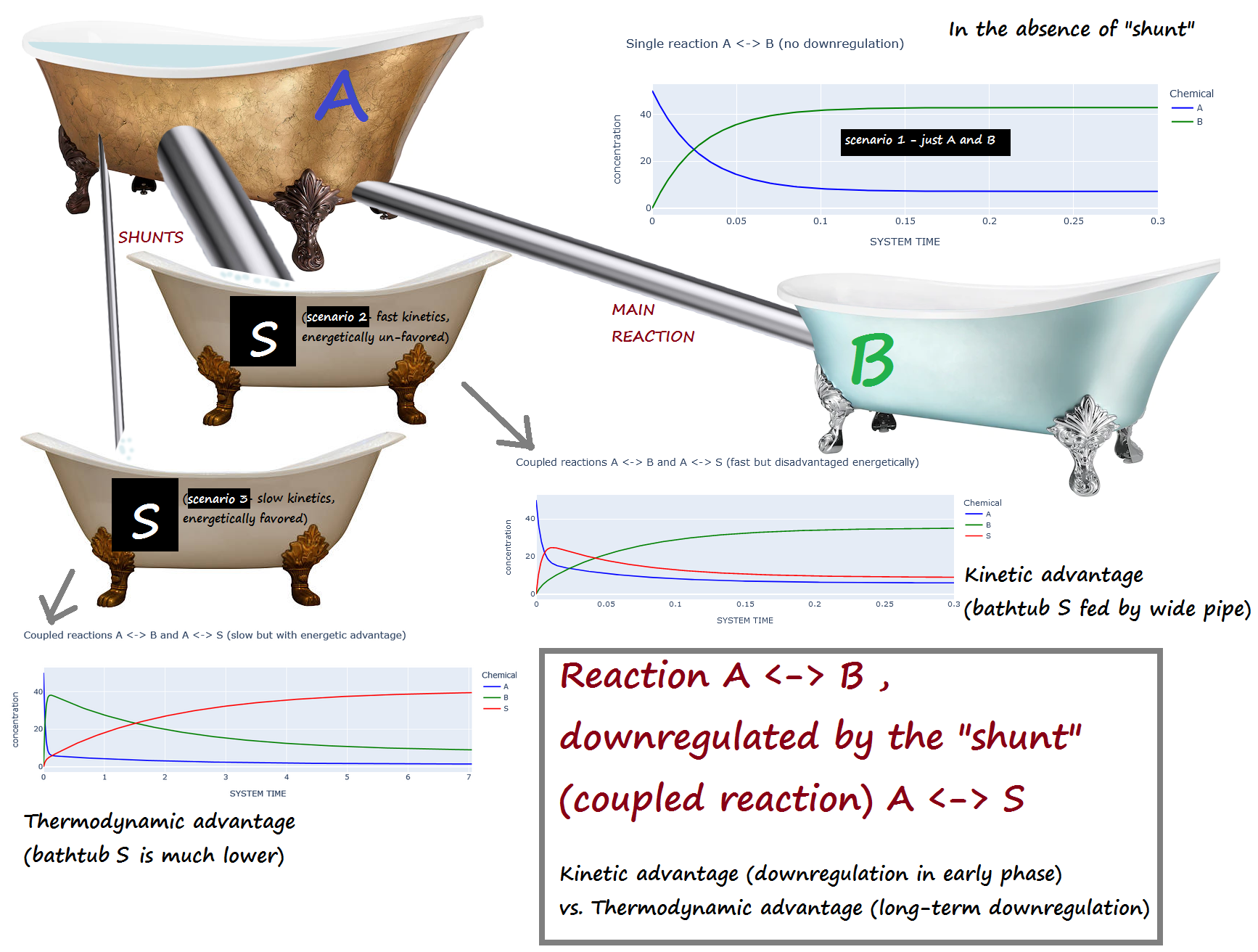
"## `A <-> B` , downregulated by the \"shunt\" (coupled reaction) `A <-> S`\n",
"### _Kinetic_ advantage (downregulation in early phase) vs. _Thermodynamic_ advantage (long-term downregulation) \n",
"\n",
"**[Scenario 1](#down_regulate_1_scenario_1)** : No downregulation on `A <-> B `\n",
"\n",
"**[Scenario 2](#down_regulate_1_scenario_2)** : The shunt (`A <-> S`) has a *kinetic* advantage but *thermodynamic* DIS-advantage compared to `A <-> B ` \n",
"(i.e. `A <-> S` is fast, but energetically unfavored) \n",
"\n",
"**[Scenario 3](#down_regulate_1_scenario_3)** : The shunt (`A <-> S`) is has a *kinetic* DIS-advantage but a *thermodynamic* advantage compared to `A <-> B` \n",
"(i.e. `A <-> S` is slow, but energetically favored) \n",
"\n",
"All reactions are 1st order, mostly forward. Taken to equilibrium.\n",
"\n",
"LAST REVISED: June 23, 2024 (using v. 1.0 beta36)"
]
},
{
"cell_type": "markdown",
"id": "61171e99-518e-4019-a731-be7437e95dfd",
"metadata": {},
"source": [
"## Bathtub analogy:\n",
"A is initially full, while B and S are empty. \n",
"If the \"shunt\" S is present, scenario 2 corresponds to a large pipe and a small elevation change... \n",
"while scenario 3 corresponds to a narrow pipe and a large elevation change."
]
},
{
"cell_type": "markdown",
"id": "832afe37-f169-41c8-a719-e739336bc5cf",
"metadata": {},
"source": [
""
]
},
{
"cell_type": "code",
"execution_count": 1,
"id": "c4231a63-e456-48e9-bf4f-d074bbd3490a",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"Added 'D:\\Docs\\- MY CODE\\BioSimulations\\life123-Win7' to sys.path\n"
]
}
],
"source": [
"import set_path # Importing this module will add the project's home directory to sys.path"
]
},
{
"cell_type": "code",
"execution_count": 2,
"id": "64933384",
"metadata": {
"tags": []
},
"outputs": [],
"source": [
"from experiments.get_notebook_info import get_notebook_basename\n",
"\n",
"from life123 import ChemData as chem\n",
"from life123 import UniformCompartment\n",
"\n",
"import plotly.express as px\n",
"from life123 import GraphicLog"
]
},
{
"cell_type": "code",
"execution_count": 3,
"id": "83c3cc5f-de21-4f66-9988-2806fbf0666d",
"metadata": {
"tags": []
},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"-> Output will be LOGGED into the file 'down_regulate_1.log.htm'\n"
]
}
],
"source": [
"# Initialize the HTML logging (for the graphics)\n",
"log_file = get_notebook_basename() + \".log.htm\" # Use the notebook base filename for the log file\n",
"\n",
"# Set up the use of some specified graphic (Vue) components\n",
"GraphicLog.config(filename=log_file,\n",
" components=[\"vue_cytoscape_2\"],\n",
" extra_js=\"https://cdnjs.cloudflare.com/ajax/libs/cytoscape/3.21.2/cytoscape.umd.js\")"
]
},
{
"cell_type": "code",
"execution_count": null,
"id": "001bc666-f2ef-40a3-b46f-30b087ea33da",
"metadata": {},
"outputs": [],
"source": []
},
{
"cell_type": "markdown",
"id": "35b5ef15-69da-4fc9-b1e9-fd34a9bafb99",
"metadata": {},
"source": [
"# Scenario 1: A <-> B in the absence of the 2nd reaction"
]
},
{
"cell_type": "markdown",
"id": "9329208b-070f-4902-8f37-0f11ddf75ed6",
"metadata": {},
"source": [
"### Initialize the System\n",
"Specify the chemicals and the reaction"
]
},
{
"cell_type": "code",
"execution_count": 4,
"id": "57d8431c-d6d0-462c-af78-e64eeb220e2e",
"metadata": {
"tags": []
},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"Number of reactions: 1 (at temp. 25 C)\n",
"0: A <-> B (kF = 30 / kR = 5 / delta_G = -4,441.7 / K = 6) | 1st order in all reactants & products\n",
"Set of chemicals involved in the above reactions: {'A', 'B'}\n"
]
}
],
"source": [
"# Specify the chemicals\n",
"chem_data = chem(names=[\"A\", \"B\"])\n",
"\n",
"# Reaction A <-> B\n",
"chem_data.add_reaction(reactants=[\"A\"], products=[\"B\"],\n",
" forward_rate=30., reverse_rate=5.)\n",
"\n",
"chem_data.describe_reactions()"
]
},
{
"cell_type": "markdown",
"id": "f5eabdf2-0e6b-4141-a886-10974dfc6c3a",
"metadata": {},
"source": [
"### Set the initial concentrations of all the chemicals"
]
},
{
"cell_type": "code",
"execution_count": 5,
"id": "67a0375f-a14f-4cbe-965b-81d4c841aeab",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"SYSTEM STATE at Time t = 0:\n",
"2 species:\n",
" Species 0 (A). Conc: 50.0\n",
" Species 1 (B). Conc: 0.0\n",
"Set of chemicals involved in reactions: {'A', 'B'}\n"
]
}
],
"source": [
"dynamics = UniformCompartment(chem_data=chem_data, preset=\"fast\")\n",
"dynamics.set_conc(conc={\"A\": 50.}, snapshot=True)\n",
"dynamics.describe_state()"
]
},
{
"cell_type": "markdown",
"id": "72a2148e-1aae-4ed7-bab3-5781b3a80fb0",
"metadata": {},
"source": [
"### Run the reaction"
]
},
{
"cell_type": "code",
"execution_count": 6,
"id": "89f23b49-2840-4517-a275-b6a4f97898af",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"37 total step(s) taken\n",
"Number of step re-do's because of negative concentrations: 0\n",
"Number of step re-do's because of elective soft aborts: 0\n",
"Norm usage: {'norm_A': 22, 'norm_B': 17, 'norm_C': 16, 'norm_D': 16}\n"
]
}
],
"source": [
"dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()\n",
"\n",
"# The changes of concentrations vary very rapidly early on; automated variable timesteps will take care of that\n",
"dynamics.single_compartment_react(initial_step=0.001, duration=0.3,\n",
" snapshots={\"initial_caption\": \"1st reaction step\",\n",
" \"final_caption\": \"last reaction step\"},\n",
" variable_steps=True)"
]
},
{
"cell_type": "code",
"execution_count": 7,
"id": "80fbaee3-bd6f-4197-9270-23374d46a4a7",
"metadata": {},
"outputs": [
{
"data": {
"text/html": [
"\n",
"\n",
"
\n",
" \n",
" \n",
" | \n",
" SYSTEM TIME | \n",
" A | \n",
" B | \n",
" caption | \n",
"
\n",
" \n",
" \n",
" \n",
" | 0 | \n",
" 0.000000 | \n",
" 50.000000 | \n",
" 0.000000 | \n",
" Initialized state | \n",
"
\n",
" \n",
" | 1 | \n",
" 0.001000 | \n",
" 48.500000 | \n",
" 1.500000 | \n",
" 1st reaction step | \n",
"
\n",
" \n",
" | 2 | \n",
" 0.002000 | \n",
" 47.052500 | \n",
" 2.947500 | \n",
" | \n",
"
\n",
" \n",
" | 3 | \n",
" 0.002800 | \n",
" 45.935030 | \n",
" 4.064970 | \n",
" | \n",
"
\n",
" \n",
" | 4 | \n",
" 0.003600 | \n",
" 44.848849 | \n",
" 5.151151 | \n",
" | \n",
"
\n",
" \n",
" | 5 | \n",
" 0.004400 | \n",
" 43.793081 | \n",
" 6.206919 | \n",
" | \n",
"
\n",
" \n",
" | 6 | \n",
" 0.005200 | \n",
" 42.766875 | \n",
" 7.233125 | \n",
" | \n",
"
\n",
" \n",
" | 7 | \n",
" 0.006000 | \n",
" 41.769403 | \n",
" 8.230597 | \n",
" | \n",
"
\n",
" \n",
" | 8 | \n",
" 0.007200 | \n",
" 40.315088 | \n",
" 9.684912 | \n",
" | \n",
"
\n",
" \n",
" | 9 | \n",
" 0.008400 | \n",
" 38.921854 | \n",
" 11.078146 | \n",
" | \n",
"
\n",
" \n",
" | 10 | \n",
" 0.009600 | \n",
" 37.587136 | \n",
" 12.412864 | \n",
" | \n",
"
\n",
" \n",
" | 11 | \n",
" 0.010800 | \n",
" 36.308476 | \n",
" 13.691524 | \n",
" | \n",
"
\n",
" \n",
" | 12 | \n",
" 0.012000 | \n",
" 35.083520 | \n",
" 14.916480 | \n",
" | \n",
"
\n",
" \n",
" | 13 | \n",
" 0.013800 | \n",
" 33.323259 | \n",
" 16.676741 | \n",
" | \n",
"
\n",
" \n",
" | 14 | \n",
" 0.015240 | \n",
" 32.003766 | \n",
" 17.996234 | \n",
" | \n",
"
\n",
" \n",
" | 15 | \n",
" 0.016680 | \n",
" 30.750777 | \n",
" 19.249223 | \n",
" | \n",
"
\n",
" \n",
" | 16 | \n",
" 0.018840 | \n",
" 28.966018 | \n",
" 21.033982 | \n",
" | \n",
"
\n",
" \n",
" | 17 | \n",
" 0.020568 | \n",
" 27.646153 | \n",
" 22.353847 | \n",
" | \n",
"
\n",
" \n",
" | 18 | \n",
" 0.022296 | \n",
" 26.406114 | \n",
" 23.593886 | \n",
" | \n",
"
\n",
" \n",
" | 19 | \n",
" 0.024888 | \n",
" 24.658551 | \n",
" 25.341449 | \n",
" | \n",
"
\n",
" \n",
" | 20 | \n",
" 0.026962 | \n",
" 23.387332 | \n",
" 26.612668 | \n",
" | \n",
"
\n",
" \n",
" | 21 | \n",
" 0.029035 | \n",
" 22.208373 | \n",
" 27.791627 | \n",
" | \n",
"
\n",
" \n",
" | 22 | \n",
" 0.032146 | \n",
" 20.568281 | \n",
" 29.431719 | \n",
" | \n",
"
\n",
" \n",
" | 23 | \n",
" 0.034634 | \n",
" 19.399045 | \n",
" 30.600955 | \n",
" | \n",
"
\n",
" \n",
" | 24 | \n",
" 0.038366 | \n",
" 17.797935 | \n",
" 32.202065 | \n",
" | \n",
"
\n",
" \n",
" | 25 | \n",
" 0.041352 | \n",
" 16.684379 | \n",
" 33.315621 | \n",
" | \n",
"
\n",
" \n",
" | 26 | \n",
" 0.045831 | \n",
" 15.188610 | \n",
" 34.811390 | \n",
" | \n",
"
\n",
" \n",
" | 27 | \n",
" 0.050310 | \n",
" 13.927325 | \n",
" 36.072675 | \n",
" | \n",
"
\n",
" \n",
" | 28 | \n",
" 0.057029 | \n",
" 12.331983 | \n",
" 37.668017 | \n",
" | \n",
"
\n",
" \n",
" | 29 | \n",
" 0.062404 | \n",
" 11.355820 | \n",
" 38.644180 | \n",
" | \n",
"
\n",
" \n",
" | 30 | \n",
" 0.070466 | \n",
" 10.167025 | \n",
" 39.832975 | \n",
" | \n",
"
\n",
" \n",
" | 31 | \n",
" 0.082559 | \n",
" 8.887006 | \n",
" 41.112994 | \n",
" | \n",
"
\n",
" \n",
" | 32 | \n",
" 0.094652 | \n",
" 8.148772 | \n",
" 41.851228 | \n",
" | \n",
"
\n",
" \n",
" | 33 | \n",
" 0.112792 | \n",
" 7.510122 | \n",
" 42.489878 | \n",
" | \n",
"
\n",
" \n",
" | 34 | \n",
" 0.140002 | \n",
" 7.160360 | \n",
" 42.839640 | \n",
" | \n",
"
\n",
" \n",
" | 35 | \n",
" 0.180816 | \n",
" 7.135357 | \n",
" 42.864643 | \n",
" | \n",
"
\n",
" \n",
" | 36 | \n",
" 0.242039 | \n",
" 7.151428 | \n",
" 42.848572 | \n",
" | \n",
"
\n",
" \n",
" | 37 | \n",
" 0.333872 | \n",
" 7.123879 | \n",
" 42.876121 | \n",
" last reaction step | \n",
"
\n",
" \n",
"
\n",
"
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "A",
"line": {
"color": "darkturquoise",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "A",
"orientation": "v",
"showlegend": true,
"type": "scatter",
"x": [
0,
0.001,
0.002,
0.0028,
0.0036,
0.0044,
0.005200000000000001,
0.006000000000000001,
0.0072000000000000015,
0.008400000000000001,
0.009600000000000001,
0.0108,
0.012,
0.0138,
0.01524,
0.01668,
0.018840000000000003,
0.020568000000000003,
0.022296000000000003,
0.024888000000000004,
0.026961600000000006,
0.029035200000000008,
0.03214560000000001,
0.03463392000000001,
0.038366400000000016,
0.04135238400000002,
0.04583136000000002,
0.050310336000000025,
0.05702880000000003,
0.062403571200000035,
0.07046572800000003,
0.08255896320000004,
0.09465219840000005,
0.11279205120000006,
0.14000183040000008,
0.1808164992000001,
0.24203850240000013,
0.3338715072000002
],
"xaxis": "x",
"y": [
50,
48.5,
47.0525,
45.935030000000005,
44.84884916000001,
43.793081383520004,
42.76687510478144,
41.76940260184756,
40.31508769256996,
38.92185400948202,
37.587136141083775,
36.308476423158254,
35.08352041338561,
33.32325862734231,
32.00376639252426,
30.750776566341038,
28.966017857925657,
27.64615309787831,
26.406113758518632,
24.658551118345823,
23.387332112380758,
22.208373096992613,
20.568280768161607,
19.39904451412549,
17.79793541574258,
16.684379152287153,
15.188610469357345,
13.927324707561334,
12.331982669319185,
11.35581998417073,
10.1670251384972,
8.887006018551627,
8.148771928336012,
7.5101218135084515,
7.16036014263505,
7.135356872772086,
7.1514282273422705,
7.1238794318490255
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=B
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "B",
"line": {
"color": "green",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "B",
"orientation": "v",
"showlegend": true,
"type": "scatter",
"x": [
0,
0.001,
0.002,
0.0028,
0.0036,
0.0044,
0.005200000000000001,
0.006000000000000001,
0.0072000000000000015,
0.008400000000000001,
0.009600000000000001,
0.0108,
0.012,
0.0138,
0.01524,
0.01668,
0.018840000000000003,
0.020568000000000003,
0.022296000000000003,
0.024888000000000004,
0.026961600000000006,
0.029035200000000008,
0.03214560000000001,
0.03463392000000001,
0.038366400000000016,
0.04135238400000002,
0.04583136000000002,
0.050310336000000025,
0.05702880000000003,
0.062403571200000035,
0.07046572800000003,
0.08255896320000004,
0.09465219840000005,
0.11279205120000006,
0.14000183040000008,
0.1808164992000001,
0.24203850240000013,
0.3338715072000002
],
"xaxis": "x",
"y": [
0,
1.5,
2.9475,
4.06497,
5.15115084,
6.206918616479999,
7.23312489521856,
8.23059739815244,
9.684912307430038,
11.078145990517976,
12.412863858916221,
13.69152357684174,
14.916479586614388,
16.67674137265768,
17.996233607475734,
19.24922343365896,
21.03398214207434,
22.353846902121685,
23.593886241481364,
25.341448881654173,
26.61266788761924,
27.791626903007383,
29.43171923183839,
30.600955485874508,
32.202064584257414,
33.315620847712836,
34.81138953064264,
36.07267529243865,
37.668017330680804,
38.644180015829264,
39.83297486150279,
41.11299398144836,
41.851228071663975,
42.48987818649154,
42.839639857364936,
42.8646431272279,
42.84857177265771,
42.87612056815096
],
"yaxis": "y"
}
],
"layout": {
"autosize": true,
"legend": {
"title": {
"text": "Chemical"
},
"tracegroupgap": 0
},
"shapes": [
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0,
"x1": 0,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.001,
"x1": 0.001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.002,
"x1": 0.002,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0028,
"x1": 0.0028,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0036,
"x1": 0.0036,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0044,
"x1": 0.0044,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.005200000000000001,
"x1": 0.005200000000000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.006000000000000001,
"x1": 0.006000000000000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0072000000000000015,
"x1": 0.0072000000000000015,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.008400000000000001,
"x1": 0.008400000000000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.009600000000000001,
"x1": 0.009600000000000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0108,
"x1": 0.0108,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.012,
"x1": 0.012,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0138,
"x1": 0.0138,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.01524,
"x1": 0.01524,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.01668,
"x1": 0.01668,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.018840000000000003,
"x1": 0.018840000000000003,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.020568000000000003,
"x1": 0.020568000000000003,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.022296000000000003,
"x1": 0.022296000000000003,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.024888000000000004,
"x1": 0.024888000000000004,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.026961600000000006,
"x1": 0.026961600000000006,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.029035200000000008,
"x1": 0.029035200000000008,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.03214560000000001,
"x1": 0.03214560000000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.03463392000000001,
"x1": 0.03463392000000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.038366400000000016,
"x1": 0.038366400000000016,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.04135238400000002,
"x1": 0.04135238400000002,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.04583136000000002,
"x1": 0.04583136000000002,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.050310336000000025,
"x1": 0.050310336000000025,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.05702880000000003,
"x1": 0.05702880000000003,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.062403571200000035,
"x1": 0.062403571200000035,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.07046572800000003,
"x1": 0.07046572800000003,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.08255896320000004,
"x1": 0.08255896320000004,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.09465219840000005,
"x1": 0.09465219840000005,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.11279205120000006,
"x1": 0.11279205120000006,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.14000183040000008,
"x1": 0.14000183040000008,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.1808164992000001,
"x1": 0.1808164992000001,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.24203850240000013,
"x1": 0.24203850240000013,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.3338715072000002,
"x1": 0.3338715072000002,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
}
],
"template": {
"data": {
"bar": [
{
"error_x": {
"color": "#2a3f5f"
},
"error_y": {
"color": "#2a3f5f"
},
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "bar"
}
],
"barpolar": [
{
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "barpolar"
}
],
"carpet": [
{
"aaxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"baxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"type": "carpet"
}
],
"choropleth": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "choropleth"
}
],
"contour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "contour"
}
],
"contourcarpet": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "contourcarpet"
}
],
"heatmap": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmap"
}
],
"heatmapgl": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmapgl"
}
],
"histogram": [
{
"marker": {
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "histogram"
}
],
"histogram2d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2d"
}
],
"histogram2dcontour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2dcontour"
}
],
"mesh3d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "mesh3d"
}
],
"parcoords": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "parcoords"
}
],
"pie": [
{
"automargin": true,
"type": "pie"
}
],
"scatter": [
{
"fillpattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
},
"type": "scatter"
}
],
"scatter3d": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatter3d"
}
],
"scattercarpet": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattercarpet"
}
],
"scattergeo": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergeo"
}
],
"scattergl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergl"
}
],
"scattermapbox": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattermapbox"
}
],
"scatterpolar": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolar"
}
],
"scatterpolargl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolargl"
}
],
"scatterternary": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterternary"
}
],
"surface": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "surface"
}
],
"table": [
{
"cells": {
"fill": {
"color": "#EBF0F8"
},
"line": {
"color": "white"
}
},
"header": {
"fill": {
"color": "#C8D4E3"
},
"line": {
"color": "white"
}
},
"type": "table"
}
]
},
"layout": {
"annotationdefaults": {
"arrowcolor": "#2a3f5f",
"arrowhead": 0,
"arrowwidth": 1
},
"autotypenumbers": "strict",
"coloraxis": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"colorscale": {
"diverging": [
[
0,
"#8e0152"
],
[
0.1,
"#c51b7d"
],
[
0.2,
"#de77ae"
],
[
0.3,
"#f1b6da"
],
[
0.4,
"#fde0ef"
],
[
0.5,
"#f7f7f7"
],
[
0.6,
"#e6f5d0"
],
[
0.7,
"#b8e186"
],
[
0.8,
"#7fbc41"
],
[
0.9,
"#4d9221"
],
[
1,
"#276419"
]
],
"sequential": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"sequentialminus": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
]
},
"colorway": [
"#636efa",
"#EF553B",
"#00cc96",
"#ab63fa",
"#FFA15A",
"#19d3f3",
"#FF6692",
"#B6E880",
"#FF97FF",
"#FECB52"
],
"font": {
"color": "#2a3f5f"
},
"geo": {
"bgcolor": "white",
"lakecolor": "white",
"landcolor": "#E5ECF6",
"showlakes": true,
"showland": true,
"subunitcolor": "white"
},
"hoverlabel": {
"align": "left"
},
"hovermode": "closest",
"mapbox": {
"style": "light"
},
"paper_bgcolor": "white",
"plot_bgcolor": "#E5ECF6",
"polar": {
"angularaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"radialaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"scene": {
"xaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"yaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"zaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
}
},
"shapedefaults": {
"line": {
"color": "#2a3f5f"
}
},
"ternary": {
"aaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"baxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"caxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"title": {
"x": 0.05
},
"xaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
},
"yaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
}
}
},
"title": {
"text": "Single reaction A <-> B (no downregulation) (time steps shown in dashed lines)"
},
"xaxis": {
"anchor": "y",
"autorange": true,
"domain": [
0,
1
],
"range": [
-0.00023121295512465388,
0.3341027201551248
],
"title": {
"text": "SYSTEM TIME"
},
"type": "linear"
},
"yaxis": {
"anchor": "x",
"autorange": true,
"domain": [
0,
1
],
"range": [
-2.7777777777777777,
52.77777777777778
],
"title": {
"text": "Concentration"
},
"type": "linear"
}
}
},
"image/png": "iVBORw0KGgoAAAANSUhEUgAAAu8AAAFoCAYAAADn3hWsAAAgAElEQVR4XuydCXwWxfnHfyQhAUIS7iOAIMolIAgKiqAi3ngAVYwnWNSqrbZWW+ttvWrr0cPWq+ofj1o8KggIXqgcotKC3PcthxwJEEhCLvjvLG5YNrvvnrPvsb/lkw/JuzPPzH7n2ZnfzvvMbJ2DygEeJEACJEACJEACJEACJEACCU+gDsV7wrcRK0gCJEACJEACJEACJEACKgGKdzoCCZAACZAACZAACZAACSQJAYr3JGkoVpMESIAESIAESIAESIAEKN7pAyRAAiRAAiRAAiRAAiSQJAQo3pOkoVhNEiABEiABEiABEiABEqB4pw+QAAmQAAmQAAmQAAmQQJIQoHhPkoZiNUmABEiABEiABEiABEiA4p0+QAIkQAIkQAIkQAIkQAJJQoDiPUkaitUkARIgARIgARIgARIgAYp3+gAJkAAJkAAJkAAJkAAJJAkBivckaShWkwRIgARIgARIgARIgAQo3ukDJEACJEACJEACJEACJJAkBCjek6ShWE0SIAESIAESIAESIAESoHinD5AACZAACZAACZAACZBAkhCgeE+ShmI1SYAESIAESIAESIAESIDinT5AAiRAAiRAAiRAAiRAAklCgOI9SRqK1SQBEiABEiABEiABEiABinf6AAmQAAmQAAmQAAmQAAkkCQGK9yRpKFaTBEiABEiABEiABEiABCje6QMkQAIkQAIkQAIkQAIkkCQEKN6TpKFYTRIgARIgARIgARIgARKgeKcPkAAJkAAJkAAJkAAJkECSEKB4T5KGYjVJgARIgARIgARIgARIgOKdPkACJEACJEACJEACJEACSUKA4j1JGorVJAESIAESIAESIAESIAGKd/oACZAACZAACZAACZAACSQJAYr3JGkoVpMESIAESIAESIAESIAEKN7pAyRAAiRAAiRAAiRAAiSQJAQo3pOkoVhNEiABEiABEiABEiABEqB4pw+QAAmQAAmQAAmQAAmQQJIQoHhPkoZiNUmABEiABEiABEiABEiA4p0+QAIkQAIkQAIkQAIkQAJJQoDiPUkaitUkARIgARIgARIgARIgAYp3+gAJkAAJkAAJkAAJkAAJJAkBivckaShWkwRIgARIgARIgARIgAQo3ukDJEACJEACJEACJEACJJAkBCjeXTbU6+9+jD/+49/48+9/jnNOP8llbiYngfgR+GT6f3H7g/9QK7Dky7Hxq4ih5CUr12PkjQ9h9OXn4Tc3FyRMvZKhIkNG/hqtWjTBv/5+XzJU94g6XvWLRzF/8WrP7a75jTD6zksPoXvnDknHIGoV5r3ur8W7nzEa55/ZH089cLM/Qya5w9Q2bspK5D5OtIc44qEHE1686wWH3t+MA/2Tz4/D2Lc/wl0/vwLXXnZu4I6tGXTjdNIqIdlwrA42LM5OLvHOh5/H1M+/TTgx6qTuTtJovmaW1ksHLjoaIfSmvfOMk+JDS8MB3TtqPwObVV7N78J4mNL6Ey+Dn6j/D9uLjngQTZa+y02La2Og7LHNTZ28pk21ez3s8ZDiHdDuBy9joFe/jZVPm4QIe0IsocW7Js6MnZZ2w+gbL6ybiOI9nIckJzeZ9tQr0gY9sGnCwO2MnuazXsSI8Zo1XzNem5fOS7s/3F6Pk3bwmybVBnS/PNzkT3bxLq5VXIM43DxUWvXDFO9uvCf8tKl2r4elO7SWonhPPPGu+XTYDxMJK941gRJr9kcIJRlfH8Xq0qIu3sPv7s1L1LeDCAXp3ePYQEIHtKdoUarXJ2n9NwJ+RLyVeBd1E/UUs45OBY/MTt+vT6TagO6Xh5v8MsS7m/KDSOtlZtnqulPRl7zwCaJdZNhIxfaRwcnKpsx+PExt46YsP31cWG0Tj8mxhBfvTmdUzZxBE1FChOlnaa2EmTFMQYQYiEMfUxrL6YxlOK27vuEf/9ubahyoOPT59YJQnDMTq8byrdKJz83CkYTNe267Wo0/Nh5aeVbXb2bPOMurz/uasnZAu05RlluRqxevGhs/s8pBiHYjM78iPpZ4dzNbGctn9YOpqL8IPdMOqwdn7VsJu3RWHafRl8X9JR5EzMqzu6fMOnazjtQogPR/L1y2tib8ynjfib+Duj+FLbP72Bj3re+39AzNrsvs+p30A8Y21MoRfaWVWNTHmGvpjfetG676a3MzQLupn74PNLsPtH5E9LXaehCzPFpdrWbXtLbR0rkJTzPrO7V7wSps1FgPu3vSbd9rtOf0esx8RF9XGf2N2f1i5SN68eulLsY+LZZP/e3R244YS50y1Mow8ynRT+p5Og0rthrz9X2uWx8RNu36Z+1azPxJXIuTcd/YN5g9AOr7xtvu+5s6nmiHmS5w0peJ/HpdoO8jjX4Qj4fShBXverhOZkBjiXcBWm8j1oCvdyatDnqhbFaOWRiDk28OzG5SM2c2i+00E2/iRjI6qvjMKPSt4kxFWsEpliOaXb+ZPbPP9A9H+uu0EitW4s9YPzesjTZliHZjGXrB5uYBw0q8xxL1ZsxE+d8tXmU6S6+/z6w6cm1httnXg26/MjSLDzRrP6efmfmOdr/or8cofPWDnv4h2Uwg6wdRP/en2bWbxZcHId6d9ANWgtlM+Jj5nNlnbrjqfdVNH6C1h9m44Lbv0tpE30fq7wmzz40PmZoNPXOncbBmDI19Z6yZd7MxyuyetOt79XU3jhlO73Gze9Z4fTL6m1hjv779jBzd1MVqLLIS7+KB3CjW3cyam01ImbWDuCaxgYb+G1gn7SCux+hndj6iv9/caB43GsSMsxvxLvIb70PjN9Rmfa7ZfWilsay+5XczAWHlT24+T1jxLi7COBOuXZjZAGo3866HYhygY3X4xgYxK8eq0WINMrHq4+ScVme72X1jHZw8IbodAK06JWNoh92svd21GB929Depm5lofcdlfLBzc/O4Tet2ga2V/8eaUTCrk2gHcZjtSBKrrY3taiWwYs3s6+tjlc6sDlZhQcZ712hTL2b012x8gLGbvY0l/L3cn26u3a94N/MBs77IjXi3SmusqxuuZr7h5OHWycOo2bc4sWZJjeFnVtfrZDzQrkvcP3aLfq3aWnAUh3hwjiXere4T7Vo1nrHuUf19Hqss4UOxdoOyCh0Q9+TcBSvUjSRk9DdmNrV2EiJaE51GJm7qYtXXu/Eppw+osdrA6QOAsc934mduxmenmsfvmCG4uxHvsb7t1yahrBjq+5VYvmF1H8Tql9xqBSfpE1q86y/A7KtgvdjzI95jdWx2nbUToWs3IMWKl4oV22w2OBi/ljcKPSdCy8k1aQ9Qdp29CMOwG0Bi2TBzYqffnMS6AfSzqU6+2RG2zMS0m69CvYp3s4caNwulBK8TenQyXR/iZgCz6rCdPBAKflYduVl+qw7WzFf094HgMn/JaoxShIIIg9B8T9T9vMH9asSHG5EZxP3p5tqDEO92/YDZoKjdL1YzlE4EsRuuXsW714dRN0LLqt81fh5LkMW677Rr13wrVj/iRcxpebR+Ola/r78m/Wy0k5AGszYUn1n1qbL6G71PaNeuhctq/adoq63bC2smMdzUJUzxHqu/seoXzcLg9D7lxM+cincn+kDf75ptZetEh2jMgxTvTvSK5rua5nQ6qUjx7uSRQ0mjgTU+VTsJxzDeHLFuFjvxbhVzpr8Mu07Qrnx9/JYRjxb/pq+HvuN0c62abSc3p5NBwepruVhxsnbbfNrxdrviOxnDZvSdmvjdbtFqUOLdauBw+rW6lSAy+lss/zP7Slg/cItrFcJd+JEm2C8YcrIae6r3OzciM4j70+m1x3rIMauHsX9y2g+4Ee+xBjzj181uuKaCeDeLidVfl5OF9PqJBC2vWVyzUUTohbbVsKnlcSreVeH943sXjDadTHKYTXA4CT0SZRn7Fzf9jf6bJSGixCE2s9ALduMDfKKK91gPhEYm2nUb/cysv7HzM6fi3W4MFuy1vtaqDeMl3u2+zRZ1N4aQGe8Dq8lYineH4t0YE+Zn5t1ucI61YNVp+Eqsy7Kb2RN57V7C4nS2zslN40a8O3mSDXLmPVbH5jTO1KwtZIh4/eyn3QOcWZ3M4vD06Zxer9eZSmPH63fm3Y2AdTPzrr9/hEjX/E0buM9VQg9EXKhZzKaVIHIaNhOLrbGtzHYHMrvXnN7LZgLcT16tvok+8x5G2IybmXer9SQOh7JaybT7WvNNu5l3u9AcUYCbb5eNFdLyOnkQMebV/FGroxvB7Ka/0X/TIO517QFeHypjfIB3UxertnTzbY7TsBk34t2KUaxv7LVrMfqZU/HuRvMkmng3fiPl5h7V8lp9S0bx/iNN4UgCktVbTJ3EUzsdyGJ1jnYz72YDqBuHEGljifdY55yIAzfx/fpYLqt4TbMbPMyYd7PFLxoHO7HrpF2CEPF+RbvT63Ea5+9V7AQd8x4rJtb4hlWnMe+ClXb/CnGhf9C1+lyfx694d3p/url2p/2WWd/jhpvVg4dZfyg75t2psNH3l1YzwW76LqciXS949A9hTiZDYvU74rqN2x1bfRNl9vW9E5EWS7wby7Iad52UY3Ytomx9P+JGMFv5hBVzbWzQx7lr5Yu+wfi5m7qEKd6drk8wso2lB5z4mVPx7kbz2K3JcDKpFWTYTCxmen6iDxT3uTESwC5MLsy3XSdszLv+6w1jJ60BDCrmXTSa2SymVo6X3WY0R9B2cInVgdsJAC2ezYzD8d06qg5mDFHRD3Lid/1XPdp16W8c/deOGg+zmUKzG9ws/tqsPm46BzNeTsR5LHHvRLxraTTmdusVjDbN2LopV5821vVq/uqk84s1GLgZwLS0Zl+DOw1XMmsfLQROP4OoCUiz8IFYW0oaBY7WjsY8bsI7grg/9SJC/y2aWf3M6qb/ytu4UFs/YLjpB6yuy6x8zYfMdiTSM3fDVe/rVg8HZvdOrMkWt32XX/GuHzuMfYXgK45Yizy1+1if16wN7WahzWbFRR6xXWH3zh1qZt7N7g9RRy30zqydnYbFmfV9xj5MZn+jsTT2RWbjuLhmN3UJU7xr4ljfLtpnxq0izXSL9pl+htiJn7kZn836ZzPNYxbmqA+7cTJ+BS3ezXxc8wexzaS4F6yuz0pfOF335VULmOVLWPGuv7mMFTf72sJP2Ixm3xi/KDo6sR+5frGf3VOxsa5ORI2dOBA29bO5+jLM4tu182KgFYd+0ah2ziz2y2x7OS190Pu8679RsRuMje0TK/bSGE4V5M0Sti27+DwnMaj6DtXMF70MYMbFUU6+ttezMy4+F35nnHnX11uf12rxkFUIkZkQEvbciMyg7k9RrvHaxeAlFtYar8vY9lb3cqzF2076AWOfF+Q+7/pZKyfrGJwuDNOETKxZLj3nWH1XEOJd1Mcsnlh87uTh32yxoTGfMQ7dbp93UbbZpJP2TgVjv66/x8yuxek9bvQnYdds+2WrB3CzPsppf2N3rxvL9NL3GccAGWEzWhnG69b6SSMjI3PBW+gW4+SbnZ+5Ee9W/bP43Fg/Y4y88EstlDEe4l3U0Wpdh9nkhL7N7d7z4OR+D0pHJLR4D+oi/dix+grWj03mJYGwCTgRoGHXieUdfohwMoilKi+n4V/66/cbrpKqLK2ui7yi1uK83rAIOP1mKuj6ULz/SNTsK043cZhBNwztkUDQBLyIpKDrEGV7YoZMLKTTf+vkJlwkFdlZzZY6uVanC7ad2Er1NBTvqd7CvL54EYhXP0TxrhPv+lfDi4/d7OEdL8dhuSTglIDVNoJO8zOdPwJmIQVOwur8lZq4uTUeTkMyjFei/+o7zK+rE5eodc0o3pOx1VjnRCeghefF45tTivdE9w7WjwRIgARIgARIgARIgAR+JEDxTlcgARIgARIgARIgARIggSQhQPGeJA3FapIACZAACZAACZAACZAAxTt9gARIgARIgARIgARIgASShADFe5I0FKtJAiRAAiRAAiRAAiRAAhTv9AESIAESIAESIAESIAESSBICFO9J0lCsJgmQAAmQAAmQAAmQAAlQvNMHSIAESIAESIAESIAESCBJCFC8J0lDsZokQAIkQAIkQAIkQAIkQPFOHyABEiABEiABEiABEiCBJCFA8Z4kDcVqkgAJkAAJkAAJkAAJkADFO32ABEiABEiABEiABEiABJKEAMV7kjQUq0kCJEACJEACJEACJEACFO/0ARIgARIgARIgARIgARJIEgIU70nSUKwmCZAACZAACZAACZAACVC80wdIgARIgARIgARIgARIIEkIULwnSUOxmiRAAiRAAiRAAiRAAiRA8U4fIAESIAESIAESIAESIIEkIUDxniQNxWqSAAmQAAmQAAmQAAmQAMU7fYAESIAESIAESIAESIAEkoQAxXuSNBSrSQIkQAIkQAIkQAIkQAIU7/QBEiABEiABEiABEiABEkgSAhTvSdJQrCYJkAAJkAAJkAAJkAAJULzTB0iABEiABEiABEiABEggSQhQvCdJQ7GaJEACJEACJEACJEACJEDxTh8gARIgARIgARIgARIggSQhQPGeJA3FapIACZAACZAACZAACZAAxTt9gARIgARIgARIgARIgASShADFe5I0FKtJAiRAAiRAAiRAAiRAAhTv9AESIAESIAESIAESIAESSBICFO9J0lCsJgmQAAmQAAmQAAmQAAlQvNMHSIAESIAESIAESIAESCBJCFC8J0lDsZokQAIkQAIkQAIkQAIkQPFOHyABEiABEiABEiABEiCBJCFA8Z4kDcVqkgAJkAAJkAAJkAAJkADFO32ABEiABEiABEiABEiABJKEAMV7kjQUq0kCJEACJEACJEACJEACFO/0ARIgARIgARIgARIgARJIEgIU70nSUKwmCZAACZAACZAACZAACVC80wdIgARIgARIgARIgARIIEkIULwnSUOxmiRAAiRAAiRAAiRAAiRA8U4fIAESIAESIAESIAESIIEkIUDxniQNxWqSAAmQAAmQAAmQAAmQAMU7fYAESIAESIAESIAESIAEkoQAxXuSNBSrSQIkQAIkQAIkQAIkQAIU7/QBEiABEiABEiABEiABEkgSAhTvSdJQrCYJkAAJkAAJkAAJkAAJULwH4AN7Syuxt6wqAEs0oRFokpOJ0vJq7K+oJpQACWTXy0BGeh3sKakM0CpN1c1IQ6Psutixp5wwAibQqkk9bN9VjgMHDwZsOdrm2MfKaf+o9bH5TevLAUmrMQlQvAfgIBTvAUA0mODAEjxTYTFqA4scirWtUrzLI03xLoct+1g5XKPWx1K8y/EjO6sU73aEHJyneHcAyWUSDiwugTlMHrWBxSEW38ko3n0jtDRA8S6HLftYOVyj1sdSvMvxIzurFO92hGzO//73v8dlBVdh1syZyG/TDhXl5Vi/bhUa5uShRcvWKCsrxcpli9C5W0+0zm+LosIdyG6Yi8qKQ1+v183MQsm+YrTKb4dVyxejR6++mDtnNi4cXoDCndsx/bMpGFEwGp9OGY9OXXsgJzev5jORf96cr1Q7ffqdavm7dgn//PufcMMvfqv+uWj+/9RyTx54pvr3uNdfxNBhBap97dDbFp9t3byxpm5GLJPHj0PffgPQus1RNaemT5uqXrO4drPDrEwtXVXpLkyePBHDRo5y1ELfzPpc5dqz94mW6devXaUyPvuC4Y5saolE+23dsgmnDznfVb5YiZ3U121hmo906NjJMqusgUUGI7fX7zW98V7wYidVxbu+z/DCJYg8FO+xKXq99yjeg/DO2jZk9bFyauvPqugfHnzwQX9GmNsTAYp3T9gOZ6J4P8SC4t2dI1G8u+MlMzXFuzVdineZnheMbYr3YDgGZYXiPSiStBOLAMW7T/+geKd49+JCFO9eqMnJQ/FO8S7Hs8KxSvEeDmenpVC8OyXFdH4IULz7ofdjXsa8BwDRYIJf6QbPVFiM0sAih6C51VQNmwmToVVZDJuR0wrsY+VwjVofy5h3OX5kZ5Xi3Y6Qg/MU7w4guUzCgcUlMIfJozawOMTiOxnFu2+ElgYo3uWwZR8rh2vU+tgwxPslo+9F0ya5ePWZu+Q0mkSri5atRcHND2Pc8w+gZ7eOgZVE8e4T5f4DB7B+bzlyuc27T5JHZufAEijOGmNRG1jkUKxtleJdHmmKdzls2cfK4Rq1PjYI8f7TX/8R385bdkSDNGmUg5kTnlU/i4d4n/DRLNz7xMt47HfXY9h5Az07C8W7Z3RyM4qY93mDz8bQlSu42wx3m3HsbIx5d4xKekLGvFsj5oJV6e7nuwDGvPtGGKiBKIn3IHab6X7GaOiFutYYQtC3bNYYf7jnxriI96CcguI9KJIB2xHi/d8DT8MNa9agS7sO3CqSW0U68jCKd0eYQklE8U7xHoqjSSqE4l0SWI9mKd6dgxMCfdXaTTUz7FY5tZl3cV6bobcS/PoZfH2oyqBht2Jgv56YNWcRinbvVYu66ZqL0a5NC3WGXTu0PGai2/gNgch/65gRMPvmYMmXY1WTFO/O/SHUlEK8jx0wCFevWY3+7TtSvFO8O/I/indHmEJJRPFO8R6Ko0kqhOJdEliPZinenYMTs+4XnzNAnV2PdQjxvnr9ZlVsC7EsDiHGO3VsWxMHLwR0YVExPhj7mHr+2VfexwtvTIQmokV6Ido1ca6dN4bniLzChlF0Gx80xPk///NdtXxx7vYbLquJaRf1tbLjnE7slIx590ny0737cO7qteiVVQ9vNW+DvLQ0nxaZXRBgPKYcP4jSwCKHoLlVxrzLo82Ydzls2cfK4Rq1PtZrzLsmjp3ElJvFvN/9+EtYunKDqdDWWlYI9pEXDVYFvzbzrj0omM2IC5tiZl7E2uvPC3ti0amTumoPDu9M+qKWHS5YlXPPebZ69sq1+KxkH55r1gqXNMjxbIcZDxPgwCLHG6I2sMihWNsqxbs80hTvctiyj5XDNWp9bCKId21xqVmLarP1VuJdL8jFbLyZ6F6zYYsaWqPN4puVo83s68+J9AybkXOfBWL1iU3bcPeObRiZnYs/N20ZiM2oG+HAIscDojawyKFI8R4WV1EOxbsc2uxj5XCNWh/rVbwL+m7CZoxbRepn3jXxbieuRcy7ceY9CPEurqN/n241ITz6kB2Kdzn3mW+rIua9/4iRGD/jC2xp3gKXZGSieMNaNMzJQ4uWrVFWVgoRk9i5W0+0zm+LosIdyG6Yi8qKcrXsuplZKNlXjFb57bBq+WL06NUXc+fMxoXDC1C4czumfzYFIwpG49Mp49Gpaw/k5ObVfCbyz5vzlWqnT79TLX/XLlK/c4Qxznfc6y9i6LAC1b526G2Lz7Zu3lhTNyO4yePHoS93m3HsT4x5d4xKekLGvFsj5m4z0t3PdwGMefeNMFADURLvfnebsVuwKgS61W4zZmEzscJa/My8CwexCpsxe3CgeA/0lpJjTIj3ywquwmtfTMOcRo1xenoGcjdtpHhXcE+fNlV9YBEPLmaH2QODlq6qdBcmT56IYSNHOWo4J2J4/dpV6gPS2RcMd2RTS+R1cIxViJP6uqqkklh7wOvQsZNlVlkDiwxGbq/fa3qKd4p3r76TCPm83nuceZfTerL6WDm19WfVr3gXpZttFakJYm0xq13Mu7Cj7fiin30XAr9/n+PUfdr9iHcRqy7qULS7uGZnHG3BqlioahT24prEwbAZf/4lNbcm3j+Y/iU+yc3FMQcPotsPW5HDmXeK9xieR/Eu9bZ0ZZzineLdlcMkWGKK98RqEIp39+1httWifhbdiXjXC3h9DfS7zXgNm9EWmmq73mj2tTqKh4SJn8yuKVbE2Ws73TBsxr0/hJZjb2klNpdU4Modm/Fd+X68oew6c2b9BqGVn4oFcVZITqtGaWCRQ9DcKhesyqPNmHc5bNnHyuEatT7WT8y7nBaIhlVuFRlAOwvxvresCk/uKcJf9hTi+pxG+H3j5gFYjq4JDixy2j5qA4scirWtUrzLI03xLoct+1g5XKPWx1K8y/EjO6sU73aEHJzXxPuc/WW4ascW5Gdk4K0WbdBGiX/n4Y0ABxZv3OxyRW1gseMR1HmK96BI1rZD8S6HLftYOVyj1sdSvMvxIzurFO92hGzOazHvs2bORH6bdvhwdyHqKQtWm+Y2Qg9lBxnuNsMFq2YuxJh3nzdegNkZ824Nk7vNBOhokkwx5l0SWI9moyTeg1iw6hFz5LNRvPt0AaN4X1BSjJ3r1iC9YQ4Gtu1A8c7dZkw9jOLd540XYHaKd4r3AN0pdFMU76Ejj1kgxXtitUeq1obi3WfLGsX7zrIyLFmzArsaNMCJbY5CXkUF93nnVpG1vIzi3eeNF2B2ineK9wDdKXRTFO+hI6d4/5EAZ97j53sU7wGw12LeNVP37dqB/9u7G7fnNcWdeU0CKCF6JhiPKafNozQrJIeguVXGvMujzZh3OWzZx8rhGrU+ljHvcvzIzirFux0hB+eN4v3TshKMVhaunpBVD28p20bmpqU5sMIkegIcWOT4Q9QGFjkUa1uleJdHmuJdDlv2sXK4Rq2PpXiX40d2Vine7Qg5OG8U7xUHD+DK7VvxdXkpXmjWGhc1aOjACpNQvMv3gagNLPKJHiqB4l0eaYp3OWwp3uVwjVofS/Eux4/srFK82xGyOW+Mea8oL8f6dauwR3lJ0+ycHPQ8cAB5a1ejsxL33VpZvFlUuAPZDXNRWVF+aNDPzELJvmK0UnamWbV8MXr06ou5c2bjwuEFKNy5HdM/m4IRBaPx6ZTx6NS1B3Jy82o+E/nnzflKtdOn36mWv2uXoN85whjnO+71FzF0WIFqXzv0tsVnWzdvrKmbEcvk8ePQt98AtFbi/LVj+rSp6jWLazc7zMrU0lWV7sLkyRMxbOQoRy3kJIZ8/dpVKuOzLxjuyKaWyGtMaaxCnNTXVSWVxJqPdOjYyTKrrIFFBiO31+81PWPerclxtxmvXhVePq/3HsW7nDaS1cfKqa0/q4x598fPT26Kd5ZU/i4AACAASURBVD/0lLxW4r2OstvM5w0bIKe8Al02rKd4N+FM8Z6Lnr1P9OmBh7NTvHtDSfFO8e7NcxIjF8V7YrSDVguK98RqDz+16X7GaBzboQ0+GPuYHzNS8lK8+8RqJd4b5uRhQV4utuzbi94bN1C8U7wfQYAz7z5vvACzU7xTvAfoTqGbongPHXnMAineE6s9vNbm2Vfex2cz56JodzGe+8Pt6Nmto1dTUvJRvAeA1Rjzrpl8T9nz/ZeF23B6/Wz8q3k+6gRQVlRM8CtdOS0dpYFFDkFzq4x5l0ebMe9y2LKPlcM1an1sqsa8XzL6Xpw1qC++W7IKLZs1xh/uuVGOw3i0SvHuEZw+m5V4315dhau2b8HSynK8qew6M1iJg+fhjAAHFmec3KZKhoGlqKwQJZX7jri0ksoS7Nq/84jP9leVY0fpD7UQbNq7sdZn20u3obz60DoT7Sgu34095XvcIjRNn6Y8mWek10FF1cFA7NHIYQKZddNQWXkAUSW7Q/Xd/YG7RFqdOjh48GBkuQYO9EeDClZ1ou5ABBx2zPE/x4NDfuML5XrlXTjrlfDisI8OWZnokJlpWuyiZWtRcPPDGPf8A1izYQuefuFtzJzwbNhVjFkexXsAzWEl3oXph3bvxD+Ld+HKnFw82bhlAKVFwwTFu5x29ireixWRu0cRu0X7C1GqCGtNYAtRXKGI4q0lm1F1oFqtdHnVfojPjcemvRtqfbZ1n8hXJediaZUESIAESEAagTv63Yunzn/Ul/2/bN+B2zdv9WXDS+ZftWiOP7dpbZpVC5nRYt1F7LsQ8okUOkPx7qXVdXlixby3aNka65WdZPasXIaV7Y/G0A6dUG9PEXeb+ZEfF6yGu2BVCO6Dafuxc/82FO3bp6zH2IxqRTiLmeqqg9UQQlr8LT4XM31ixk/MgIt8dkdv9EYH5d8E5Z/fo0n9psiue+T2qtl1s9G4XrMjTNfLyELzBq1qFdcu5/COR9rJ5g1aIis964i0uVmNkJeVh52rt6KqrAKterb3XPX09DQ0rJeBPSXhzyB5rrSDjIvHf4Mew092kFJeEvEgv2tfpTpLHMXjkO/Ws7z0zavXoUgRQD0H9HOFp1HDuiirOIDyikMP3TyCIdAgKwPpyrdwYlIv1Y+PXn8bDz74oK/LHFtYhNeKdvuy4SXzqCaNMLqp+Us0tZCZW8eMUE3/9Nd/TLjQGYp3L62uy2Mn3svKSiEWFM0/qj2Oa9sefUtLKd5/5Efx7k+8i9ltIbi1n5KFhdjesBA/1FPEuSK4RVhIccWhGXMxc+7nEGJaiOrczDxoolf836SeENrZaFLcCHX2HET94/KQlVEPLRTBYTza5tQWx60btkFGWoafqvnOywWr1gi5VaRv95JugAtWpSN2VYDXbzddFZIgiVNxq0gtZMaIuEmjnIQKnaF493kTuBHvRc2a4xfVB5Gr7ETDfd4Bindr8S7EtpgB14T5pn0b1d+/L95Q85kxDrYABZiv/Fuu/DM7cpVZ5kb1GqFZ/WbKTF62KrCF0FYFdJ10tP1xxrpdbnuk18lQP7cS4kb7XgWEz9svkOwU7xTvgThSnIx4vfcYmiinwSje5XANy6oxZEYrV4TOPPa76zHsvIFhVSVmORTvATRDrJh3Yb5K+br3gh++xxJl4aqIexfx7zxiE0j1gWX9njVHiHFNnAuBLkS7k5lyIcbzFYEtRLb4advwqCMEdwMxW67OjB+aNRdHlAaWMO8x7jYjjzZ3m5HDNtX7WDnU7K1GrY9Ntd1mBg27FSMvGgwtZEZrcRE6I45Xn7nL3glCSEHxroN89+MvYeIns2stTBDxT6vXb1ZTmm3YbyfeRb439+3BXUXbcUpWfbyqbBuZm5YWQvMmbxGpNLAIob5kx0LM2/ZfLNm5EAu2z7UV5yLGVRPlYiZcxHG3yj4s1IVoF+Ld7RG1gcUtH6/pKd69krPPR/Fuz8hLilTqY71cv6w8UetjU028y/KLoO1SvP9IdMJHs/B/46aqIl2/qlg8bRUWFde8YUsI+aZNco94+nIi3ncqO3Fcv2ML/lu+H083aYGChu6FV9CNn8j2knVgcSrUhfDu2rS7IsrbqyK9VXa++r8Q6q0Vka7NlAfdRlEbWILmZ2WP4l0eaYp3OWyTtY+VQyM4q1HrYyneg/MdN5Yo3n+kpW0FpO3tqW0JJL5CueOmy2vinITI1+/56TTmvXO3nljRpAm+3boZLXLz8JPMehBz73Uzs1Ci7EjTKr8dVi1fjB69+mLunNm4cHgBCndux/TPpmBEwWh8OmU8OnXtgRwlr/aZqPq8OV+pV9Cn36mWv2sOoV98ZozzNYs/19sWNrZu3lhTN6OTTR4/Dn37DUDrNod3+pg+bSpa57dV3y5rdiRzzHtldQVWFCm7CO1SfgqVH+V38bcQ78ajfV5HdG7cFZ2bdFN+uqKT8v/e5TuRl9sEPXuf6OZ+jZlW85EOHTtZppM1sHiNuw3s4n0YYsy7NTwuWPXhWCFl9XrvUbzLaSBZfayc2vqzmooLVv0RCS83xbvCWsymX1dwPo5pn1+zMb8Q7/qN+jUxb/xMiPeRBVdj1qwZyFeEa7kys75+7SpVZIutIsVuMyuWLkKX43ois2U+3t+0HmuUlwNcpoj3LnWzkKmI931KSE3r/KOwcvki9Ox1Iv6nCPKLR1yBnTu248vPPsSlV1yHjz98H5279lTtap8JN/nft4fE+4n9T7X8XXOnF/72R9x026F4rYXf/U8td8CgIerf/xr7glqmsK8detvisy2bNtbUzeiiE9//N05UHiDy2x4W718oDx75ykOJuHazw6xMLV15SREmT5qInygPLk6O2TOnoaHybcbxJ1iL4XVrVqmMzx16aPsnp8fSJfOxYt0SHDg2XRXrK4RYV/5fu2t1LRMdGx1bI9KFWO8ifpp2Q6Zhm0In9XVaPy2d5iNHH2Mt3mVtYyZ8fMuW7zH4rAvcVjvu6Y33gpcKiZn33AYZKCxOra0i9X2GFy5B5GneKAuFeyqUl95Ec6tIO4Ze7z1uFWlH1tt5WX2st9rIzSX6B79bRcqtYepaj7x4F3Hu23buUsNgjMLcqXi/9tprMX36DLTv0B779+/HyhUrkJfXCG3a5KOkpBQLFsxHr169cVT7ozBp3Tp8pIxBJ6ZnYESjXGRlZaF4T7F6buGChejXvx9mKLauUWxu++EHTFQE7A033Ih333kHx/c6Ho0Uu9pnwi1nzJiueudpp51u+bvmvo8+8jDuu/8B9c9vv/1GLffsc85R//77s3/D1ddci0aNGtV4u962+HDDhvU1dTPeEm+8/jpOO/00tG/foebUpIkT1esS1252mJWppdu+bRsmTvwA1yvX7uT49JNPkJuXi/79rfekXrFiucr4spEjbU0u3b4E3275Ft9u+hZb12xF3eKMWnuYd23WFd2aHYfjmh+Hbs274Tjld/F33fS6tvad1NfWiCGB5iNdunS1zCrr7X/Cxzdu2IiLLr7YbbXjnt54L3itkHhjZaoJTH2f4ZWL33zpyutrq6PwukqPoLzee3zDqkfgNtlk9bFyauvPqugfKN79MfSaOy7iXYSiFO3ea1rnJV+O9XotrvMZQ2C8iHdRqJOYd61yyysrMEaJfd9SXY1XmrfGmfUauK53FDKE/ZWueMvnvB/m4MvvP8OXGz5RFpXOOwKziEHv07IfujfvheObn4BeLfqoMerJdkTpK90w24Yx7/JoM+ZdDtuw+1g5V5F4VqPWxzLmPT4+GLp4N1vwGZ9LB4R4v/eJl02Lv+mai9Wtgsxi3kUe/UOGG/EuCnt01048v3eXumWk2DqSR20CYQwsQqB/p+wAI0T7XOVHH6/eIruVItZPQp9W/Wr+j/WWw2Rpw6gNLGG1C8W7PNIU73LYhtHHyql5YluNWh9L8R4ffwxdvCfaRvd67GZhMkHtNqMvZ64SFy92nqnAQbyszL6fksXZd6P7yxhYxEuNZm+aienff4ovNnyK1btWHFHssY27YFC7wRjS/nz0yz9F3R891Y6oDSxhtR/FuzzSFO9y2MroY+XUNLmsRq2PpXiPj39SvOu4m4l3cTrWPu9udpsRO68UFe5AdsNcTN1ThAUV5ejdMAeDqg9wtxmD/1eV7sLkyRMxbOQoR3fGN7M+V7kad28Rgl0I9SlrJmDN6mXoWt0V45R/4hDi/Iz2Z2NQ28E446iz1W0azQ6vuznEqrhVfR1drEUi7jbjjR53m7Hmxt1mvPlUmLm89k8U73JaKUrinbvNyPEhJ1ZDF+9CCJ81qG+tt1c5qWwipvEq3leX7sNE5adOZl0MP5iGnu06cKtIXQP7Ee8ifv3rzTMwcfV7mLx6fM0LkbqiK86oNxhZPfNwRruz1JCYjLQMW7fyOjhSvNuiTYgEFO8U7wnhiB4r4bV/onj3CNwmG8W7HK60eiSB0MW7cZFosjeIV/Feqcy6Ty0rwbyD1Tij+iAuOroTxbsP8T5j+kfYXr0dizMXY9amL7Fs52LVmohTP7Xt6epPtzrHoWJLCc6+YLgrt/M6OFK8u8Ict8QU7xTvcXO+AAr22j9RvAcA38QExbscrmFZ1SIwjOU99rvra973E1ZdYpUTungXMe+xjjB3mwmqAdwuWNXK/UgR7yL2vZOy3/vLzVvhmIzMoKqU9HacDixLdy7Cx2sn4eN1k7Fox3z1utPqpGFAm0OCXfz0bdU/6XkEdQFRGliCYubEDmPenVDyloYx79642eVy2sfa2eH5IwlErY9NtZh3s/BpsaX4rDmLMHPCswnj7qGL94S58gAr4lW8iyrcsHMrpijhM3c3aoZf5DYOsFbJbcpuYBE7w7zw3V/xzrI3IeLaxSEWnA49djhGdL5c/Z1HbQJRG1jC8gGKd3mkKd7lsLXrY+WUmvpWo9bHRkG8azsTJtLkMsV7AH2JH/H+n5Ji3Fa4Db2VN62+0rwNWqWnB1Cj5DdhNrCIWHYxw/7J2snq/3sripGTmYtzO16Ec4++UP1JTyO/WK0ftYElrDuB4l0eaYp3OWwp3uVwjVof61e8r9+9HuIn7KNDow4QP8bDatdBkU68zDNRjriId7P91RMtnshpA/mJeVfLyMzEB9u3YlZeHq5Q3io6tO/JmDtnNi4cXoDCndsx/bMpGFEwGtpOIjm5eTWfiezz5nylmunT71TL37Vr0e8cYYzzHff6ixg6rADCvnbobYvPtm7eWFM3I5/J48ehb78BaN3mqJpT06dNhdhhp3O3nqY4zcrUEuoXrO4s26GK9Y/XTMK0DR+pSY7K7aCIdkWwK8L9lPxBcLJ7y/q1q9R1BYx5r4M9JZVOXdxROq9xt46MS07EmHdrwNxtRrLzBWDe671H8R4AfBMTURLvQew285dv/oLbP75dTmPEsPqrk3+FP5/751oprGLetXf/hF5RiwJDF+/PvvI+XnhjIsY9/wB6duuoVkuDlWhwnDSSX/FeV5lx/65oB17LboABmzfhlv6DKN4V8EK8/2fCu9jZda8S0z5ZeYnSt2pziDebqrPsimjv2rR7TRNRvKPmAa9Dx06WritrYPEqIJzcY7LTULxTvMv2MZn2vd57FO9yWkVWHyuntv6sBiHex84fi9cWvOavIh5yj+o1CqN7j7YU73qNyrAZBZN4Y+nIiwbX2ipSiPp3Jn2RUAsCnPhDEOJ9795i/LV+FtpuWIe+vfoid9GCSM+8F5fvwYuzn0Tl0nL84+A/1Gbo2vQ4/Kz3L3FJ58vUHWSMB8U7xbuT+9UsDcU7xbtX30mEfBTvidAKh+tA8Z5Y7eG2Nlbv+xGbregFvVu7QacPfebd6g2rifhk4xS2n5h3rYx/Fu/GQ7t34JR6DfBc01ZoEcHYd7Hw9B9zn8Y/F/y9Zm/2wcpLlMYc/wuI/3n4JxClgcU/LecWGPPunJXblIx5d0vMWXrOvDvj5DZV1PpYvzHvbvnKTm8m3rWIkUgvWE21mXfhSEGI9+3VVfiFsnD1q/2leEjZeeaGCO08s61kKz5Y+S4mrHoHC7bPU+/Nn3S7DBcfeynOan+R7Hs1UvajNrCE1bgU7/JIU7zLYUvxLodr1PrYVBXvRu9IJOEu6hb6zHuqxbwHJd6FnQ+VLSNvVLaOFLPu3+R3QJayX3kqH2K3mAkr31GF+9dbZqqXes7RQ3Fxp0txXZ+rsL/ioPJTncoIQr+2qA0sYQGmeJdHmuJdDluKdzlco9bHppp4l+MVwVsNXbyLS+BuM7kQb1gVh1iwWrKvGK3y26k7oYxt2w4dli5Ei/MvwdWV1Sm728zM7z/H3Mkz8M/ql7Bb+Sf2Zb/7lIdxnrIQVRz63WacuD1j3hnz7sRPzNIw5t2aHHeb8epV4eVjzHt4rJ2UFCXxHsSCVSdMmaY2gbiI91RqiCAWrOrF+64u3bDwv7Px5WlD8IQy8759+rSU2iqyqKwQD836Lf6z4t/4lfLvs5wvMKbfLRjR5QpkpGXUuAbFu/u7RNtOlLvNuGNH8U7x7s5jEis1xXtitQfFe2K1R6rWhuLdZ8sGLd57KLvNvDP7S/yp/ym4oeoAjp/zdcqI97eWjMUjs+9WF6Nm122I36T9BpeNHINGeU1qtQLFu3vHpHh3z0zkoHinePfmOYmRi+I9MdpBqwXFe2K1R6rWJjTxLnaZEfu4iz3eYx2JtijAScMHsWBVX85sZdGqeOtq8YED+GvTlji/QUMn1UjYNDO+n4ZxS1/HB6veVet4adercMVxo3By/kDLOjMeU05zRmlgkUPQ3Cpj3uXRZsy7HLbsY+VwjVofy5h3OX5kZzU08W5XkWQ+H7R4Fyye2F2IZ4uLVOEuBHx2Ei5e3bpvM/69dCzGLXsdm/d+j94tT1RFe0G3UUeEyJi1PQcWOXdE1AYWORRrW6V4l0ea4l0OW/axcrhGrY+leJfjR3ZWQxfvVvu8J+tLmgRgGeJ9dVUFfrlzG+ZX7McfmrTAtQ3z7Noyoc4vL1yKn39yLcT/4rimx/W455RHkJvl7Do4sMhpzqgNLHIoUryHxVWUQ/Euhzb7WDlco9bHUrzL8SM7qwkj3pP1JU0yYt7nzpmtvmH1lQ1rsGXGZ5hz9gUYM3cOenc7Hjm5eTU70IjGnTfnK7WN+/Q71fJ3zQn0O0cY43zHvf4ihg4rUO1rh962+Gzr5o3Q6mZ0rMnjx6FvvwFo3eYovLPsDdw34w6cXXkWShvsx83n3Yl++QNq+aJZmVoixrzb3bq1zzPm3T0zkYMx79bcuNuMN58KMxdj3sOkbV9WlMQ7d5ux9wdZKRJGvN/9+EuYNWcRZk54Vta1SrErU7xv3P4D3v9kEp487Qzc+913uKDnCQkt3tt274gJ29/DKwuew4GDB/CbJnfhpK4DcWqfIabsKd5z0bP3iYH5JcW7N5QU7xTv3jwnMXJRvCdGO2i1oHhPrPZI1dqEIt7N9nU3A/rY767HsPOsFzEmYiPIFO+FO7dj0qeT8cjA03HFf7/BaT16o2+T5gk58/6vcc9jTsYcvPfD2zgqtwN+evwt6LSzI/LbtEPnbj0p3g0EnOxL79bfKd7dEjuUnuKd4t2b5yRGLor3xGgHivfEaodUr00o4l0P0SrmPZlBy4h51/O4q2g73ty3B1c2zMWTTVomHKo3Fr+szrav2rUcg9qdiTGKcD/76At81ZPxmL7wWWaO0qyQHILmVrlgVR5txrzLYcs+Vg7XqPWxjHmX40d2VkMX73YVSsbzssX7nP1l+GXRNvxQVYW/KTvPXJSdkxCYtuzbpIr2VxY+h8rqClzdfQzG9LoFnZt0810/Diy+EZoaiNrAIodibasU7/JIU7zLYcs+Vg7XqPWxFO9y/MjOKsW7HSEH52WLd1GFp/YU4s97inBW/WxFwLdCXlqag5rJS/L15hmqcJ+6diLyc9qqs+0/Pf5mZKZnBVIoB5ZAMNYyErWBRQ5FivewuIpyKN7l0GYfK4dr1PpYinc5fmRnNXTxvmjZWhTc/LBlvZLtJU2yY96nfzZFfcPqBx++jymt8/G/uhm4adEC3HDlGJVhPHabKe68X5lt/4eyDeQSnNLmNHW2vXpBac1uM1rjTp82Fa3z2zLm3cTbGfNu1zWFd54x79asudtMeH7otSTGvHslJydflMQ7d5uR40NOrIYu3gcNuxUD+/VE/z7H4ekX3q7ZXeaS0ffirEF9ceuYEU7qnTBpwhLvYjHizg4d8VR1JS6dPw+3XX29+uKmMMX70jXz8eWsqXim7GmUVZUqL1warc64d2vWA/qtIine7d2T4t2eUVgpKN4p3sPyNRnlULzLoOrdJsW7d3bM6ZxA6OJdW7B6TPt83HL3n2vEu9iRRi/mnV9CfFOGKd47de2B+yrL0GXO18BFP8H9jZqFJt5LKvfhtveuQ6vC5ngz7U38pv8D+EXfO2vgU7y780OKd3e8ZKameKd4l+lfsm1TvMsm7M4+xbs7XkztjUDcxLvYElIIeS1MJllf0iSwhxHzrjXvF2WluFNZvFp18CD+pCxePVeJgZd9bC/dhis+GKq+LbVFg5Z48bx/mb50Kch6MB4zSJqHbUVpYJFD0NwqF6zKo82Ydzls2cfK4Rq1PpYx73L8yM5q6OJdhMcc17k9/nDPjdD/nqwvaQpbvIvynlIWrv5ZWcA6pF4DPKUsXm2Rnm7Xzp7PT9vwEf4+9ynM2TIbZ7Y/F7848U70b32qZ3tOM3JgcUrKXbqoDSzu6HhPTfHunZ1dTop3O0LezrOP9cbNLlfU+liKdzuPkHM+dPFuvAwx+64d455/AD27dZRzpRKthjnzLi5jc1WlMvu+HTP2K7PweU1we15TKVcndpQZNflSiJCZfvkD8OZFE5Bdt6GUsoxGObDIwRy1gUUOxdpWKd7lkaZ4l8OWfawcrlHrYyne5fiRndW4i3e7Cib6+bBj3nNy89Q3rGYoMe9CwJ+5cjnOVkJnfjLgDMv4d42hfucIY5zvuNdfxNBhBRD2xSFm2p//4E+oUPZvzz62MZ475zXs2LoFc+fMxoXDC2o1C2Pe3XkqY97d8ZKZmjHv1nS524xMzwvGNmPeg+EYlJUoiXfuNhOU17i3E7p4T7U3rMZLvIvtI3+zaxu2/G8OWmdk4IkzzsPC/85WPaBPv1OPEPJuxbsQ7ldPGoaTKk9E31b98OtLH1JNbN28keLd/T1mmoPiPSCQAZiheKd4D8CN4maC4j1u6E0LpnhPrPZI1dpQvPts2XiK9+IDB/DbL6Zir/J/334DcMaK5b7F+7RtH+Opbx7Fuj2r8ZvWdyv7uA9C/5PPoHjfsgmnDznfp7cczk7xHhhK34Yo3inefTtRHA1QvMcRvknRFO+J1R6pWpvQxXuy7uceywHCjnnX12VKWQnu37UdZYqAf6Rxc/wkO9ezr05a9R88NedRrN61AjeecBvu7HdfaDHuxkozHtNzM8bMGKWBRQ5Bc6uMeZdHmzHvctiyj5XDNWp9LGPe5fiRndXQxbt4w6p+f3e7CibD+XiKd8Hn78VF+MPuQvTOrIdHFQF/QlY919g+XD0eT//3UawoXIbre/0cd/a/HzmZ3h8EXFfAkIEDi1+C5vmjNrDIoVjbKsW7PNIU73LYso+VwzVqfSzFuxw/srMaunjX7y5jVjlt33e7iifS+XiL99KDB5TZ9x0Yt68YIxrkKDPwLdAoPc0xoqlrJ+Kpbx9R9nFfgp8efzPuUGbcG9Vr7Di/jIQcWGRQBaI2sMihSPEeFldRDsW7HNrsY+VwjVofS/Eux4/srIYu3u0qlGzn4xnzLljNm/OViizrhBPx+qzPsamqCn1PUuLflV1oxCEWr2qH2W4zxW3KVOF+9s4zsbdLOX418B40qX9o60nNtmaDC1YZ8251f3qNu02E+50x79atwN1mEsFDY9fB671H8S6nbaMk3rnbjBwfcmI1dPFutdvMs6+8j3cmfYGZE551Uu+ESZMo4l0I7HFffYHP95fg667H4e71G9C1bmZM8b5sy0L8375XsHD7d7iv7n24YEQB2jc/vM8+xfshN/M6OMZyUi5YTZhbGBTvFO+J443ua+K1f6J4d8/aSQ6KdyeUmMYvgYQR7xM+moV7n3gZyRY2k0jiXYjtb8vL8Gj79rhyzWoMrpeNC045zXTm/f0v/oU562bijdLXcVX369BrQ3dcPOKqmn3eOfN++NbyOjhSvPvtnsLJT/FO8R6Op8kpxWv/RPEupz0o3uVwpdUjCSSMeL/78Zcwa86ipJt5FzjjHfOub9J9avz7Tryzbw8uVXaeEQtYc9KOjH/fum8zLnhnILaXbsPQY4erL2DKSMtIqHuDA4uc5ojSwCKHoLlVLliVR5sx73LYso+VwzVqfSxj3uX4kZ3VUMS7NqtuV5nHfnc9hp030C5Zwp1PJPEu4Cys2K8uYP1f+X78Nq8pfpnXpIZZSeU+RbgPUreDHNTuTLx50YSEE+6ishxY5Lh51AYWORRrW6V4l0ea4l0OW/axcrhGrY+leJfjR3ZWQxHv+kqk2htWE23mXWP9Qele3F+0A3WUD8T+7xdn52DL3k14/Ov7MX7l2ziz/bm495RH0bVZdzsfict5DixysEdtYJFDkeI9LK6iHIp3ObTZx8rhGrU+luJdjh/ZWQ1dvNtVKNnOJ1rMu+AnFq+K+PdvlPj3x5T495OUfd/vaZiNBf/3Mh5S/p3Q8iT8rOXP0TytGU4eeKaKfNzrL2LosALGvJs4oNeY0li+zAWriXOnM+bdui2420zi+KlVTbz2TxTvcto2SuKdu83I8SEnVinenVCKkSaRxXv5wYMYd2wnvFdSjF7fv43h32XiybpP4d3hHyFtywGUKPvCU7zbO4DXwZHi3Z5tIqSgeKd4TwQ/9FoHr/0TxbtX4rHzUbzL4UqrRxKIi3gfNOxWFO3ea9oW3G2mL+bOmY0LhxegcOd2TP9sCkYUjManU8ajezg2yAAAIABJREFUU9ce6sy49pkAqN/O0ez3tN4n4o61n2P5tJF46OD9aHdBN5zX8aJa2+Nx5t26a/A6OFK8J0d3S/FO8Z4cnmpeS6/9E8W7nFaneJfDlVbjLN4vGX0vmjbJxavP3JUybZFoC1b1YJftXIzbpv8KS7fORlqXm/HOoEdwSr0GCc+eA4ucJorSwCKHoLlVLliVR5sx73LYso+VwzVqfSxj3uX4kZ3V0GfeuWDVrkmCO7+/qgz3z7wDby0Zi85HD8PKrrcjO7MJprRqh2OVFzgl8sGBRU7rRG1gkUOxtlWKd3mkKd7lsGUfK4dr1PpYinc5fmRnleLdjpCD84k68/7cvGfw2Oz70LtlX/zm1Ccxud7R+LeIc8+qj3sbNUMfZSFroh4cWOS0TNQGFjkUKd7D4irKoXiXQ5t9rByuUetjKd7l+JGd1dDFuwibOWtQX9w6ZoRd3ZLifKIuWH3741fxxYZP8KXy79HTnsalXa+CWBn+34Jr8GHpPlz3/fcYVH0A5552lsqZMe/W7uY1pjSWA3O3mcS5vRnzbt0W3G0mcfzUqiZe+yeKdzltGyXxzt1m5PiQE6uhi3fxwqanX3g7Kd+kagY0EcV7g2MaY+yHf8f3xevR66ST8dv+D6hVFzfaJTffgeHbNqHVyqXoXVmFu8+6EBl16lC8x7hbvA6OFO9OuqD4p6F4p3iPvxd6r4HX/oni3TvzWDkp3uVwpdUjCYQu3kXMe6yDu834222m+mA13it9B5uWrEWXpsfhlmF3oVn95jXi/YZf/BarKyvw+9lfILOkBOn9TsFzTVvhvTde4j7vFo7pdXCkeE+O7pbineI9OTzVvJZe+yeKdzmtTvEuhyutxlm8p2IDJFLM+4vz/4qHZ92Nns174xElXOak1qeYIp+qhM48XVyEZRXl+GlOI9yZ1xR5aWkJ0zwcWOQ0RZQGFjkEza1ywao82ox5l8OWfawcrlHrYxnzLseP7KyGPvNuV6FkPJ8o4v2TdR+qu8vs3r8Ljwx6CiO7XRMTp4h9f3pPIVYoM/HX/yjgcxJEwHNgkXMnRG1gkUOxtlWKd3mkKd7lsGUfK4dr1PpYinc5fmRnNS7iXSxaXb1+s1q3x353PYadNxAinKZ/n25Juf97Ioj3VbuW4/4Zd2Lm95/jthN/i7tOfsiu7dXzk0r24qk9RVhdVYEbcxsrM/BNkF0n/jPwHFgcNZ/rRFEbWFwD8piB4t0jOAfZKN4dQPKQhH2sB2gOskStj6V4d+AUEpKELt71L2kSb1q946bLVfH+7Cvv451JXyTdQtZEWLB64OAB/KfsPWxcvAqdm3TDzZf8Rol5X6O6S59+p9a4jX7nCH2c74TSvVg37nW8PGAgrmjVBncoAr6+IuD1b2wVRrZu3ljz9lejL04ePw59+w1A6zZH1ZyaPm0qWue3ReduPU1d12yHGy1hVekuTJ48EcNGjnLk9k52b1m/dhVWLV+Msy8Y7simlshrTGmsQpzU11UllcTaW3g7dOxkmVXWwCKDkdvr95qeMe/W5LjbjFevCi+f13uP4l1OG8nqY+XU1p9V7jbjj5+f3KGLdzHDPu75B9CzW0foxbvYhebeJ14GF6y6X7A694c5eGjjvbisweU4s/05uHjIFbWEt3ASK/Euzr0y9nm8PfA0LMnMxM/FDHyjplg8Z/YRDwAU7+f7udeOyEvxHhhK34Yo3inefTtRHA1QvMcRvknRFO+J1R6pWpvQxbsQ7M/94fZa4j1eM+8//fUf8e28ZUe0r/EBQh/mc2yHNvhg7GM16eM98/7eJ6/h8w0f45PqT3B/u9+rC1XFbLtx1txOvItZ8LRzL8Qz1RX4vqoKtymz70NWLIcIoNFm7yneKd6tOkKvAiIROlaKd4r3RPBDr3Xweu9x5t0r8dj5KN7lcKXVIwmELt7vfvwlzJqzSA2P0Wbej2mfj4KbH8bF5wzAH+65MdQ2EnUQddEOff3EZ0LcFxYV1wh2fdiPlideMe8llfsw5N8nKfu5b8ANvX+Bhwb+yTe7t/ftUXah2YXNVZX4pSLgxS408YiA58DiuylNDURpYJFD0NwqY97l0WbMuxy27GPlcI1aH8uYdzl+ZGc1dPEuKqSFyOgrd9M1FyfEW1cXLVurPkiYhfZodTe+ZCpe4v2hWb/FP+f/Hd2bHY8pI2chIy3Drr0dnRe70NxS+AOqDh7EDUoIzUONmjnKF2QiDixB0jxsK2oDixyKta1SvMsjTfEuhy37WDlco9bHUrzL8SM7q3ER73aViud5ffiOUciLepl9Fg/xPnXNRNz5xc3K21Ez8OSZz+Gco4cGik0v4K9pmIdHGzdX38Qa1sGBRQ7pqA0scihSvIfFVZRD8S6HNvtYOVyj1sdSvMvxIzuroYt3LcbcGFeeCFtFasJc277SiXgXMe8FV16NmTNmoE27o1Bevh/rVq9CTm4eWrbOR1lpCZYtWYRu3XsiX9mJpXDndjTMyUWF8nIkcWRmZmHf3mLlXDssX7YYvU44EXO+noXhl12JnTu24bOPp6Dg6uswZdL76NqtB3Lz8vDR1A/wYe5UfLpuKu5ucx8GHXUG+p08EHO+maXaNP6uOcHf//wEfnH779Q/58/7r1ruwNOHqH+//srzGKaUmavUWzv+NeNzfFiyD5907oJzshvikcpKLPxmtlo34zH+3bfQ75SBaNP28G4z0z75UL1mce1mh1mZWrrSvYWYPHEiRl51nZ0Pq+dnTZ+mcu3d5yTL9GtXr1QZX3DRCEc2tUSi/bYoO+0MOSe4ByQn9XVVSSWx5iMdj+1smbVeZjrS0+qgZH+VW/Mx08tgFGgFYxgz3gteyk1PT0PDehnYU1LhJXvC5tH3GfGqpBCZu/ZV4qDyTSCP2gS83ns5DeqivOIAKqqqiTVAArL62ACrGJgp0T88+OCDgdmjIecEQhfvIsZ85EWDa4XIxGvBqoZKE+r68B2n4v3qa67FjBnTcdRRHbB//36sWrkCeY3ykJ/fBiUlJVi0cAF6Ht9LOd8e27dvUwVyefkh8Z6VlYXi4j1op5xbrKQ78aT+mDVrOq686lps2/YDpkyehOvG3ID333sHPRQbeYp4/7+3X8ZD+x7EkKPPwi3Nf468rEYYOOg0zJo5Q7Vp/F27xicefwS/u+d+9c//zvlWLXfIWeeofz//j2dx5dXXKPYb1XiPsLdMeRgZe/SxmFtWiquVn5OWLcXPrh1dy8Pe+tfrGDjwdBzVvn3NuQ+VrR7FNYtrNzvMytTSFSoPLhMV8S6u3ckx7bNPVK4n9etvmXyl0i6C8YhLRzoxWZNGtN/GjRsw9MKLXeWLldhJfd0WpvlIZ+Vhy+oQwl18gVJVHawQksHI7fV7TW+8F7zYUbAiI72OIoSC5eqlLkHm0fcZQdp1YyuzbhoqKw8gtci6IRA7rdd7r67ir9XKA9GBA8HVhZagTo7I6GMTka3oHyje49MyoYt3McOuzWzrLzmeW0VqZWtx7vp66bezFJ8b6xn2bjNrylbhf8os/9iM1/Dk4H8gv6ilWl3jDjNedpsZOqxA/cZAOzQbuSechDE7t6Jq6xact2oFRoy4At3rZh3hsdzn3d0NzK0i3fGSmZq7zVjT5T7vMj0vGNvcbSYYjkFZiVLYDPd5D8pr3NsJXbwn2sy7EOPGBah6jIm228yl48/D15tn4Lrjb8Kjpz3jvsU95theXa0I+C2Yp8zEZynTCo82aoErlTAVWQfjMeWQjdLAIoeguVUuWJVHmzHvctiyj5XDNWp9LGPe5fiRndXQxbsIj3nhjYk1u7mICpqFrNhVPIjzWrlmtvTfDsTa513kDWvB6oerx+PGj65CblYevr12ufp/mMeG6kq8Wrwbr+zdrX6FfV1OI4xRfo7OqBt4NTiwBI5UNRi1gUUOxdpWKd7lkaZ4l8OWfawcrlHrYyne5fiRndXQxbuokNlWkWahNHaVT5TzYYj3Lfs24bZPr1dn3e8/9XHcdMKv4nb5ryriXQj49cpe8GfWb4CfNmyMwcr/QR4cWIKkedhW1AYWORQp3sPiKsqheJdDm32sHK5R62Mp3uX4kZ3VuIh3u0ol0/mwYt5feOtPeLdoHI5p1QVnVQ7ByCuuVzHpY9utftd46uNXjXG+4g2rVjHvZm9Y/bysBK/u240vlEWsHTMyMWr2TAw95TS0VnaX0Y7p06aidX5bdO5mvtuMWZla3qrSXZisLHgdNnKUI3dwEkO+fu0qrFq+GGdfMNyRTS2R15jSWIU4qa+rSiqJP50yHp269kCHjp0ss8oaWGQwcnv9XtMz5t2aHGPevXpVePm83nsU73LaSFYfK6e2/qwy5t0fPz+5Kd790FPyhiHeOw7ugfcnvI7ZlV/hxpNvQ8bqgxhRMDqu4l0UvraqQp2BH7t3D66bPQvZyjaXV3XsiqPqHgqjoXi3di6Kd583XoDZKd4p3gN0p9BNUbyHjjxmgRTvidUeqVqbuIh3sWi1aPdeU6bG/d8THXwY4n1e/kKULdyNBm3y8KtBd2P6Z1MSQryLthFbjYkwmq3K3vMfd+qCTso+72NyG2FQvWyK9xjOS/GeOHc2xTvFe+J4o/uaULy7ZyYzB8W7TLq0rREIXbyLxZ9Nm+Ti1WfuSplWkBnz/vHayfjltOvRICMbfzv7ZQxsOzghuX0iwmiUGfiZ+0vQuW6mupD1auXNrF4PfqXrlVzsfFEaWOQQNLfKBavyaDPmXQ5b9rFyuEatj2XMuxw/srMauni32ufdrqKJfF6WeC+rKsUvlUWqH66ZgJ/3uQP3DHgkkTFgZeWhMJo39+1BJuqoAl7MwrdOz3Bdbw4srpE5yhC1gcURlAASUbwHANHCBMW7HLbsY+VwjVofS/Eux4/srFK82xFycF6WeH9zySu464tbcXyLE/CXs15GlybdHNQm/kn+WbwLj+4pRJUSUtMrsx6ebNqi1kud7GrJgcWOkLfzURtYvFFyn4vi3T0zpzko3p2ScpeOfaw7Xk5TR62PpXh36hnBpgtdvIuwmbMG9cWtY0YEeyVxsiYr5r3nkP64f+rt6LSjI1oPOgbtN+erO4mIN6AmUsy7ht34htUFFfvxxscTsaxxEyxu3wHXK7Pwv81rorzgKa2mpbjbTC569j4xMM/lbjPeUDLm3Zobd5vx5lNh5mLMe5i07cuKknjnbjP2/iArReji3e6NprIuVJZdWeJ9RZs1eGvOK7gm81qMvvaX+O/nXyaVeBe8J3/6IVY0aYKxrVqiSHlD6ylZ9VHQMBeXZh96MyvFO8W7rPvSjV2Kd4p3N/6SaGkp3hOrRSjeE6s9UrU2oYt3EfMe6+BuM33x5aypeLb8b6jeV4Wbsm/GdaN+VbOHd7LMvIs21raKXNehI8YpcfBiUWtGnTq4QhHvQsQvH/darb3lNd/gPu/uuxzOvLtnJnJQvFO8e/OcxMhF8Z4Y7aDVguI9sdojVWsTunhPRZBBx7yLOHcR735l99F4cvBzKYFMxL+/rCxmfWZPEUoOHkBuWhruz2uOK3MOzcIbD8Zjymn2KA0scgiaW2XMuzzajHmXw5Z9rByuUetjGfMux4/srFK82xFycD5I8f7h6vG4TdlhplmD5virsjXkyfkDHdQgeZJ8X1WJ+3btwGfKLLw4utfNwq+VWPjzGjQ84iI4sMhp06gNLHIo1rZK8S6PNMW7HLbsY+VwjVofS/Eux4/srMZFvIu493ufePmIuj32u+sx7LzkFKpBivdLx5+HrzfPwC9P+h1+2/8Bu/ZL2vMflu7D47sLsV55S6s4+mTVwz15TXFKvQbq3xxY5DRt1AYWORQp3sPiKsqheJdDm32sHK5R62Mp3uX4kZ3V0MX7s6+8jxfemIhxzz+Ant06qvVbtGwtCm5+GDddc3HS7UIT5ILVWXM/w/PbnsU56efgzusfxf49pTU7y2jxzMkY8965W09TP/zXay+g5OwLMEHZgGZBRbma5pIGObgkOwdDDlZg8uSJGDZylJ0Pq+edvLF0/dpVWLV8Mc6+YLgjm1oirzGlsQpxUl9XlVQSM+bdLbFD6Rnzbs2Nu81486kwc3ntnyje5bRSlMQ7d5uR40NOrIYu3gcNuxUjLxpcS6QLUf/OpC8wc8KzTuqdMGmCEu85zRpj4uxxmFw6CaPyrsON1/wGhTu3p7R413abKWzQABNKivFBaYnyoqdyZCkveLr6QDU6fTMb1xRc56itnYhhindA1sDiVUA4alzJiSjeKd4lu5hU817vPYp3Oc0iq4+VU1t/Vine/fHzkzt08W71hlUtlCaqu80srVyKJUvmYVfTYlxY9yKMuHRUZMS7+DZBHMUHDuCZ4iK8rixsbbx7N4bPn4eN51+En+c2Vl/25Hcmm+Kd4t3MhyjeKd79DKLxzkvxHu8WOLJ8ivfEao9UrU3o4j3VZt6FY/iNea86UIXT/3UC1u9Zgz8PeREju12Tqv7m6Lq2VlXhr2W78c6e3Sg/cFDN069effysYSOcVT9b3W6ShzcCURpYvBHylosLVr1xc5KLMe9OKLlPw5l398yc5IhaH8uYdydeEXya0MV7qsW8ByHeP1o7CWOmXI4Oecdg+lXfISMtI/iWTjKLYmD5urgEE4uL8XHZPiyrOLSw9ZSsBji3QTbOUX7ap9dNsquKf3WjNrCERZziXR5pinc5bCne5XCNWh9L8S7Hj+yshi7eRYW428zhZqmsrlCEewGmbfgID576BG484Ta7NovEef3AslHZXvJjJR5eiPivy8vU6++amYVz6zdUfrKVkJqsSDAJ4iKjNrAEwcyJDYp3J5S8paF498bNLhfFux0hb+ej1sdSvHvzE7+54iLe/VY6kfL7XbC6sng5Pl3+IQ4o7yoalvcT9DtxEObOmY0LhxdELuZd365mb1jdq8TECwGvCflqJUOrjAycUy8bfRbOR9dGTdCz94mW7sGYd8a8mzkHY96te1TuNpNIo415XRjznlhtFCXxzgWr8fM9inef7P2K9w/WvYdNOzagd+d+6FrVBT169aV4V9rETLzrm2pBxX68WLwbHypiXry99bzFi5Cbm4vjju+LC5XtJlsrot54ULxTvFO8u+vwKN7d8YpHaor3eFC3LpPiPbHaI1VrE5p412LdzfZyj3Uu0cH7Ee9r96zGOyvfRJvMdri039Uo3byH4v3HBrcT75pf/E8JoxEz8YXfzsKGeln4puOxyElLxwDlpU8DlPj4gcpLn7pmZqrJKd4p3ine3fWoFO/ueMUjNcV7PKhTvAsCnHmPn++FJt4vGX0vmjbJxavP3GV6tT/99R9RWKTs9T32sfjR8Fiy191m7vz8Fvx76Vjc0ufXuHfAox5LT81sbuMxtyv7wn+1vxSzlZ+vysqwobpSBdMkPQ2nKmE1p2bVV8R8fRxT95CQj+oRpVmhMNuYMe/yaDPmXQ5bt32snFqkntWo9bGMeY+PD4cm3q32d9cuO1n3eRf19yLev94yE9crC1Uz07LwygXj0KdVv/h4QIKW6mdgKT94AB+VKTvVlO7DF8r/5UpYjTiylC0mByhbTg5UZuTPUAS9NiOfoAikVCtqA4sUiCZGKd7lkaZ4l8PWTx8rp0apYTVqfSzFe3z8luI9AO5exPsDM+/EKwuew6ieN+Lx0/8SQC1Sy0RQA0uJEPKKiH+3ZK+6U42Ij9eO1ukZGKTsVnOGIugHK2I+Ny0ttSCaXE3UBpawGpTiXR5pinc5bIPqY+XULnmtRq2PpXiPj6+GJt7Fy5nuuOlyDDtvoOmVipn3p194GzMnPBsfEh5L9RLzvgfFGL90HCqqy3FJt5FomdESrfLbYdXyxYx5/7EdnMa8a832zazPkd0wN+ZuM4tXr8C8ZQvx3amn40tloev2arFfzeGjjxInP1iJkRcz8+J37WVQXmNKY7mUk/q6dclPp4xHp6490KFjJ8ussgYWGYzcXr/X9NxtxpocY969elV4+bzeexTvctpIVh8rp7b+rDLm3R8/P7lDE+93P/4Slq7cYBnTbhcT7+ciZeb1It6/LZyNOd/PRvdmx+OirpeiZF8xxbuhkWSId/2C1f3KjPx3FeWYq8zGzyvfr/6+vbqqphZHZ9RFXyVG/gRlD/l269chY/sPOH3I+YG5EsV7YCh9G6J4p3j37URxNEDxHkf4JkVTvCdWe6RqbUIT7wKgmH0Xh3F2XXxetHsvlnw5Nuk4uxXvazevxIQN72Lf/r24uNOl6NaiJ8V7bl6tdpct3o0FivCar/eX4UtlwesXZaVYX3Xoja7i6L1xA47ftQslA05DT2XBa6/MeuielYXsOt7DbCjeE+dWp3ineE8cb3RfE4p398xk5qB4l0mXtjUCoYp3UaiYgZ/4yewjWqB/n26Wu9AkQ1O5iXl/du6TeOLrBzH0mGF46fy3kuHy4lLHeH+l+52yj/xcMSOv/MxTfhdvedUfDRTh3lkR8p2Un8516yo/Werv7ZUZ+0Q+ojSwhNkOjHmXR5sx73LYxruPlXNV8bcatT6WMe/x8bnQxXt8LlNuqU7F+9Z9m3H91ALM3zYXz57zKkZ0LpBbsSS2nkgDyy4lNn6lMhO/svLQzwrxvxJms0PZnlJ/ZKGOKui7KKE2h0T9oZ8OCSToozawhHULULzLI03xLodtIvWxcq4wPlaj1sdSvMfHzyjeA+DuVLy/uvB53D/jDpxx1Fl4Wdkesn5GgwBKT00TiT6w7DpwQBHx5VhVWan+v0IR86sUgb/DsAi2rrI9ZeeMw0JeE/Xx2m8+agNLWHcHxbs80hTvctgmeh8r56rlW41aH0vxLt+nzEqgePfJ3WnM+9Gdu2DSjgnYt2sPBh17Jo5vcoJacl1llpYLVuMf8x7LDZzGlIptKFcoAn7B/nIsrSpXF8IuV4S9ts+8voyfLF2C7JwcHOzWE20zMiC2rcxXftoqs/Tid22nGzfuyd1m3NA6nJYx79bcuNuMN58KM5fT/slYJ4p3Oa0UJfHO3Wbk+JATqxTvTijFSONUvNdtXR/vbv03ejc8ARd1vwyZBw/FRlO8FyAnARasBiHezWxYCfrBixZid4P6+KbjsaZFt0hPrxHy+Yqgb5WWjtaKyG+Xfkjci9+NB8W7t5uZ4p3i3ZvnJEYuivfEaAetFhTvidUeqVobinefLetUvK+vtwHz93+HYW0uw8lHD0KlEmZB8f4ihg5LbfFuJeg/nvEZ9jVogOIux+EHJXZ+a1UVvq+uVP/fqtuyMpZ7thOz9GLWPk0R9crvzWdMQ/axXdR93rOUBbXNlQcAsSNOE+V/2QOLVwHh8/YLJDvFO8V7II4UJyNe7z3OvMtpMIp3OVxp9UgCFO8BeIRdzPtn66di1OSfoEWDlvhm1DJkpdcLoNTUNhH1geV7ZXebGkGviPkflFh6Ieq1z4oMsfV23iDeHpunzN43U8R+Q+X3JkhDlvJ/C+WzrDpQZvbrKqE6B9WHgHRl4a14KBAPAOIbAB72BBjzbs/IawrGvHslFztf1PtYOVSBKIl3wZAx77I8KbZdivcAuNuJ95s/vhYTV72HO/vdh9v73RNAialvggNL7DYuV/alrxHzitDfJIT9gSqUVB9U3xxbqpwvUmb09yg/xcriWj9HlrLotoUSqpOdVgeN62QgLz0NuYrob6KI/2zlXHPlXJZSgBD8GYrw1x+NlW8FRD79IR4ixMNEKh0U7/Jak+JdDlv2sXK4UrzL4UqrRxKgeA/AI2KJ9683z1Bm3S9Fdt2GeO3C/+D4FocWqvLgrFCYPiBm6sWLqPYqE+lVygz7qpL9qFYW2IoHALHh5ffKefH3FiVspxwH1F1zRHq3M/xer0l7QDDmb6vE+BsP8W2A+NZAf2gPEvrPxF784nP9ka48bIiFwbXK8bmdpxDvuQ0yUFh8+OVeXlkw35EEmjfKQuGeChxQ/JNHcAQaNayLsooDKK84csvb4EqIpqUGWRlIT68DoQtS/RATMV2bZ6f6ZSbk9VG8+2wWu5j3bzbMRP3CTBTl7sbwk65AUeEOZDfMZcy7wn3c69GMeRcul0xvWNVEfLEi5osVUV+kzOSXKDP625WfCvEAoMz4N1yzBnW2bcHCE0854o7adVD5NuDAkaIriG8DfN62R2Q/ee1qNCotw0c9egZpNiVsPTRxPB66eHhKXEuqXoR4A3SHokJM6N0nVS+R15WgBET/8OCDDyZo7VK7WhTvPts3lnjPbdYYn6yYhPb7j0JOu8bo0/lkincdb4r3XPTsfaJPDzycPZl3mxFhQCLcx3hsMrzZVpwXDw3lhlAg8UAhQoX0R4nyYCE+1x/atwv6z9qvXI7qkr345vjevtoiTZnVT7XZ4THvv4NXRoz0xcVv5nQl7Kra8ADo12Yq5e+0YR1a79yJGX1PcnVZwl8PKvcIv89whc02sYJVDR6MgsuK/oHi3dYlpCSgePeJNZZ43113N75ePwO9lX+dunZHfpujKN4p3lUCyTTz7uQW8brjhRPbstNwtxlrwtznXbb3+bfv9d5jzLt/9mYWohTzzn3e5fiQE6sU704o2aSxinn/1Wc34t3lb+KeAY/g533uCKCk6JjgwCKnraM0sMghaG6VC1bl0eaCVTls2cfK4Rq1Ppa7zcjxIzurFO92hBycNxPvy3YuxpUTL1K+OjuAty6ZhO7NjndgiUk0AhxY5PhC1AYWORRrW6V4l0ea4l0OW/axcrhGrY+leJfjR3ZWKd7tCDk4bybeX/juL3jkq3twaZcr8dezX3ZghUn0BDiwyPGHqA0scihSvIfFVZRD8S6HNvtYOVyj1sdSvMvxIzurFO92hGzOm8W8r1u7Ct9XbMSS8kUY0uZclG8uQeduPdE6vy1j3nU8uWCVC1Z93n6BZGfMuzVGxrwH4mJSjTDmXSpe18ajJN4Z8+7aPQLLQPHuE6WZeF+2aiFWl65EeYMKnNNmKDasWkXxbsKZ4p0iupwVAAASDUlEQVTi3eftF0h2ineK90AcKU5GKN7jBN6iWIr3xGqPVK0NxbvPljUT7/OXzVFn3lu2aoNejftAdK6cea8NmuKd4t3n7RdIdop3ivdAHClORije4wSe4h2ceY+f71G8B8BeH/O+Ze8mXDXpEmzYsw7/uuQDnJI/KIASomeC8Zhy2jxKs0JyCJpb5YJVebQZ8y6HLftYOVyj1scy5l2OH9lZpXi3I+TgvF68/3vpWNz5+S045+ih+L+h7zrIzSRmBDiwyPGLqA0scijWtkrxLo80xbsctuxj5XCNWh9L8S7Hj+ysUrzbEXJwXi/ef/bR1Zi8+n08fvpfMKrnjQ5yMwnFe3g+ELWBJSyyFO/ySFO8y2FL8S6Ha9T6WIp3OX5kZ5Xi3Y6QzXl9zHudRumYvPw/6HywM45qfjTatemIsrJSxrwrO+2YHYx5Z8y7z9svkOyMebfGyN1mAnExqUYY8y4Vr2vjURLvjHl37R6BZaB494lSL943YD3mbZ6DEzNPQjtFvLdo2ZriXdkeUyzWpXg/ksA3sz5HdkOKd5+3XyDZKd4p3gNxpDgZoXiPE3iLYineE6s9UrU2FO8+W1YT7zNmfIlZe2egcO8ODKw/CM2btKZ4nzZV3due4r22k1G8+7zxAsxO8U7xHqA7hW6K4j105DELpHhPrPZI1dpQvAfQsiLm/e3F/8ENU6/AKW1Ow1sXf4DM9KwALEfXBOMx5bR9lAYWOQTNrTLmXR5txrzLYcs+Vg7XqPWxjHmX40d2Vine7Qg5OC/E+y1TbsabS17B3ac8jF/0vdNBLiaJRYADixz/iNrAIodibasU7/JIU7zLYcs+Vg7XqPWxFO9y/MjOKsW7HSEH57/btBSXvHsBSitKlL3dJ6Jn894OcjEJxXv4PhC1gSUswhTv8khTvMthS/Euh2vU+liKdzl+ZGeV4t2OkM15EfNe74RsLP9uCRo0y8GQNudi/bpVaJiTx5h3xrxbeg9j3n3eeAFmZ8y7NUzuNhOgo0kyxZh3SWA9mo2SeOduMx6dJIBsFO8OIF4y+l6sXr9ZTXlshzb4YOxjNbmEeF+avxwNttRDt2N64diGnSjef6QzneKd4t3B/RXvJBTvFO/x9kE/5VO8+6EXfF6K9+CZ0mJtAhTvNl7x01//EYVFxTWCXQj5pk1y8eozd6k5hXh/M+1NnJt+Hk7reTYyqtMp3inebfsazrzbIgotAcU7xXtoziahIIp3CVB9mKR49wGPWR0ToHi3QTVo2K2446bLMey8gWrKCR/NwtMvvI2ZE55V/7572t14YtYT6iJVsViVRzAEGI8ZDEejlSgNLHIImltlzLs82ox5l8OWfawcrlHrYxnzLseP7KxSvMcgtGjZWhTc/DDGPf8AenbrqKY0ftb3pb5YvH2xsj3kRHWbSB7BEODAEgxHinc5HI1WKd7lcaZ4l8OWfawcrhTvcrjS6pEEKN59ivf6j9VH71a98fWYr+lbJEACJEACJEACJEACJCCVAMW7T/EuYt4HDxuMdfPXoUOHDti/fz+WL1+ORo0aoU2bNigpKcH8+fPRu3dvtG/fHtu2bUNeXh7Ky8vVkrOysrBnzx713IIFC9C/f39Mnz4do0aNwg8//IAPPvgAP/vZz/D222+jV69eql3tM5FfpBXH6aefbvm7domirg8++KD65zfffKOWe+6556p///Wvf1XLFPa1Q29bfLZ+/fqauhmxvfbaa2odBAPtEPUU1yWu3ewwK1NLp792J3fAxx9/rHI9+eSTLZOLdhGML7/8cicma9KI9tuwYQMuueQSV/liJXZSX7eFaT7StWtXt1l9p5fByHelHBow3gsOs0Uimb7PiMQFJ+FFJvO9l4S4WWUdAfYP8XMHincb9mYx7/c+8TKWfDlWzSmc97KCqzBr5kzkt2mHCkWUc6vIQ1C524y1c3HBavw6PWPJXLBq3RbcKjJx/NSqJlywmlhtFKWwGW4VGT/fo3i3Ye9ktxmKd2Dy+HHo228AWrc5qoYoxTvFe/y6NuclU7xTvDv3lsRLSfGeWG1C8Z5Y7ZGqtaF4d9CysfZ5F9n3llZib1mVA0tM4pQAF1M5JeUuXZQGFndk/KXmglV//GLl5oJVOWzZx8rhGrU+lrvNyPEjO6sU73aEHJyneHcAyWUSDiwugTlMHrWBxSEW38ko3n0jtDRA8S6HLftYOVyj1sdSvMvxIzurFO92hBycp3h3AMllEg4sLoE5TB61gcUhFt/JKN59I6R4l4fQ1DL7WDnAo9bHUrzL8SM7qxTvdoRsznPB6iFAjHl350hcsOqOl8zUjHm3pssFqzI9LxjbjHkPhmNQVqIk3rlgNSivcW+H4t09syNyULxTvHtxIYp3L9Tk5KF4p3iX41nhWKV4D4ez01Io3p2SYjo/BCje/dBT8lK8U7x7cSGKdy/U5OSheKd4l+NZ4VileA+Hs9NSKN6dkmI6PwQo3v3QY14SIAESIAESIAESIAESCJEAxXuIsFkUCZAACZAACZAACZAACfghQPHuhx7zkgAJkAAJkAAJkAAJkECIBCjeQ4TNokiABEiABEiABEiABEjADwGKdx/07N686sN0SmZ1yytW+gkfzcK9T7xci9OSL8emJDs3F+WWs7C9aNlaFNz8MMY9/wB6duvopriUTRskR/prbDdxw/qnv/4jvp237AiDvO8P4QiSI33W2mfdcL778Zcw8ZPZ9NeUHSnic2EU7x65iwGksKgYH4x9rKbTbNokF68+c5dHi6mdzS0vu/RiYHn6hbcxc8KzqQ3O5dXZcTMzN2jYrSjavVc9RfF+iFDQHOmv1o7slrXwV/19L8TRrDmLIt8XBM2RPmvus245C6H/6F1jaiZFnn3lfbwz6YvI+6vLoY3JDQQo3j26hBhA7rjpcgw7b6BqgR1dbJBuedmlJ29z3nbcrFqJM+9HkgmaI/3Vun/wylqzSN89RCJojvTZYPtY+qtHscVspgQo3j04htlgwQHEGqRbXk7Sm32lG/Wvzp1wo3i3v+FlcKS/mnP3w1qzyJlM87A3t2OSkSN9trbPBuGvYuZ+1dpNnHm374qZIgYBincP7hHEDeyh2KTN4paX2/QCjPGrzKSF5aPiXrhxNijYAdqpYKK/HuLux2f1+R/73fU134L6uIWSNmsYHOmz/vxVH54Y9YmmpL3REqjiFO8eGsNvR+mhyKTO4paX2/T6QTzKnaIXbhTv8RHvWltF2V/9ineN4U3XXIxbx4xI6j7Sb+WDuPftONJn/Yl3/TdFL7wxEVG/9/36fNTzU7x79ACz+EKx+wlvSHOgbnm5Ta99xRt1/m65UbwH469uOdJfD3P34rMaPy6wDo8jffYQay/+auxlup8xmpsDeNRezHaIAMW7R09wu+LcYzEpk82Ol1iRLw5t9x679MYdJ0R+7vZjv0uKkbNb0ZkyDmlzIXb+55Yj/dUauFvWXEhpzjJojvTZYDhzd6SojBrhXifFuw/ebvZ69VFMymSNxctMDNmlX71+cw2b/n26cZvOH2nYcdM/JGkzSdpWkeLvJo1yuJhK4RAkR70twZj+emS35pS1Frph1ilGPe5dMAmSI33Weuh1ytnYJprFqH9DnDKiJo4XQvEeR/gsmgRIgARIgARIgARIgATcEKB4d0OLaUmABEiABEiABEiABEggjgQo3uMIn0WTAAmQAAmQAAmQAAmQgBsCFO9uaDEtCZAACZAACZAACZAACcSRAMV7HOGzaBIgARIgARIgARIgARJwQ4Di3Q0tpiUBEiABEiABEiABEiCBOBKgeI8jfBZNAiRAAiRAAiRAAiRAAm4IULy7ocW0JEACJEACJEACJEACJBBHAhTvcYTPokmABEiABEiABEiABEjADQGKdze0mJYESIAESIAESIAESIAE4kiA4j2O8Fk0CZAACZAACZAACZAACbghQPHuhhbTkgAJkAAJkAAJkAAJkEAcCVC8xxE+iyYBEiABEiABEiABEiABNwQo3t3QYloSIAESIAESIAESIAESiCMBivc4wmfRJEACJEACJEACJEACJOCGAMW7G1pMSwIkQAIkQAIkQAIkQAJxJEDxHkf4LJoESIAESIAESIAESIAE3BCgeHdDi2lJgARIgARIgARIgARIII4EKN7jCJ9FkwAJkAAJ/H979/NiZRXGAfz8A4mNQYuCQjTQcOMiIQxcRSsTV20CqYhauGlTJrQI1NrYwkXRDxEEEYIyVxEuAiUoqI2Ei0Ja6EJIg/6CODfO5Z3X+973HGeu+tz53N3Mfeec5/2cZ+A75557hwABAgQItAgI7y1ariVAYCkFTn31Tfrs7MW77u2tV/enw68fTC8cODx57vKFU3ddk59b2bwpfXfm2OS5sbGe3XdoruHK5kcm87z2zsfp59+uzbz22HtvpAMv7U0vHzqa/vzrZipfl4svfH8lHf3oy7Tt6SemdfUHqqlj73O70sUffpr+6P4Xn08n3n+zad6a+1jKpnJTBAgQWJCA8L4gWMMSIBBDoITL859+kHbt2DotOofwS5d/nYbfHHb37N6RTp98d3rNkeOfpyu/XJ2G+tqx+iG7H77z83ms23f+HQzf+ZoS3vt1le/PC+/d1Slhf1Yds55rmbfmPmJ0iioJECDwcAgI7w/HOqiCAIEHJJBDedlRnldCP8RevXY9vfL2h6t2vWvHWs/wvmVl02SHvvzxUerKgX4s/NfUMRTea+cV3h9QY5uWAIGlFRDel3Zp3RgBAjUC+djL9q1PrtpRH/q5HET/uH5jstOed59zgO3uxLeMleeYt+NdE3pzDTufeSrd+vuf9Phjj06OtORXA/Ijf2+R4b123pr7qFkn1xAgQIDA/wLCu04gQGBDC5QAXRDKmfMhlO5Z8d9/PLPqstaxxsJ7zZn3HKL37N45OeOe68n15V34T774euHhvWZeZ9439K+XmydAYAECwvsCUA1JgEBMgXLkpFQ/6zhNCdzlzaxDd9oy1lrOvOfwXt5Emmsprwa07Hjfy5n32nlb6ojZNaomQIDA/RUQ3u+vt9kIEAgikI+f5E9a6e+uzzrrPnZLQ2ON7byPHXspx2ZyeC+fclP+EGgJzWsJ72PzttQx5uh5AgQIEHBsRg8QILCBBXIQP/ftpcnOdf9RQmn/U2iGwvu9jLWe4T3Xn8/cl4+zbAnNawnvY/O21LGBW9GtEyBAoFrAzns1lQsJEFg2ge7Rlu4Oe/cTW7pvSM33Py+850+fyY/asdY7vHfXpyU0rzW8z5u3pY5l6y/3Q4AAgUUICO+LUDUmAQKhBGb9w6KhM+1jx2ZaxhoL77VvWJ31ykFLaB6qoxz3KYvZ/SdN5cx7f6H783rDaqhfBcUSIBBAQHgPsEhKJECAAAECBAgQIJAFhHd9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIgPAeZKGUSYAAAQIECBAgQEB41wMECBAgQIAAAQIEgggI70EWSpkECBAgQIAAAQIEhHc9QIAAAQIECBAgQCCIwH9xS0N3VRqLJQAAAABJRU5ErkJggg==",
"text/html": [
""
]
},
"metadata": {},
"output_type": "display_data"
}
],
"source": [
"dynamics.plot_history(colors=[\"darkturquoise\", \"green\"], title=\"Single reaction A <-> B (no downregulation)\",\n",
" show_intervals=True)"
]
},
{
"cell_type": "markdown",
"id": "ce63f6dc-f9fc-4875-82aa-bbd05488ecb5",
"metadata": {},
"source": [
"#### Notice the intersection at the exact midpoint of the 2 initial concentrations (50 and 0):"
]
},
{
"cell_type": "code",
"execution_count": 10,
"id": "477e29f3-17ee-4037-920a-b3e33920f04a",
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(0.024381560069949713, 25.0)"
]
},
"execution_count": 10,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"dynamics.curve_intersect('A', 'B', t_start=0, t_end=0.1)"
]
},
{
"cell_type": "code",
"execution_count": 11,
"id": "58b4b5e5-ebd0-4bb6-8059-6a460d555106",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"0: A <-> B\n",
"Final concentrations: [A] = 7.124 ; [B] = 42.88\n",
"1. Ratio of reactant/product concentrations, adjusted for reaction orders: 6.01865\n",
" Formula used: [B] / [A]\n",
"2. Ratio of forward/reverse reaction rates: 6\n",
"Discrepancy between the two values: 0.3108 %\n",
"Reaction IS in equilibrium (within 1% tolerance)\n",
"\n"
]
},
{
"data": {
"text/plain": [
"True"
]
},
"execution_count": 11,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"# Verify that all the reactions have reached equilibrium\n",
"dynamics.is_in_equilibrium()"
]
},
{
"cell_type": "code",
"execution_count": null,
"id": "00363317-fe95-4d18-acab-5e6116e01a17",
"metadata": {},
"outputs": [],
"source": []
},
{
"cell_type": "markdown",
"id": "3f67c102-0d90-4624-98f3-d6f50719d8c1",
"metadata": {},
"source": [
"# Scenario 2 - downregulated by shunt: \n",
"### kinetically fast, \n",
"### but with thermodynamical dis-advantage (i.e. energetically un-favored)"
]
},
{
"cell_type": "code",
"execution_count": 12,
"id": "589fe009-4bf5-454e-ac37-9abd6ba7ffeb",
"metadata": {
"tags": []
},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"Number of reactions: 2 (at temp. 25 C)\n",
"0: A <-> B (kF = 30 / kR = 5 / delta_G = -4,441.7 / K = 6) | 1st order in all reactants & products\n",
"1: A <-> S (kF = 150 / kR = 100 / delta_G = -1,005.1 / K = 1.5) | 1st order in all reactants & products\n",
"Set of chemicals involved in the above reactions: {'S', 'A', 'B'}\n"
]
}
],
"source": [
"# Register the new chemical (\"S\")\n",
"chem_data.add_chemical(\"S\")\n",
"\n",
"# Add the reaction A <-> S (fast shunt, poor thermodynical energetic advantage)\n",
"chem_data.add_reaction(reactants=[\"A\"], products=[\"S\"],\n",
" forward_rate=150., reverse_rate=100.) \n",
"\n",
"chem_data.describe_reactions()"
]
},
{
"cell_type": "code",
"execution_count": 13,
"id": "cb582868-431c-4022-aa0e-a2f554f80d6c",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"[GRAPHIC ELEMENT SENT TO LOG FILE `down_regulate_1.log.htm`]\n"
]
}
],
"source": [
"# Send a plot of the network of reactions to the HTML log file\n",
"chem_data.plot_reaction_network(\"vue_cytoscape_2\")"
]
},
{
"cell_type": "code",
"execution_count": 14,
"id": "ae304704-c8d9-4cef-9e0b-2587bb3909ef",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"SYSTEM STATE at Time t = 0:\n",
"3 species:\n",
" Species 0 (A). Conc: 50.0\n",
" Species 1 (B). Conc: 0.0\n",
" Species 2 (S). Conc: 0.0\n",
"Set of chemicals involved in reactions: {'S', 'A', 'B'}\n"
]
}
],
"source": [
"dynamics = UniformCompartment(chem_data=chem_data, preset=\"mid\") # Notice we're over-writing the earlier \"dynamics\" object\n",
"dynamics.set_conc(conc={\"A\": 50.}, snapshot=True)\n",
"dynamics.describe_state()"
]
},
{
"cell_type": "markdown",
"id": "fc516ca2-e62d-4784-b826-5372ff7f4c75",
"metadata": {
"tags": []
},
"source": [
"### Run the reaction"
]
},
{
"cell_type": "code",
"execution_count": 15,
"id": "2502cd11-0df9-4303-8895-98401a1df7b8",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"57 total step(s) taken\n",
"Number of step re-do's because of negative concentrations: 0\n",
"Number of step re-do's because of elective soft aborts: 2\n",
"Norm usage: {'norm_A': 38, 'norm_B': 37, 'norm_C': 36, 'norm_D': 36}\n"
]
}
],
"source": [
"dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()\n",
"\n",
"# The changes of concentrations vary very rapidly early on; automated variable timesteps will take care of that\n",
"dynamics.single_compartment_react(initial_step=0.001, duration=0.3,\n",
" snapshots={\"initial_caption\": \"1st reaction step\",\n",
" \"final_caption\": \"last reaction step\"},\n",
" variable_steps=True)"
]
},
{
"cell_type": "code",
"execution_count": 16,
"id": "a41e5009-e2b3-4bea-9892-7b9bd8295a07",
"metadata": {},
"outputs": [
{
"data": {
"application/vnd.plotly.v1+json": {
"config": {
"plotlyServerURL": "https://plot.ly"
},
"data": [
{
"hovertemplate": "Chemical=A
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "A",
"line": {
"color": "darkturquoise",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "A",
"orientation": "v",
"showlegend": true,
"type": "scatter",
"x": [
0,
0.00016,
0.000352,
0.000448,
0.000544,
0.0006399999999999999,
0.0007359999999999999,
0.0008319999999999998,
0.0009279999999999998,
0.0010239999999999997,
0.0011199999999999997,
0.0012159999999999996,
0.0013311999999999996,
0.0014463999999999996,
0.0015846399999999996,
0.0017228799999999996,
0.0018887679999999994,
0.0020546559999999993,
0.002253721599999999,
0.002452787199999999,
0.0026916659199999987,
0.0029783203839999985,
0.0032649748479999983,
0.003608960204799998,
0.004021742632959998,
0.004434525061119998,
0.004929863974911998,
0.005524270671462397,
0.006118677368012797,
0.006831965403873277,
0.007687911046905852,
0.008715045818544943,
0.009742180590184033,
0.010974742316150941,
0.012453816387311231,
0.01422870527270358,
0.016003594158095928,
0.018133460820566747,
0.02068930081553173,
0.023245140810496712,
0.02631214880445469,
0.029992558397204265,
0.03367296798995384,
0.03808945950125332,
0.04338924931481271,
0.04974899709108397,
0.05610874486735523,
0.06374044219888074,
0.07289847899671136,
0.0838881231541081,
0.09707569614298418,
0.11290078372963548,
0.13189088883361705,
0.15467901495839492,
0.18202476630812836,
0.2148396679278085,
0.25421754987142464,
0.30147100820376405
],
"xaxis": "x",
"y": [
50,
48.56,
46.905036800000005,
46.11949179699201,
45.35383252878493,
44.607545689901286,
43.880131240017015,
43.17110206113199,
42.47998362360085,
41.806313660795034,
41.14964185217302,
40.509529514541434,
39.760753258646176,
39.03460778554155,
38.18954921943425,
37.37509202354907,
36.43310062025945,
35.531961527923315,
34.49743189242318,
33.51660150109652,
32.40059733325611,
31.144523830183388,
29.981806308284227,
28.68994661258535,
27.277150107971387,
26.01404298592347,
24.658037510153605,
23.23535470465236,
22.025489070832347,
20.78833150543161,
19.563652381053977,
18.39684605185298,
17.510259052230232,
16.690460412531404,
15.962184189736089,
15.336059619437233,
14.89170237056441,
14.476318939971941,
14.07199615444411,
13.723599199808127,
13.337219176461387,
12.902672816605815,
12.494767570609763,
12.033736604354653,
11.519072453465899,
10.953147538913296,
10.44404133929037,
9.894450801787363,
9.314399538695683,
8.71897146542408,
8.128419312766091,
7.567292117856376,
7.0621616790657855,
6.637722086682146,
6.311648162271255,
6.089109655617535,
5.961954653046801,
5.8867319831351415
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=B
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "B",
"line": {
"color": "green",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "B",
"orientation": "v",
"showlegend": true,
"type": "scatter",
"x": [
0,
0.00016,
0.000352,
0.000448,
0.000544,
0.0006399999999999999,
0.0007359999999999999,
0.0008319999999999998,
0.0009279999999999998,
0.0010239999999999997,
0.0011199999999999997,
0.0012159999999999996,
0.0013311999999999996,
0.0014463999999999996,
0.0015846399999999996,
0.0017228799999999996,
0.0018887679999999994,
0.0020546559999999993,
0.002253721599999999,
0.002452787199999999,
0.0026916659199999987,
0.0029783203839999985,
0.0032649748479999983,
0.003608960204799998,
0.004021742632959998,
0.004434525061119998,
0.004929863974911998,
0.005524270671462397,
0.006118677368012797,
0.006831965403873277,
0.007687911046905852,
0.008715045818544943,
0.009742180590184033,
0.010974742316150941,
0.012453816387311231,
0.01422870527270358,
0.016003594158095928,
0.018133460820566747,
0.02068930081553173,
0.023245140810496712,
0.02631214880445469,
0.029992558397204265,
0.03367296798995384,
0.03808945950125332,
0.04338924931481271,
0.04974899709108397,
0.05610874486735523,
0.06374044219888074,
0.07289847899671136,
0.0838881231541081,
0.09707569614298418,
0.11290078372963548,
0.13189088883361705,
0.15467901495839492,
0.18202476630812836,
0.2148396679278085,
0.25421754987142464,
0.30147100820376405
],
"xaxis": "x",
"y": [
0,
0.24000000000000002,
0.5194752,
0.654312357888,
0.7868224243315508,
0.9170637872507723,
1.0450933282198076,
1.1709664613935111,
1.2947371714281024,
1.4164580504217874,
1.5361803339006745,
1.6539539358746604,
1.7930021924098518,
1.929382586408905,
2.089933322573377,
2.246868459183652,
2.4310071945769423,
2.6103052458402805,
2.8199028731625377,
3.0231137042193295,
3.259695002518898,
3.5336562477554763,
3.7964230596194666,
4.099292560129741,
4.44611315524191,
4.774722615967033,
5.149470120275348,
5.573872801254246,
5.971644577655393,
6.421662599643144,
6.927990082366925,
7.4952454134362885,
8.02363348555623,
8.621659621455152,
9.298492072429907,
10.065906220427024,
10.793170947994442,
11.629751085867117,
12.591106818882864,
13.509175664355627,
14.564723569134426,
15.769295709366046,
16.903721997558126,
18.18593732133796,
19.61731733475414,
21.191263245892706,
22.607185470483728,
24.135702357923897,
25.748936442359923,
27.404936291797696,
29.047375481426027,
30.607977602612998,
32.01284422426204,
33.193283496969016,
34.10021204335371,
34.718719931378274,
35.07625889274106,
35.24057547583993
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=S
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "S",
"line": {
"color": "red",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "S",
"orientation": "v",
"showlegend": true,
"type": "scatter",
"x": [
0,
0.00016,
0.000352,
0.000448,
0.000544,
0.0006399999999999999,
0.0007359999999999999,
0.0008319999999999998,
0.0009279999999999998,
0.0010239999999999997,
0.0011199999999999997,
0.0012159999999999996,
0.0013311999999999996,
0.0014463999999999996,
0.0015846399999999996,
0.0017228799999999996,
0.0018887679999999994,
0.0020546559999999993,
0.002253721599999999,
0.002452787199999999,
0.0026916659199999987,
0.0029783203839999985,
0.0032649748479999983,
0.003608960204799998,
0.004021742632959998,
0.004434525061119998,
0.004929863974911998,
0.005524270671462397,
0.006118677368012797,
0.006831965403873277,
0.007687911046905852,
0.008715045818544943,
0.009742180590184033,
0.010974742316150941,
0.012453816387311231,
0.01422870527270358,
0.016003594158095928,
0.018133460820566747,
0.02068930081553173,
0.023245140810496712,
0.02631214880445469,
0.029992558397204265,
0.03367296798995384,
0.03808945950125332,
0.04338924931481271,
0.04974899709108397,
0.05610874486735523,
0.06374044219888074,
0.07289847899671136,
0.0838881231541081,
0.09707569614298418,
0.11290078372963548,
0.13189088883361705,
0.15467901495839492,
0.18202476630812836,
0.2148396679278085,
0.25421754987142464,
0.30147100820376405
],
"xaxis": "x",
"y": [
0,
1.2000000000000002,
2.575488,
3.22619584512,
3.859345046883533,
4.475390522847954,
5.074775431763192,
5.65793147747451,
6.225279204971056,
6.777228288783186,
7.314177813926316,
7.836516549583915,
8.446244548943984,
9.036009628049555,
9.720517457992388,
10.37803951726729,
11.135892185163621,
11.85773322623642,
12.682665234414296,
13.460284794684167,
14.339707664225008,
15.32181992206115,
16.221770632096323,
17.21076082728493,
18.276736736786724,
19.211234398109518,
20.19249236957107,
21.19077249409342,
22.002866351512285,
22.790005894925272,
23.508357536579123,
24.10790853471076,
24.466107462213564,
24.68787996601347,
24.73932373783403,
24.598034160135768,
24.31512668144117,
23.893929974160965,
23.33689702667305,
22.767225135836267,
22.09805725440421,
21.328031474028162,
20.601510431832136,
19.78032607430741,
18.86361021177998,
17.855589215194016,
16.94877319022592,
15.96984684028876,
14.93666401894441,
13.876092242778242,
12.8242052058079,
11.824730279530645,
10.924994096672194,
10.168994416348857,
9.58813979437506,
9.192170413004213,
8.961786454212165,
8.87269254102495
],
"yaxis": "y"
}
],
"layout": {
"autosize": true,
"legend": {
"title": {
"text": "Chemical"
},
"tracegroupgap": 0
},
"shapes": [
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0,
"x1": 0,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.00016,
"x1": 0.00016,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.000352,
"x1": 0.000352,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.000448,
"x1": 0.000448,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.000544,
"x1": 0.000544,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0006399999999999999,
"x1": 0.0006399999999999999,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0007359999999999999,
"x1": 0.0007359999999999999,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0008319999999999998,
"x1": 0.0008319999999999998,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0009279999999999998,
"x1": 0.0009279999999999998,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0010239999999999997,
"x1": 0.0010239999999999997,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0011199999999999997,
"x1": 0.0011199999999999997,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0012159999999999996,
"x1": 0.0012159999999999996,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0013311999999999996,
"x1": 0.0013311999999999996,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0014463999999999996,
"x1": 0.0014463999999999996,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0015846399999999996,
"x1": 0.0015846399999999996,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0017228799999999996,
"x1": 0.0017228799999999996,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0018887679999999994,
"x1": 0.0018887679999999994,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0020546559999999993,
"x1": 0.0020546559999999993,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.002253721599999999,
"x1": 0.002253721599999999,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.002452787199999999,
"x1": 0.002452787199999999,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0026916659199999987,
"x1": 0.0026916659199999987,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0029783203839999985,
"x1": 0.0029783203839999985,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0032649748479999983,
"x1": 0.0032649748479999983,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.003608960204799998,
"x1": 0.003608960204799998,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.004021742632959998,
"x1": 0.004021742632959998,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.004434525061119998,
"x1": 0.004434525061119998,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.004929863974911998,
"x1": 0.004929863974911998,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.005524270671462397,
"x1": 0.005524270671462397,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.006118677368012797,
"x1": 0.006118677368012797,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.006831965403873277,
"x1": 0.006831965403873277,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.007687911046905852,
"x1": 0.007687911046905852,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.008715045818544943,
"x1": 0.008715045818544943,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.009742180590184033,
"x1": 0.009742180590184033,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.010974742316150941,
"x1": 0.010974742316150941,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.012453816387311231,
"x1": 0.012453816387311231,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.01422870527270358,
"x1": 0.01422870527270358,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.016003594158095928,
"x1": 0.016003594158095928,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.018133460820566747,
"x1": 0.018133460820566747,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.02068930081553173,
"x1": 0.02068930081553173,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.023245140810496712,
"x1": 0.023245140810496712,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.02631214880445469,
"x1": 0.02631214880445469,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.029992558397204265,
"x1": 0.029992558397204265,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.03367296798995384,
"x1": 0.03367296798995384,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.03808945950125332,
"x1": 0.03808945950125332,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.04338924931481271,
"x1": 0.04338924931481271,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.04974899709108397,
"x1": 0.04974899709108397,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.05610874486735523,
"x1": 0.05610874486735523,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.06374044219888074,
"x1": 0.06374044219888074,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.07289847899671136,
"x1": 0.07289847899671136,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.0838881231541081,
"x1": 0.0838881231541081,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.09707569614298418,
"x1": 0.09707569614298418,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.11290078372963548,
"x1": 0.11290078372963548,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.13189088883361705,
"x1": 0.13189088883361705,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.15467901495839492,
"x1": 0.15467901495839492,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.18202476630812836,
"x1": 0.18202476630812836,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.2148396679278085,
"x1": 0.2148396679278085,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.25421754987142464,
"x1": 0.25421754987142464,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
},
{
"line": {
"color": "gray",
"dash": "dot",
"width": 1
},
"type": "line",
"x0": 0.30147100820376405,
"x1": 0.30147100820376405,
"xref": "x",
"y0": 0,
"y1": 1,
"yref": "y domain"
}
],
"template": {
"data": {
"bar": [
{
"error_x": {
"color": "#2a3f5f"
},
"error_y": {
"color": "#2a3f5f"
},
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "bar"
}
],
"barpolar": [
{
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "barpolar"
}
],
"carpet": [
{
"aaxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"baxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"type": "carpet"
}
],
"choropleth": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "choropleth"
}
],
"contour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "contour"
}
],
"contourcarpet": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "contourcarpet"
}
],
"heatmap": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmap"
}
],
"heatmapgl": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmapgl"
}
],
"histogram": [
{
"marker": {
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "histogram"
}
],
"histogram2d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2d"
}
],
"histogram2dcontour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2dcontour"
}
],
"mesh3d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "mesh3d"
}
],
"parcoords": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "parcoords"
}
],
"pie": [
{
"automargin": true,
"type": "pie"
}
],
"scatter": [
{
"fillpattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
},
"type": "scatter"
}
],
"scatter3d": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatter3d"
}
],
"scattercarpet": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattercarpet"
}
],
"scattergeo": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergeo"
}
],
"scattergl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergl"
}
],
"scattermapbox": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattermapbox"
}
],
"scatterpolar": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolar"
}
],
"scatterpolargl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolargl"
}
],
"scatterternary": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterternary"
}
],
"surface": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "surface"
}
],
"table": [
{
"cells": {
"fill": {
"color": "#EBF0F8"
},
"line": {
"color": "white"
}
},
"header": {
"fill": {
"color": "#C8D4E3"
},
"line": {
"color": "white"
}
},
"type": "table"
}
]
},
"layout": {
"annotationdefaults": {
"arrowcolor": "#2a3f5f",
"arrowhead": 0,
"arrowwidth": 1
},
"autotypenumbers": "strict",
"coloraxis": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"colorscale": {
"diverging": [
[
0,
"#8e0152"
],
[
0.1,
"#c51b7d"
],
[
0.2,
"#de77ae"
],
[
0.3,
"#f1b6da"
],
[
0.4,
"#fde0ef"
],
[
0.5,
"#f7f7f7"
],
[
0.6,
"#e6f5d0"
],
[
0.7,
"#b8e186"
],
[
0.8,
"#7fbc41"
],
[
0.9,
"#4d9221"
],
[
1,
"#276419"
]
],
"sequential": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"sequentialminus": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
]
},
"colorway": [
"#636efa",
"#EF553B",
"#00cc96",
"#ab63fa",
"#FFA15A",
"#19d3f3",
"#FF6692",
"#B6E880",
"#FF97FF",
"#FECB52"
],
"font": {
"color": "#2a3f5f"
},
"geo": {
"bgcolor": "white",
"lakecolor": "white",
"landcolor": "#E5ECF6",
"showlakes": true,
"showland": true,
"subunitcolor": "white"
},
"hoverlabel": {
"align": "left"
},
"hovermode": "closest",
"mapbox": {
"style": "light"
},
"paper_bgcolor": "white",
"plot_bgcolor": "#E5ECF6",
"polar": {
"angularaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"radialaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"scene": {
"xaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"yaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"zaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
}
},
"shapedefaults": {
"line": {
"color": "#2a3f5f"
}
},
"ternary": {
"aaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"baxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"caxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"title": {
"x": 0.05
},
"xaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
},
"yaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
}
}
},
"title": {
"text": "Coupled reactions A <-> B and A <-> S (fast but disadvantaged energetically) (time steps shown in dashed lines)"
},
"xaxis": {
"anchor": "y",
"autorange": true,
"domain": [
0,
1
],
"range": [
-0.00020877493642919949,
0.3016797831401932
],
"title": {
"text": "SYSTEM TIME"
},
"type": "linear"
},
"yaxis": {
"anchor": "x",
"autorange": true,
"domain": [
0,
1
],
"range": [
-2.7777777777777777,
52.77777777777778
],
"title": {
"text": "Concentration"
},
"type": "linear"
}
}
},
"image/png": "iVBORw0KGgoAAAANSUhEUgAAAu8AAAFoCAYAAADn3hWsAAAgAElEQVR4XuydCXwV1dn/n+RmD0kgrAk7AoKyKYioIAKiYl2Q14XWWuhbtdrF1vb9t1rb2vatrfVtq6+2lWrti3QRl6pFixugCFUEZAvIDmELEPaEkD35n2eSuZlMZu6cmXvmzr3hN3zu55J7zzznnN85M/c7zzznmaRGsRE2KAAFoAAUgAJQAApAASgABeJegSTAe9yPERoIBaAAFIACUAAKQAEoAAU0BQDvmAhQAApAASgABaAAFIACUCBBFAC8J8hAoZlQAApAASgABaAAFIACUADwjjkABaAAFIACUAAKQAEoAAUSRAHAe4IMFJoJBaAAFIACUAAKQAEoAAUA75gDUAAKQAEoAAWgABSAAlAgQRQAvCfIQKGZUAAKQAEoAAWgABSAAlAA8I45AAWgABSAAlAACkABKAAFEkQBwHuCDBSaCQWgABSAAlAACkABKAAFAO+YA1AACkABKAAFoAAUgAJQIEEUALwnyEChmVAACkABKAAFoAAUgAJQAPCOOQAFoAAUgAJQAApAASgABRJEAcB7ggwUmgkFoAAUgAJQAApAASgABQDvmANQAApAASgABaAAFIACUCBBFAC8J8hAoZlQAApAASgABaAAFIACUADwjjkABaAAFIACUAAKQAEoAAUSRAHAe4IMFJoJBaAAFIACUAAKQAEoAAUA75gDUAAKQAEoAAWgABSAAlAgQRQAvCfIQKGZUAAKQAEoAAWgABSAAlAA8I45AAWgABSAAlAACkABKAAFEkQBwHuCDBSaCQWgABSAAlAACkABKAAFAO+YA1AACkABKAAFoAAUgAJQIEEUALwnyEChmVAACkABKAAFoAAUgAJQAPCOOQAFoAAUgAJQAApAASgABRJEAcB7ggwUmgkFoAAUgAJQAApAASgABQDvmANQAApAASgABaAAFIACUCBBFAC8J8hAoZlQAApAASgABaAAFIACUADwjjkABaAAFIACUAAKQAEoAAUSRAHAe4IMFJoJBaAAFIACUAAKQAEoAAUA75gDUAAKQAEoAAWgABSAAlAgQRQAvCfIQKGZUAAKQAEoAAWgABSAAlAA8I45AAWgABSAAlAACkABKAAFEkQBwHuCDBSaCQWgABSAAlAACkABKAAFAO+YA1AACkABKAAFoAAUgAJQIEEUALwnyEChmVAACkABKAAFoAAUgAJQAPCOOQAFoAAUgAJQAApAASgABRJEAcC7ooGa9/I79Kvfv0CP//TrdNXEi5RY/a+fPU1vLfmENn0wV4k9GIEC7UEBHBexGUVd52mTL6Zf//jeVpWef8Xs8N9W38emhS21TLn1O9SjWz797Xc/jHXVUvXd/o2f06HS47T4pd9Klfer0P88PZ/mvvg2vfTMT+j8wf38qqZd2oV28TWsVuPh9Tyg8xufQ7weo3yMr9u4g2bfdg39v3tn+i5WQsD7u0tX0f0P/76NGN//+ufpS7dc7btIMhUA3lurFAmw+Ic/moNEZjxUldHHle2pvDBT1T4VdowgZrYXjxeOXuFd72c8wKaKcbOysWlbMd1690/afOW2z/q8t/oh4h/IC4YNagP0qvrkBZK8/mhHarOXdtjZM8O7/pvmdlyi1Vhln6Jtiz5XYwU70bY3nrSLti8q97c79iKdQ1TUrxLeuT36fPRyTOptMTOp3W+VXteoYQM9OxziHt55YrDHwiyKPjGi6byKCaTbALy3T3jXr6a5d14Oahk4cPvjpfqkyFBrdRzpx168AbwXeDdehGknaoV3s/QTsZcLUpUXsvoPiHk+eTlX2v0g69Dp54WsF0gCvMv9mnnRVs6y+1KAd/eaxeMe7QXeWVv9+HB7fuPzuBUfnLXwrnc80i0+hqt4uFUKeJeH93g8AUXyYjIMrdu0Q7slpgL6jCDp9e6R0YZb+Df31Q7e/ZjTKsbeC7zrns9Z4k4dh7d51d1KO/4sGieCfkfAC/zr7ZHxGrFu5vAXq/GINO76d36GXXgBzESDdxXHgRcbXrT1Uo/MPoB3GZXiv4wfx55Mr1V73vU63fYn0jHl5bdKpu9cJm497zI/Rlad1MXSv7P6UbWLP7QSWr+iGjF0gPajr2/mqyy7Hzyr29hWV3VW5fjHnO86yACj3s6rJo4JhxgZYUCmHfokNOtqdxVq9Err+3DZd5eu1mL17ezYHRxme1ZXsvq+DGHGUCqrcbYKt3IDusaD8tP1W6OGPhXQbtZUBcTbwbsbL4TZs6230+6OGc+T58U6Eb4gMs4d83qRaI8Ltm2GBLv+yp40uZwO3NFAu7m+aCBeJQjZnR/tzg98fnJz7rAqq59j7Ow4XWzZnRfM5xD9nGC2Z9bPazt4TK3OO3wu5k2Pp7UbL/PvF+9jvFByo7NxnupzTf9N0W3q51yr3xiz88wqvM7ujh3XE+kcbRfepduTPZ/o/TL/drAdPrdYzRtzP+zmlrmcWTun84XVPDBf9OrH2pM/v69VuJvdhbyVLuax04+FH9z3xVY29XJW2ut6mW051affnTVrwXbsjjUua7Wfro3VMWA+DvhvGXiPdK7nNhiPSb0Pbi9w7XjGrh/6uc7srTfq9Y4IFbf6bTTqFrfwrk8ap5O2cdJYDYbVZ27hnesw/gjoIhtB0ArercIbrPplZY/rjHRitfvhtzqZyraDJ+3hIydaeefsAM7qoOCyvPFijUhXnFaT3cqe1Wf65DX20+qH0EpnNzCqn2CMi+C8Qp8f0G4e/2jqsOuXm/5y/XzCMd4FsxoDuzUEVvNFxXFhdZKXuaNn98PsB7S7OZYjAYPetmi94mzH7iLXzkkhe+6w0t58nnP742mEAavzgtW52wne7eDACdjsfrfMv0NW5yyr8715P1md7ZxfZv0jAZZ5HvDf5rkVzTk60gWn7PlEH3szhFnFIFutM7A6x8hqF2kuWNVv9fujj7kZ1q3CMPSxMzrTrI4n/TfS6gLA7pxqFSLppj6rxeJWcyuStuwg5TWMXG/3rp1aLfq06qcMvNv95ke6uxjpmGhzkdK8xsiOU53W/lmdm7gOqzE2fx638O4WtJw838YfI7fwLhPLZFV/pDiotRu3h70wTldubjzvVrfFZdsRCViM+sncCnID73Y/1lYHUSSvvTGbg139bJM3p4xAVnW7hT6jh8PNRagTIET63g6OZaDPqoybOxVW+5sXN9odp27GWmb+GdtinjN2P2CRNNJ/ZFV62p3G2e0CWxUL+/U5azdf3YZSmQHQ7vjlc4CeocErvFsBhNmWrOfdK7zbnWvNvzlW4Gq3r1Eb2XO0090TI4RbObhkw6N0fY2/UbLnaC93i8znExW/HeY+uNHOajwi9Ytt86Y7OWRZxMmLbVxAHinsQ5Y1VNRnZcNr1iWr85IMvLPWVseVeRysHCgyv31Oa4C8wHuku9VGbmk38O4GFmUPGLuB58/NP2LmvyNNfuPJItKB7gZS7E78su3QJ69MtgqZmDAV42Glv+wPg66x1zhiN55gux9Ur/BudXtadgGNV3i3glIvK+Kdbq3Lwruq48Ju/tvdMrUbSy/wbhVWJvODoLfBLbybL1j4Yta4ydTt9GMUCd5lzh26JpEWf6uEd/P4+wnvkc61MvCuez+dxklG50gXAuZUkVZj6gR55uPEfDFgdSElo4EVRJk/M56r7H7LzWPh5OXnkFi9D260szpfRDpG+HfF6LiTZZFIx4QZRO3Gzs05VUV9VseDnbZmHe0cEcZjQxbezb/lTg4KnTucjkMjB9rd7VQB73bnlbiHd1mPZaQrOvMPtewBEw2828XsGScpD/iBg0e02G2rfqqAd9l2cM5f/YfVPGnNB5zMAegG3iNdDJhvy8rCu9FzZtRcNmOMFYTqdtxeEEQT0mIHk+bPo6kjUniM7B0wfbzN+prHSxbeI4GQm+PCLu5Q10/2osgM1H564P0IzdHB0Cmkxiu8y547WEerixrj+S9R4T0StMmCqzGm1WqOyugcCdLstDWe063mgBGmjF52WYDSx914d1QGJp3OJ3a/B3YXbZHOp3wu6FnQVYsTtwI32Xmpl4tUl66hLIs4nceMv0mymhjbZz6n+lGf7J0W/RgwnhOs9pWde+Z9zRdQVuMUKXTQWN7pDtVZCe92sVF2B4QbWJQ9YCLBu3ni2HneneBAFaQ4ed6d2hHpwDLbTgTPu9U80X/4nC4II/0Iy8JsJI8Mf+fUBi/QLuMpsDtR2cGoXQyvlYcs0gJj/TZxrOHd7rhwe36x6i9/phLidWh3e3EoM1dk562TV8pq/NycO+zmH3+uX1jIQpLRlt05SfaOqCwcRNJaBbwb7Rs97Ax7sjp7gXfj76cV3Nj9vsoClFt4tztuzeMsC6pO81rX3Yt25jnhBHTG8rIsYhWeZDcXZTWJBO9+1efk+Iv2zlikO0b6HQ8ZMJcpw/o5OTvOSnjXD3ZecRsJPFkcjvOOdcy7+aBzE/MuCz5uPIyRDgqnA8Y4Ce1W5xvBLFK79PjMSAe/+QBTEbdoHg99Xlj9GDpBrg75VmsNooU+bk80XnK9P0YbTv1xgrtoPe+RfvC8et65zZHWasg8edjpwiPSODtppn+vwkuuAtr5R4S9mnYPrXN7LnGzYDWSA8I8hlbHpV0IotNdAhl4N/fbDRy4gTBui1OscySvM++7cPGKNk9mNJ4bI92llXWw2J1rjW3nsBrz+LuJA3dzd9QKktycT+zmtZs1NOZjPdK6DJmn08p6mK0uaPS2yM5bq/OU051smTV8kY5pc5128eNuYt5ZM87oxpvVk+plL66dLly4706/HW767nRRGImDzMesXb0JFzajTxCrWyhGADJ6vvSy5sUzXN74yFsr0DbeJjLubwU2Vl4sK5v6Z+YTIQ/4fT98MtwmK3vGW2/RLliVbYdVX63ibvUJa/Y6Gk/ETgtezDGRvK/VintzHbI/DDqYWaVZi3QxKAPnKqCP56Q+xm7h2248ZYHT6uIxUpYimVAjqxhy/Xg02pb1vBv1MY6Xm+PCaZyc4F5WT33OePGYW8172XqN5YxhDWbodTvPIoUg2o2f7LnDqr/mcXIDP8bfCbvzvNkhYT6HGD3cxmPRSzv03xHjGOifWaXu1euzO6eajytZna3mtvGC3+rCyBjOZP7NcfqN8hLzHgleZc8ndudr/XfLOPb6MWJ1PuPyep+9aGc+ZnW9zHOP28Apct0uWGX7+jwy/35xezds3hXOEhcJ3iOxi3YBanh4nWx9bpxvVmNgBODRI8/VwpaMY2R3fLq566PPNXYGO/3eur3zF0lvN46NdgfvRlA3HyBWEGaO1bK7rW2OS+ODjA8A81WZ/mNjXvxlPvk5ef7NbTdPIHN7+HtO2+h0lajbdfKuGw8AY1us4tuN3/PBHOkWplEX80nR3Cd9vCIBuDG3qUwYht5WK+Cwih918ubJhBeogj4vkObHPvoPnZVtN+E9Zr1Za75INV6ouYF3I8DrbZM9LmQuwthmpLsOfmjtt027sZRxAOhtixT+Eek7c91W5w6r85DVRY8RNLldTvNQhz3zedruQt3YVq5fz7Nt93RaXRundhghy7gPp1F1ive2WqRn9fslo7PV7yafTzkFn533WNfc7mLd6jeK6zHbk3WwGH+79P9bOeP076zOJxpwNqfrMx5bPO68lsxq/K2OEbvntug2nbSzOq7tFl0a55Bs2Iz52DTXZ+ynU1ireRy5b7wZF9K6qY/LmtexOOV5N4+B1UWz3ga749MtvMtCuZN+Zu2d7NpxULv3vPv9Y+dk3wmKnfbH91AACkCBRFPA7Q9YovUP7W3fCrgNeWrfajj3zi70xXnPxCkhw3IyzjurHsvYVq1U3GabUd1Rr/aCGBSvbcV+UAAKQAEVCuheQ6dbzCrqgg0oEI0CfKHJd044Y5q+4XfbWlFz2A6XinQ3LZpxiad9ZaBc9m6tVb+CuBsPeHeYYTgJxNMhiLZAASgQKwXs0n/Gqn7UAwVkFLAKj8RFpz28cziReXMKJ5UZh3gu4xQeqcO3l3VLer/t0rj6pQvg3S9lYRcKQAEoAAWgABSAAlAACihWAPCuWFCYgwJQAApAASgABaAAFIACfikAePdLWdiFAlAACkABKAAFoAAUgAKKFQC8KxYU5qAAFIACUAAKQAEoAAWggF8KAN79UhZ2oQAUgAJQAApAASgABaCAYgUA74oFhTkoAAWgABSAAlAACkABKOCXAoB3v5SFXSgABaAAFIACUAAKQAEooFgBwLtiQWEOCkABKAAFoAAUgAJQAAr4pQDg3S9lYRcKQAEoAAWgABSAAlAACihWAPCuWFCYgwJQAApAASgABaAAFIACfikAePdLWdiFAlAACkABKAAFoAAUgAKKFQC8KxYU5qAAFIACUAAKQAEoAAWggF8KAN79UhZ2oQAUgAJQAApAASgABaCAYgUA74oFhTkoAAWgABSAAlAACkABKOCXAoB3v5SFXSgABaAAFIACUAAKQAEooFgBwLtiQWEOCkABKAAFoAAUgAJQAAr4pQDg3S9lYRcKQAEoAAWgABSAAlAACihWAPCuWFCYgwJQAApAASgABaAAFIACfikAePdLWdiFAlAACkABKAAFoAAUgAKKFQC8KxYU5qAAFIACUAAKQAEoAAWggF8KAN79UhZ2oQAUgAJQAApAASgABaCAYgUA74oFhTkoAAWgABSAAlAACkABKOCXAoB3v5SFXSgABaAAFIACUAAKQAEooFgBwLtiQWEOCkABKAAFoAAUgAJQAAr4pQDg3S9lYRcKQAEoAAWgABSAAlAACihWAPCuWFCYgwJQAApAASgABaAAFIACfikAePdLWdiFAlAACkABKAAFoAAUgAKKFQC8KxYU5qAAFIACUAAKQAEoAAWggF8KAN79UhZ2oQAUgAJQAApAASgABaCAYgUA74oFhTkoAAWgABSAAlAACkABKOCXAoB3v5SFXSgABaAAFIACUAAKQAEooFgBwLtiQWEOCkABKAAFoAAUgAJQAAr4pQDg3S9lYRcKQAEoAAWgABSAAlAACihWAPCuWFCYgwJQAApAASgABaAAFIACfikAePdLWdiFAlAACkABKAAFoAAUgAKKFQC8KxYU5qAAFIACUAAKQAEoAAWggF8KAN79UhZ2oQAUgAJQAApAASgABaCAYgUA74oFhTkoAAWgABSAAlAACkABKOCXAoB3v5SFXSgABaAAFIACUAAKQAEooFgBwLtiQWEOCkABKAAFoAAUgAJQAAr4pQDg3S9lYRcKQAEoAAWgABSAAlAACihWAPCuWFCYgwJQAApAASgABaAAFIACfikAePdLWdiFAlAACkABKAAFoAAUgAKKFQC8KxYU5qAAFIACUAAKQAEoAAWggF8KAN79UhZ2oQAUgAJQAApAASgABaCAYgUA74oFhTkoAAWgABSAAlAACkABKOCXAoB3v5SFXSgABaAAFIACUAAKQAEooFgBwLsCQcvP1FJ5ZZ0CSzDhRoHsjBRKCSXRqYpaN7uhrCIFeuRnUOmJampobFRkEWZkFchKD1FaaohOnq6R3QXlFCrQNS+dTorzTm1dg0KrMCWjQEZaiHj+Hy/H3JfRS3WZzrnpdLqylqprm+Z+YedM1VXAnoQCgHcJkZyKAN6dFPLne8C7P7rKWgW8yyqlvhzgXb2mbiwC3t2opbYs4F2tnm6tAd7dKuZPecC7Al0B7wpE9GAC8O5BNIW7AN4ViunSFODdpWCKiwPeFQvqwhzg3YVYPhQFvPsgqgeTgHcPohl3+elPf9rKQlJSEiUlJ1MoFKKG+gbKyc2j9PR0ysjMFp8lU2VlJR07epgmTb2ONqxdRcNGjqbtWzZS3/4D6WDJfpo4ZRo9+7vH6K5vfE+UK6WlixbSjJmztTrefG0+jR57KRX07EPbNheFy/N3xbu2a3amXntTuD1F61ZTxekyGjd+cvizFcuXUHaHXBo+akyrdr+38DUaNGQY9RswqI0ikb5buvgtKijsRYOHDrdU0twHO7lrqqvphXlzaNZd35IekXnPPkl33/t1qqoLSe1j1E9qh+ZCsn1wshlJR6d97b73w6a5rjUr/619dOHYy1p95Te8Hzywlz5d+RFdd9NMr/Io308/NpUbdmlQBt6tjn+X1cRVcVXHoYpOxRre7Y5BFX1JNBtO8B6P541E0zhSe43wzufDhx9+uD11L2H6AniPcqgA74B32SnkB2j7YRPwbj+igHfZ2a6+HOC97QW0epXj3yLgPdgxArwHq79eO+A9ynEAvAPeZaeQH6Dth03AO+Bddk7HshzgHfDO8w3wHsujrm1dgPdg9Qe8K9QfMe8KxXRhCjHvLsTyoajfYTM+NLndmJQJm2k3nY3DjsQ6bCYOJQisSU7wHljDzpKKEfMeHwMNz7uCcQC8KxDRgwnAuwfRFO4CeFcopktTgHeXgikuDnhXLKgLc4B3F2L5UDQIeL9x9kPUOT+X/vzb7/vQI39NFm3eRTPv/RnNf/rHNHzoAGWVAd4VSAl4VyCiBxOAdw+iKdwF8K5QTJemAO8uBVNcHPCuWFAX5gDvLsTyoagf8P6f3/kVfbJmc6vW5nfMoWWvP6V9FgS8v/72cnro0T/RIw/cSdOvGe9ZScC7Z+n83REx74h5l51hfsSn+2HT3B9km2lRBAtWZWe7+nKIeUfMO88qJ3hHthn1x57RouqY9/OvmE1GUNfrYqDv3qUT/fIHdwcC76pUBLyrUlKxHcA74F12SvkB2n7YBLzbjyjgXXa2qy8HeAe8A97VH1duLaqEdwb07bv2hz3sdm3RPe/8ve6htwN+owffGKoyYfo3afzY4bR8ZREdP1muVXXPHTdQ757dNA+7vun7WEG3+Q4B7//Nr8wgqzsHmz6Yq5kEvLudYTEqD3gHvMtONT9A2w+bgHfAu+ycjmU5wDvgHfAeyyPOui6V8M5e9xuuulTzrkfaGN53FB/QYJthmTeG8UEDeoXj4Bmgjx0vo3/OfUT7/qnnXqU5f1lAOkRzeYZ2Hc71783hObwv2zBDt/lCg79//NmXtfr5u/vvuiUc087ttbOjagQR8x6lkklrN9C4rCz6R5eeUVrC7m4VQMy7W8XUlkfMu1o93VhDzLsbtdSXRcy7ek1lLTqFzcjaQTlvCqiKedfhWCam3Crm/cFfPEOfbdtjCdp6zxjYb71+kgb8uuddv1Cw8oizTfbMc6y98Xu2x4tOZdqqXzi89Mb7bexgwaq3OefLXgzvg1LT6IOCvr7Yh1F7BQDvwc4OwHtw+gPeg9Oeawa8B6c/4D047bnmeIR3fXGplTK6t94O3o1Azt54K+jeuadEC63RvfhW9eiefeN3XB5hM8HOV9vas9YVUVZyiDb07B+nLWy/zQK8Bzu2gPfg9Ae8B6c94D1Y7QHvweqvCt65F27CZsypIo2edx3eneCaY97NnncV8M79uPjCoeEQHmPIDuA92PlqWzti3hHzLjs1/YhP98OmuT/INtOiCBasys529eUQ846Yd55VTvCObDPqjz2jRZUx704LVhnQ7bLNWIXNRApricbzzv23C5uxunAAvPs7B5VYB7wD3mUnkh+g7YdNwLv9iALeZWe7+nKAd8A74F39ceXWokp4173v5swxOhDri1mdYt7Zjp7xxeh9Z8C/+MLztDzt0cA7x6pzG46fLAtnxtEXrPJCVTPYsyeeN4TNuJ1dMSwPeAe8y043P0DbD5uAd8C77JyOZTnAO+Ad8B7LI866LtXwbgRvY41GL7oMvNvZMWab8Ro2oy801bPe6O3U28gXCQve/SjcfI6z1zPdIGwm+Dlr2YLbivfSSydO0tyuhTQ1MztOW9k+m4WY92DHFTHvwemPmPfgtOeasWA1OP2dwmaCa9nZUbPKmPezQzF/eolUkVHq+rV9B+jpo8fot/nd6LYOeVFaw+5uFAC8u1FLfVnAu3pNZS0C3mWV8qcc4N0fXWWsAt5lVPKvDODdP23dWAa8u1HLouyPDh6inx8qpR916kL35HSK0hp2d6MA4N2NWurLAt7VayprEfAuq5Q/5QDv/ugqYxXwLqOSf2UA7/5p68Yy4N2NWhZlEfOOmHfZKeRHfLofNs39QbaZFkWwYFV2tqsvh5h3xLzzrHKCd2SbUX/sGS36EfPub4vbp3XAe5TjCngHvMtOIT9A2w+bgHf7EQW8y8529eUA74B3wLv648qtRcC7W8X8KQ94j1JXwDvgXXYK+QHaftgEvAPeZed0LMsB3gHvgPdYHnHWdQHegx8DbgHgPcpx+KTiDI3btoMuTs+kV7v3itIadnejAGLe3ailvixi3tVrKmsRMe+ySvlTDjHv/ugqY9UpbEbGBsp4VwAx7961U7kn4D1KNXfW1NDATVtocGo6vV/QJ0pr2N2NAoB3N2qpLwt4V6+prEXAu6xS/pQDvPujq4xVwLuMSv6VAbz7p60by4B3N2pZlD1ZX09dij6jTknJtL7XgCitYXc3CgDe3ailvizgXb2mshYB77JK+VMO8O6PrjJWAe8yKvlXBvDun7ZuLAPe3ahlURYx74h5l51CfsSn+2HT3B9km2lRBAtWZWe7+nKIeUfMO88qJ3hHthn1x57RImLe/dVX1jrgXVYpm3KAd8C77BTyA7T9sAl4tx9RwLvsbFdfDvAOeAe8qz+u3Fo8m+D9/Ctm08B+Pemfcx9xK5Pv5QHvUUoMeAe8y04hP0DbD5uAd8C77JyOZTnAO+Ad8B7LI866rrMF3p967lVatOxTOn6yjP7wy/tp+ND4CosGvCs4Fq7fsZveLC+neV0LaUpmtgKLMCGjAGLeZVTyrwxi3v3T1skyYt6dFPL3e8S8+6tvJOtOYTPBtezsqPlsiXm/cfZDdOWE0bR203bq3qUT/fIHd8fVAAPeFQzHrF17ad6pk/S/nbvTzdm5CizChIwCgHcZlfwrA3j3T1sny4B3J4X8/R7w7q++gPfg9HWqWTW8F4uMfeM4dCIAACAASURBVMXVNU7VKv++X3oa9UtLs7RbtHkXzbz3ZzT/6R/Tzj0l9Js5L9Ky159S3oZoDALeo1Gved/79xygJ44fo5906kp35XRUYBEmZBQAvMuo5F8ZwLt/2jpZBrw7KeTv94B3f/UFvAenr1PNquH9idIjdP+Bg07VKv/+29260uM9Cyzt6iEzeqw7x74zyMdT6AzgPcopgZh3xLzLTiE/4tP9sGnuD7LNtCiCBauys119OcS8I+adZ5VT2Ayyzag/9owWVce8zz12nJ4/ftLfRltYn5XfkWZ3zresVw+Z+eZXZmjf/+d3fhV3oTOA9yinDOAd8C47hfwAbT9sAt7tRxTwLjvb1ZcDvAPeAe/qjyu3FlXDu9v6/S6vh8yY68nvmBNXoTOA9yhnAuAd8C47hfwAbT9sAt4B77JzOpblAO+Ad8B7LI8467raO7ybQ2Z0FTh05pEH7qTp14wPfhBECwDvCobhnaOn6Jp9e+iS9Cx6pXtPBRZhQkYBxLzLqORfGcS8+6etk2XEvDsp5O/3iHn3V99I1p3CZoJr2dlRs+qY93hTbcL0b9Kt108iPWRGbx+HzvD2599+Py6aDHg3DMODv3iGFrz7UZuFCRz/tKP4gFbSKmH/qpOnaezuXTQkNY0WF/SNi4E9GxoBeA92lAHvwekPeA9Oe64Z8B6c/oD34LTnmts7vAerrnztgPdmrV5/ezn93/y3NEg3rirmq61jx8vCT9hikO+cn9vq6mtn2RkauHMHdQul0Nqe/eXVR8moFAC8RyVf1DsD3qOW0LMBwLtn6ZTsCHhXIqMnI4B3T7Ip2wnwrkzKqAwB3pvl01MB6bk99ZRAfAvlu/fcFo5zYsg35vw0x7xTUhIlJydTKBSihvoGysnNo/T0dMoQD28KhZKpsrKSjh09TJOmXkcb1q6iYSNH0/YtG6lv/4F0sGQ/TZwyjfRFceYYzzdfm0+jx15KBT370LbNReHy3IXiXds1O1OvvSk8IYrWraaK02U0bvzk8Gcrli+h7A65NHzUmFYTJ1LsdKTvli5GzLvsEehHfLofNs39QbaZFkWwYFV2tqsvh5h3xLzzrHKCd2SbUX/sGS2295h3f9VTZx3wLrRkb/qXZ06jc/oWhhPzM7wbE/XrMG/+zAnec/PyKC0tgzKzsihZwHsVw/uRwzT5qutp/dqVNHzkGNq2pYj69R9EJSX7aNKV19KcJ39F99z3fTp6pJQ+WPQvuvnzX9ZGfMGrL9CYsZdRYa8+tPWzonB5/m73zu2anas/15TaiLcNa1fT6dOn6NIJU8KffbRsMXXokEcjLmgN7+/861UaPGQ49T9nUJvZFem79xctpMLC3nTuecMtZ6W5D3ZTt7q6mv4+92n68le/LT27/++P/0tfvfcbVNMQktrHqJ/UDs2FZPvgZDOSjk772n3vh01zXas/+bf20ZiLL2v1VdeO6XTsVA01NDZ6bX7E/Ur276XVK/9NN8z4vC/2vRjVj00v+6rcJzM9RKkpyVRWUWtr1ur4V9mGWNtSdRyqaHfn3DQqO1NHtXUNKsw52rA7Bh13bIcF0tNClJmWTCdPW8/9eDxvtKdh6JSTRhVVdVRT26CxysMPP9yeupcwfTnr4Z3j3A8fPaGFwZjB3Cu8hzTPewo1NNRTx44dKT0jg7Kzhec9OURnzpyhw4cP0Y3Tp9OKj1fQ2IvH0ob1G2jwuYNp7569dP0NN9DP//tn9MMf/ZgOHzpEC95YQHfd1fRY3r/Mm0eXT7yc+vbtR+vXrwuX5++2bt2i2bnl1lvDk++TT1ZQ2akymnrVVeHP3nv3XcrNy6WLLx7XapK+/NJLNGLkCDr33CFtJm+k795YsID69O1DI0eOspz05j7YHRlVVVX0u6eepP/6f9+TPnh+/T+P0X333Udp6RlS+xj1k9qhuZBsH5xsRtLRaV+77/2waa7rww+Xah9dfvnEVl+FkpOovsEfcOeK9uwppg+Xfkh3fOlLXuVRvp9+bCo37NJgkiifJO7yRbpwsjr+XVYTV8VVHYcqOpXsoL2KOow27I5B1fUkgj2nuR+P541E0FW2jTz3G4XDhs/8fD4EvMsqp7ZcIPDOoSjHT5Zb9mTTB3PV9jCCNXMIjBd4Z/PlZ2pp+t49tKTyDP21a0+alJkVsz6czRUh5j3Y0UfMe3D6I+Y9OO25ZsS8B6e/U9hMcC07O2pGzHt8jHPM4d1qwWdQUjC8P/Tonyyrv+eOG7RUQVYx77yP8SKD4f3L+/bTP86U01NdetCMrJygunRW1Qt4D3a4Ae/B6Q94D057wHuw2gPeg9Uf8B6s/nrtMYf3eEt0bxwGqzAZmWwzDO/fKTlIfyo/ST/t1JXuzOkYH6PbzlsBeA92gAHvwekPeA9Oe8B7sNoD3oPVH/AerP6Adwv9reCdi0XK895mwSqJbDNiYSqyzTQJLJshokYsWH1h3hyadde3pI+Mec8+SXff+3WqqpNbsGrM1iNdiYs+ONn0IzOMHzbN/UC2mRZFkG3GaZb7973sucS/FrRYjnXYjN0xGIu+xlsdTvCObDP+jhiyzfirr6z1mHveGYSvnDC6zdOrZBscb+UA70gVKTsn/QBtP2wC3u1HFPAuO9vVlwO8I1UkzyrAu/pjy41FwLsbtfwrG3N4Ny8S9a9rsbHcFt5JeN5D8Lw3yy/7gwvPu7f5Cnj3ppvXvQDvXpWLfj/Zc0n0NTlbgOfdWSO/SgDe/VJWzm57h3c9AsOsxiMP3Bl+3o+cUv6Wijm8c8x7pC2W2WZUScsx72+dKKPbSg/QZelZ9FL3nqpMw04EBRDzHuz0QMx7cPoj5j047bnmWMN7sL2Nr9qd4D2+Wtv+WtPeY96twqc5pfjylUW07PWn4mZAYw7vcdNzhQ1heF9RVkFXHdxL56Wm0XsFfRVahyk7BQDvwc4NwHtw+gPeg9Me8B6s9oD3YPU/G+Fdz0wYT85lwLuC44DhfWt5FV1UspsKUlJodWF/BVZhwkkBwLuTQv5+D3j3V99I1gHvwWkPeA9We8B7sPqrhvfik8XEr1hv/Tr2I36ZN7usg1yOH+YZL1sg8G6VXz3e4olkB8gy5l08STWUEqKG+gbKyc2j9PR0ysgUT1gVWWgqKytFBpbDNGnqdbRh7SoaNnI0bd+ykfr2H0gHS/bTxCnTSI+rNcd4GrOlbNtcFC7PbS3etV2zM/Xam8JNL1q3mipOl9G48ZPDn61YvoSyO+TS8FFjWnUxUux0pO+WLsaCVdm54kd8uh82zf1BtpkWRRDzLjvb1ZdDzDsWrPKscoJ3ZJtRf+wZLaqOeX9ixRN0/zv3+9toC+vfHvdtevzqx23h3fyF/uyfmDfUpsKYw/tTz71Kc/6ygOY//WMaPnSA1iz9SifexJEZJMA74F1mnnAZP0DbD5uAd/sRBbzLznb15QDvgHfAu/rjyq1F1fA+d91cen79826bEXX5WSNn0exRs23h3cioCJsRMvETS2+9flKbVJEM9S+98X5cLQiQmR1W8J4kPO8p8Lxr8sn+4CLbjMxsa1sG8O5NN697Ad69Khf9frLnkuhrcrYQ6wWryPPeMibwvDvPTz9LqIZ3P9vqxbbd83442YoR6L3YVrlPzD3vdk9YjccrG1mhOea9vLKOZh0poUWVFfSnLgU0LauD7O4o51EBxLx7FE7Rboh5VySkBzOIefcgmsJdYg3vCpue8Kac4D3hOxjnHVAd8x5v3bWCdz1i5KxesNrePO888XR4v/94Kb10+hQ9mt+N7uiQF29zst21B/Ae7JAC3oPTH/AenPZcM+A9OP0B7/5qX3rmMFXXVdlW0iknjSqq6qimtoHy0jvSkMIe/jYoxtbt8rzHE7izJDH3vLe3mHcjvD9TdoJ+evIofS23Ez3UsUuMp9zZVx3gPdgxB7wHpz/gPTjtAe9tta+oPU3HK4/ZDkpZzSkqqz5p+/2p6sjfH686RmdqK7T9U0Tih9SUJKqsrg/b04CzvtrTpOB2cf1+bAdPH6D6xjrlpp30Vl6hjcHvjn2Ifj3t57GqDvUYFIg5vHPd7T3bTGNyMqWJlJHINoOYd+PZxo/4dD9sms+QyDbToghi3oP7/WxvMe9mD6cZ9EoY/BqawK9692kBv6eoumcLsBpHoq6xnnh/u43tsD27rbq+io4IALbb4gUWZWafSABIV4h/c8U/bO4V6JbVndJTMmx3TE5KosbGRmoUJb5S9mV6+OGH3VeCPaJWIBB4j7rVcWTAasFqg4D3dMC7NkqyP7hYsOptUgPevenmdS/Au1flot9P9lwiW1OdAFoj8JoBtqquWgDtobC5CuH5PSE8wLxlZaRQaflx8XeLN9nswT1RdZR4H30rrWDvsH04QqR2M4zy9oH4F49bdmoHys/sbNu03LQ8yhUhFnZbXnrk7/MzOlN2ara2u5XnvSsDZyhd+77uRI242Cmn7Avt22NsB7eL6/djK+jQk0JJKcpNM1wzZAextfcFq0Fo6qVOwLsX1Qz7AN6RKlJ2CvkB2n7YNPcHnvcWRQDvsrPdezndy2uE6X1le6i+XCQG2HSUTp9fGzZ+sOIA1TW0eKMZxhnK9W1/+Z7w/82w7r2FavY0ezibALTF41nI4JfcBH5dj+ZTngBgO897KClEDIp2G9the3Yb18v1221OcK5GETkrTjHvyPMup6PXUoB3r8qp3S9m8M5ZZjiPO+d4j7TF26IAGbn1BatcdvLBPbS1toY+LOhL56SmyeyOMh4VQMy7R+EU7YaYd0VCejATjzHvDNvsXeZNh2Y9NIRjistF3DN7ojl+2RjGoYeLcMw0g3usthQBtEbgZQ+p8e+MlHQBtC2L8djzyx5g3tjzHmrMEECdH26u2YPLf7PHWd/YM80QjC06BZzgPTrr2NtJgfaebcap//HyfczgPV467Ec7jPB+R+kBWlJ1hv7etZAmiqeqYvNPAcC7f9rKWAa8y6jkTxnV8M6ebd447pkhXF9AqIeKGGOqde+2HhaiOh6avcDdsrtr4QY6TOteaGP4BLfXGC7Bf3N5hnJ9M4ct9M7tq2RAkG1GiYyejADePcmmbCfAuzIpozIUc3i3y/OeqA9pYvWN8P6DE6X0fPkpeqxzd7o9OzeqwcHOkRUAvAc7QwDvwelvBe8M4OzV5thrBmz2eteIDBx6aMnxyqYYbP6es39wOjguo2ozerJ1aNa9zbrXOk3EJeuxujpI6+EinHYu16fYY1V91O0A3lUrKm8P8C6vlR8lAe9+qOreZtzAe6I+pKltzHsS1ScnUXJIPGW1oZFycvMoPT2dMoQXPiRSXFVWVopFnIdp0tTraMPaVTRs5GjavmUj9e0/kA6W7KeJU6aRHldrXqD15mvzafTYS6mgZx/atrkoXJ6HvXjXds3O1GtvCs+ConWrqeJ0GY0bPzn82YrlSyi7Qy4NHzWm1WyJFDsd6bulixHzLnvY+RGf7odNc38Q896iSCxj3nWPNoegcEgJe8MPiRhvBu6KujLNS77v1H4B4ict0/SNo3HUUfx7W/xz2nSPNXu2s0RohwbbmV20RYA6bPfK6aOZ4VASDinRw0JitXhO9YJVJ00ifR9reMcTVltGwwneEfMezcx23hcx784axaJE3MD7g794hpavLKJlrz8Vi34rq6MNvIs0SvXilSTgPRXwjmwzhpnmB2j7YRPwbn96iBbe2SPOEM4hJ2FPeU2Z5innzCb8HUO5eeGlzAmLAZzhm6Gavdq9ygopsz6dcoY0Qbgek81wzuXMMd8ydQRZBvBOdOHYy4IcgrioG/Ae7DAA3oPVX689JvBuldfdqvuPPHAnTb9mfHwoI9kKO3hvFPCeDngHvAPeJY+ktsXi0YNmhnf2iB+rPELHq8X7maNa2Mpx/lu8a39XH9Xej4mQFf6sqq5SWg8OOemc0VUD7c5ZXTRPeH5GF+os3gtyulKPnO6URnna31wmIyWzlW2rO2/SlcdhQcA74J2nJeA92IMT8B6s/jGFd2NX7WLe40MOb60wxryX1tfTxQd2U2cB758U9hOLrpK8GcVejgog5t1RIl8LtPeYdwbzfSJkZb+IJdfey/dq3nKOF9diyj3k7WYgZwDnOG9ehMkx3gzeHJLCnnH+P3vQecGmMW2geSBVL1j1daK0Q+OxDptphxJ67pITvHs2jB2lFDhbYt6ZVY3bwH496Z9zH5HSKBaFYuJ5j0VHgqzDCO/cjstFusidIl3kRz37Ud9QapBNa9d1A96DHd5Eh3d+YiVDOQM5v/aU7dLeOcZ8X9leqbSFeugJQzeDOceDM3znpuVqIM7hK0YoVzVigHdVSnqzA3j3ppuKvQDvKlT0buNsgPcJ079J48cOp1/+4O6wUDfOfgjw7n3axOeeZnj/gkgXuVSki3ypW0+6LCMrPhvdDloFeA92EOMd3s1wfuB0k+dcewlA5+8jbZyTu3duH+EV79sUQy68473F/xnS+bMgs6MA3oOd+4D34PQHvAenPdfc3uG9aPMumnnvz2j+0z+m4UMHBCt2hNpj7nnXhbFrU6I9pMkc854kwmQaxKs2OZnSGxspL7cjss0sWkgzZs6OeBDUVFfTC/Pm0Ky7viV9sMx79km6+96vizjikNQ+xmw9Ujs0F1IVa+vH4lI/bJq1iddsM4crDlJJ+X4qERlY+J1DWUoqmt/5b/F5ozgG7bYumV1F6EovzVNemNP8rv8t3guzRc5w052zaBesupl3kcrKwDti3lWp3dZOrOEd2WZaxsAJ3uNxrYx/MzH2lpXHvBcXi3R54hXrrV8/In5ZbOx5z++YG1eednMzYw7v+u2Iiy88j34z58Vwdhm+JXHlhNH0za/MiPUQRlWfFbw3CnivEfCeKsChE+CdlgLetTnmB2j7YTNe4H3Pvh30ycdLKX9sYXM4i/CcG2LQOSY90sYx4+wx173nmuec/xZe817iXU+B6OYEAHh3o5basqouolW0CvCuQkVvNgDv3nRTtZdyeH/iCaL771fVPHk73/420eOP25Y3x7zfc8cNccWnMYd3fcHqOX0L6WsPPh6Gd85IY4R5+REItqQVvJOA92oB7ykC3vMB74D35inqB2j7YTPW8M5P9NxxYpt4baXPjhZp7/x33YlqukL8myv+WW0cb87hK/07DtDeVcC509kE8O6kkH/fA96RbYZnF+Ddv2NMxrJyeJ8rzu/PPy9Ttdoys2YRzZ4tZZMfIjrnLwsonjIiBgbvnBKSQV4Pk0nUhzTxyJtj3ldVV9H0w/vokvQseqV7T6nJgULuFUDMu3vNVO7hNubdDtIZ1u029pT3yzunyXveQbyaPef8OYe7nK2bTNjM2apNLPoda897LPqUKHU4wXui9CNR29neY97txoWjRm69flLceN9jDu8cHnPe4L7aKl7j/xP1IU1W8H6wvk5LF9kzNZU+LuiXqMdo3Lcb8B7sENnB+zHxAKLdJ3ZQ8amdtEu8ik/upN0nd9Bu8f9y8UAi85aZkkX9BaD37zSw6V28+nU8hwZ0HChCW3oE28k4rR3wHuzAAN6D0x/wHpz2XHN7h3d2JP/f/LdaxbvHo3M55vBunnbGuKJ4X91rd8iYPe9c7tKSPbS3rkbkeu9PPVNSgj3a2mntgPdgBzYl4zSt2rOZdp7cLkB9lwbsu081QbpVJheG9H4ixGVA3kDNm95fADq/M6R3zy4ItjMJVjvgPdgBA7wHpz/gPTjtzwZ45z6yY3lH8YFWQsdbMpXA4T3YaRh97VYx70ki3p2zzTQ2NFB2Ti51zMikjMxsCoWSqbKyUjx19DBNmnodbVi7ioaNHE3bt2ykvv0H0sGS/TRxyjTS42rNMZ7GbCnbNheFy3Mvindt1+xMvfamcKessk2sWL6Esjvk0vBRY1p1PlLsdKTvli5+iwoKe9HgocMtxZSNU0W2GW9zMRYx7ys/WUonq05Qbc9G2nr8M9oiXjv1uPSGOsuGc1jLufnn0cBO54r3oXRu5/PF/wcTp1+U3eIxawRi3mVHT3052XOJ+prbWow1vCPbTMsYOMF7PJ43YjEnY1WH8pj3WDW8ndUTc3hvb09YtYP3OgHvDQLe0wW8dwG8I1WkOHH4Adp+2OQY9PWla6joyFpaWfIRdT2aT3UC0j8Q/8ybCki3O6fG448w4D24X0DAOxas8uwDvAd3DHLNgPdg9ddrB7xHOQ528N4g4L1OwHuoQw71yMyC5x153uMS3kvPHKY1h1bSBgHq/L6+9NM2IS+TkyZrTwk93aOylSf9sgEj6ExFKjVEyKUezeEFeLdXTyZsBnneo5l9kfeF590/bZ0sA96dFPL3e8C7v/rKWo85vCdqPvdIglrFvL9cUUbfPnaYPi9CVH6d3112PFDOhQKIeXchlii6v3wvbTm2qeV1fBNtPfaZgO+GsCFOvzhEhLgM6TxMvM6jIfnDaGiX86lHdmGbytxmm3HXWpSOpIAMvENB/xSINbz715PEs+wE74nXo8RqcXtfsJoooxFzeOcnrBrzuyeKUG7h/eOqM3Rz6QGakJFF87udvSnt/BxfwLu9uvwAo60CzjcLWN8qXk3vn9Hp2vJWO/XNGyAAXUB6FwHp+QLaBagP6jREatgA71Iy+VII8O6LrNJGAe/SUikvCHhXLqkrg4B3V3L5Vjjm8G5+apW5Z/G2oldGeSvP+766OhpXspv6p6TR8sK+MmZQxqUCgPcmwTh/+uajAs7DsC4WlR7bSBwSY9w6Z3YRi0fP1zzpukedveycBcbLBnj3opqafQDvanT0agXw7lW56PcDvEevYTQWAO/RqKdu35jDu7qmx4clu5j3UChElfX1dCoriwZmdaBc8UK2GfsxQ7YZ+fnMi0c3iAWlHx34kMrXH6WPapfTmto1bQxwGsaR3S6kC7pfRKN7jKXzu46g9FCGfEXNJe0yXfgN74h5tx8qGXhHzLvrqS69Q6zhHdlmWobGCd7j8bwhPbESoCBi3uNjkGIO73bZZvjxsy+98T4te/2p+FBGshWR4L1KwPsJAe/9BbjnA94jKgp4jzzhNh3dQO/veZdWlCwTGWA+pora09oOM8W/deJfSfpBGltwqQbrI8Trwu5jKT+zs+QsjlwM8N6iD7LNKJlSnowg2wyyzfDEAbx7OnyU7QR4VyZlVIbiBt7j8QlWMspGgvdqAe/HBbx3E3HvvcXCVXje4XkfNGQY9RswyHFqMax/tP9DDdbZw25+6BHnT7+k5wQ6r/RcGnLeCBo7bIKjTa8FAO+Ad69zR+V+gHfAO+Bd5RHlzRbg3ZtuqveKG3h/8BfP0PKVRQnneecBsYp558//WH6CfnbiKM3ukEeP5HdTPXZnvb32EvNeV1+r5VXn14aja0VIzKdicenm8PgmUVLYo86e9ZHdRou49WGBj7/fYTOBdzCOGyATNhPHzU/4psU6bCbhBVPYASfPu8KqYMpCAcS8x8e0iAm86151py4/8sCdNP2a8U7F4u57O3hfVFlBs46U0KXpmfRy915x1+5Eb1Aiw7sG6s3Avv7wp/TZsaJWw3F+lxE0snsTqHMYzIiuF8TdcAHegxsSwHtw2nPNgPfg9Ae8B6c91wx4D1Z/vfaYwLuxq+3tCavcNzt4Py7CZoYf2EXpSUm0udcA8Z4cH6PeTlqRSPDOqRsXFS+kj0uW08ciDGZf2Z5Wo9Atq7sIg7lcC4XhF4fFxPsGeA9uhADvwWkPeA9We8B7sPoD3oPVPzB4j49uq2tFpJj3hvoGOpmdRadCKTQ4N496pKZRZWUlHTt6mCZNvY42rF1Fw0aOpu1bNlLf/gPpYMl+mjhlGumL4swxnm++Np9Gj72UCnr2oW2bi8LluTfFu7ZrdqZee1O4c1bZJlYsX0LZIv5++KgxrUR4b+FrZBePHem7pYvfooLCXjR46HBLUWXjVNvjgtUtIq/6O7sW0Af7FolFph+FF5duoS3iiaV5dKmA9XGFE2hS36meYT3S2Kia5Yh5b1ESC1ZVzSr3dmTPJe4tu98j1p53ZJtpGSMneEe2Gffz2c0eiHl3o5Z/ZWPuefevK8FYdoL3quxsOizSRvbokEOD0zMA7zbD1B7gnfOtM6S/s/tN4WV/q5V3nZ9cenf6V6lL3wIaP+pK4rAYFRvgXYWK8jYA7/JaqS4JeMeCVZ5TgHfVR5Y7e4B3d3r5VToQeJ8w/Zt0/GTrJz3qHUy0hzQ5wXtjTg7tFWEzWVnZNCYzyxW8n9i2md5f+W+a8cU7NXngeW99GMx79km6+96vU1VdSOr4MOontUNzoUjQwN711Yc+plUifeOqgx/TnrLdYdPndBxMFxVeQhf1GKe97/pok+3dDTftMZYFvHtVztt+gHdvuqnYC/AOeAe8qziSorMBeI9OP1V7xxzeb5z9EHXOz6U///b7qvoQuB27mHduWFFNNU07tJd6iNCZtwv6UJdke9BM2fwZZT83h7Je+CslVVeF+9XQuTPVjB5LtcNHUu3IC6h21IVUX9gz8H4H3YAgYt45M8wqHdYPrdD+X15dFpbiogIB682vMQLaVeVaD1prq/oR8x7cqCDmPTjtueZYh80E29v4qt3J8x5frW1/rUHMe3yMaczh/WxasKoP8bUC3tcLiP9L1540WXjfzVvGu29R9h9/T+lLl7T6iqE9qaqakiqaHshj3Pi76slXUeX106l6ylRqFCE5Z9sWK3jnJ5ryIlO7cJiLxMORrux3DV0z4Hrip5qeLRvgPbiRBrwHpz3gPVjtAe/B6g94D1Z/vXbAu4JxiOR5Z/PfP3aY/lpRRt/P60z35eWHa2Qo73zbdEpb8ZH2WUNeHp2ZfRdVfOWrrTzr7JFP3bie0tavpdR1ayi1aEMroG/M7kBV115/1oG83/DO8euvbptP/9r5GnG2GH3jxaZT+11LE/tcqb3z32fjBngPbtQB78FpD3gPVnvAe7D6A96D1T8weOewmSsnjKZvfmVG5F/tyAAAIABJREFUfCgQZSucYt5zRJaZ8pQU2hhKpu4paXR+Y6OWbWbyhCn02Ut/o0vefYfWXXIp9R08lPb07UuXX3WdVLaZHUveocNbNtGNCxdq8L9lyBBaP2oU3fLeexrIn7nti/RpRhpVnC6jceMnh3uJbDNN2Xrsth0ntgpgf5Fe3fpCeMFpD+pBM1NmUpVIqMMedva08wJUt5sf8el+2DT3C9lmWhRBzLvbWa+uPGLeEfPOs8kJ3pFtRt0xZ2UJMe/+6itrPeaed35g02/mvJiQT1K1ElUG3uvT0mhVElG6yDozVjwt89TRUrphzTpanZ1JY/bso9VfuJ36Djnfc6rIUMkBOvDai7T1wF76/DPPhJv50dVX07ExY+ni62ZQ3aCmvOGA97bwXnRkHS3f/z4t3yde+z8gDpPhbUzBOBrfaxKNyRlLJzYcpBkzZ8seV5bl/ABtP2wC3u2HGfAe1SEQ1c6Ad8A74D2qQ0jJzoB3JTJGbSTm8M4x75G29pZthj3v6enptC45iU40CiAUna8/JEDwb3+ljyZOpCG3fpG2HtyvLM/7tP7nUubrr4hFr3+hVX1606mOHenqt9+m2mEj6MzML9KSfn0oq0fhWZ/nvfjUTg3UlwloX7Z3CZ2qPqlNyyGdz9OAfULvyTS+9xWUEcoUd0pKaemihYD3sZe1OnT9DpuJRw8a4D3q3xzPBlQdh54bYNgx1gtWkee9RXx43lXMYO82AO/etVO5Z8zhXWXj48WWU8w7t/PBE6U0r/wUPVF6lL724Pcode2ndOpXj2vx7X5sHE+fvvR9Sv9QvMR7yvatWjUNAuarL59ENeJVffkVVDdgoB/Vx8Sm25j3I2dKmzzsAtrZy76/fK/Wzp45vZuBfZIA9knUNbNbTNqf6JX4De+Jro+f7UfMu5/qOtuONbw7t+jsKeEE72ePEsH0FDHvwehurhXwrmAcZOD9hdOn6L+Ol9JrTz1J0//vOaq86WY6+eQcarTIPqOgSa1MhA6WaJls0j/8gNIEzIeE55+3+l69qXriZA3iGegbuiYWtMrAe1V9pQB1AevsYRevLUc3aX3nRaYThIdd87L3mUT98xL3Ikb1fJG1B3iXVUp9OcC7ek3dWAS8u1FLbVnAu1o93VoDvLtVzJ/ygcA7L1rdUXxA69EjD9xJ068ZTxxOc/GFQxMy/7sMvG+qraY5LzxP//fwjyhLPG31lAD36klX+jOqEaymbN2sQbzukU86U6GVrht6ngbwOsw3ZmTGvG1uK4wE7x8fWNbkYRfAvvrgCs10clKyBuscDsNhMSO6XuC2SpQ3KAB4D246AN6D055rBrwHpz/gPTjtuWbAe7D667XHHN6ND2niJ61+957bNHh/6rlX6aU3hHf09afiQxnJVsgsWOWY94zMbKr591JKr6mhEpFVZtL1N9OGtato2MjRtH3LRmUx71OvvSnc8qJ1qyNmmwnt3UNZL79Ama+8SP8YfQGNXLeOhmzZQnX9z6HKGbdQ5czbtf9HWhS5dPFbVFDYiwYPFalYLDbZONWa6mp6Yd4cmnXXtySVJzI/YXXj0fXNi07Zyy4WnooHKvF2QfeLBKxPot7FPWjShGsjZpuJpg9ODfdjcakfNs39QLaZFkUQ8+40y/37XvZc4l8LWizHGt4R896ivRO8x+NamVjMyVjVgZj3WCkduZ6Ywzt72Oc//WMaPnQAGeGds9A89OifqL0uWM2qq6NM4fE+lZtL+/sPoGuuvj5weDdOjUXzn6cR27fTiBf+Rsmlh8Nf8dNcX7zpJhp46eXU54KL2symoOH96i98jt4rXkrL9i3RvOwnq05obRzUaYjmYW9afDqJslM70JuvzafRYyOnigS8t1UA8A54j4efK8A7ss3wPAS8B3s0At6D1V+vPebwzsD+h1/e3wbeg/K8/+d3fkWfrNncajTMFxDGMJ+B/XrSP+c+Ei4v63nP2b2L0sRryznnULUIm5kq8rlvFZ7xID3vxk7rHtwBnbo0h9QsEfHxH1BK8S56ceZMGrFjB/Xr1bd5satY6NpvgLZ7UPD+/p73aOvC1fRk0pN0qu6U1pb0UAZN6juVbhnyRe2d/zZugHdvJx3AO+Dd28xRuxfgHfAOeFd7THmxBnj3opr6fWIO7w/+4hlavrJIC4/RPe/n9C2kmff+jG646lL65Q/uVt/LCBa5DcZQHWP7eDeG+2PHy8LAbgz70c06xbxz5pceg/tQUnUVXbN4Kb3TqSP9pWshTRahNPG8cXszFrxGWa/M1zLWkLh7wFtjegZV3XCTtui2avJUIvEQqlhsZdWn6B/i4Ulzi/5I/DAlfbuk5+U0Y/BMum7gTWft005job+5DsS8B6F6U52IeQ9Oe6451mEzwfY2vmp38rzHV2vbX2sQ8x4fYxpzeOdu6yEyRgnuueOGuHjqatHmXdqFhFVoj95280OmnOA9Y+EblP+l27Rc619//U16tuwE3dohjx7PT5zsLsnHjlGmiI9nkE9dtyY8dA2dO1Pl9TdR1fXTqfqyy30BeQZ1BvaXNv+VKmpPa3V3y+pOd11wL31p5CzqkNw9Po6ms6wVgPfgBhzwHpz2gPdgtQe8B6s/4D1Y/fXaA4H3+Oi6dSuM4TtmkOc9rD5zgveO93+Nsv4yl8q/+wAV/b8HaOLBvRQStlYU9qdu4qmribal7N5JmfP/Rpmvvkz8f31jkK+efJWAeQHyU6ZqHnqvG8euLypeKOLZ36LFxW9TZd0ZLQzmyn7X0BTtNY36diqglJB4Ym1F08JUbLFVAPAeW72NtQHeg9Me8B6s9oD3YPUHvAerf2DwrseYm+PK4yFVpA7mevpKGXi3inlPTk6mkIDy+voGys3Lo+zNn1GH48epXuRTPyMyz+wrPUTzLxhNt+4pphsuuoS2bN5I/QcMopIDe2nKVZ+j3z3+KH3j/gfo6JHDtOidhTTzi1/Wxuu1l/9OYy8ZTz179aHNm4rC5fm7XTu2aXauvX5GeGatW7OKTpeX0fiJU8KfLV+6mDrk5NKoC1svPl34xqs0ZOgwGjBwcJuZGem7JX+dS3337KHRr7xEoV0tIN+Y3YFqJ02m/dM+R2831NJtX7oz4oyvFiE6856bQ5feOoXe3SWgffdbtLKkKcXjOZ0G0VX9p9HUAdPocrH4VN+e/cMTdO/XvkF1JBe2Y9TPzeFnHgc3+xrLRtIxnmya27JyxXLto7Hjxrf6Kj8njU6crqXGRvHoYB+2A/v30sqPl9NNt3zBB+veTOrHpre91e2Vnhqi1JRkOl1pf+Fqdfyra0HsLak6DlW0PC87jU5X1Wnn+FhsdsdgLOqOtzrSUkKUnpZM7DSz2uLxvBFvGkbTntysNKqsqaPaugaNVR5++OFozGFfjwrE3PPOMea3Xj+pTYhMUAtWdd10UDeG70QL7w0NDZSXlk6Z27ZStogXT7pmGp2pPEMHDh2iv48aTZN27aAbLr2Mjgu4HzT4XNorUjd+7rob6NFf/Dc98IMf0eHDh2jhm2/Ql79yl9bMv/9tHo0fP5H6iFSTRRvWh8vzd9tEHRvFZzNuvjU8FVat/ITKyk7RlCuvCn+2eNG7lJubRxeNvbjVlHlVwPewESNpsGiHeYv03b/eXEB9+vSl4WLfpK1bKPkf/6DktxZS8idN4H2oRw/654wZdNemTdQw9WpquPpqahw5qk0dq/etonf/vpAeS3qMquqqtO/P7zqM7rzgbrpjhAiNSevQZp8nfvs/9I1v3kcpqelS09+on9QOzYXM4+BmX2PZSDrGk01zW5Yv+1D7aPwEERZl2NJSk6m2toH8QXeiveKicPnypfSF27/kVR7l++nHpnLDLg2GkomSkpJEOlR79a2Of5fVxFVxVcehik6lpTRp3+DX5Dc10u4YVNGXRLMhfGMUEnO/1mbux+N5I9E0jtTeVDH365vnPp8PAe/BjG7M4Z097Lpn29jlIFNF6nXrce7GdhnTWfLn5nY6ZZvJq62lrJIDlNEpnxpGXkCVlZV07Ohh2ihSFqZtLqJewy6ggXuLA8nzbuxnpHzhXvK8h0Sf0xe/R2WL3qa3+vSirz79h3B1Dd26U9WUq7Qnu/77vBx6bMccWrXvI/q2+Pfr5F/T1QOupy8P/yrxQtRImznPu9MhhGwzTgpZf49sMy26IM+7tzmkYi9km0G2GZ5HTmEzyPOu4mizt4FsM/7qK2s95vAeb553hnHzAlSjeNFmm+ly/VWUJm79n3h2npadRd9eO1NO3zh6iC5Iy6A/dy0Qse9yoR+yAxtP5RjkUz9dRWmrV1Lap+Il3vXMNeXCaf5JT6LVvVOow4Rr6Ir/+Cnl9x4q1fxIT1iVMoBCUSmAmPeo5ItqZ8S8RyVf1Dsj20zUEno24ATvng1jRykFEPMuJZPvhWIO7xweM+cvC8LZXLiHViErvvfcUK9VXca7A5HyvPO+dgtW01Z9Ql0+N4Xqe/Wmo/9aTPUFheGqqkSc8J1HS+h9EUbzS5F15ksi+0x7345XHqOFu/5JuxbPo1QB8OP2E00oSaM+R2vCXa/v259qxoylmtEXUa148bvdBngPdsYA3oPTH/AenPZcM+A9OP0B78FpzzUD3oPVX6895vDOFVulirQKpYkPiZxbYQfvHX73OOX+5CGqvOXzdOLp59oY+kdFGd137DANSUujxT36OleUoCVKyvfTwp3/pLd3L6CPDyzTejGq+xi69pwb6cZOl9PAbUc0bzwDfZrw0CedqWjqqYhrDEM8A71YZFvft19YBcB7sBMC8B6c/oD34LQHvAerPeA9WP0B78HqHyi8x0fX1bQiUsx7Y1UVdRRZZlKFxz1NPJ00JFaZ6THvk6ZeR+vXrqS/9u5DvfbspvNEtpmBouzEKdNIj6s1x3gaY7a3iXj5gyX7tfK8Fe/aTtu3bKSp194U7liReIJrxekyGjd+cvizFcuXUHaHXBo+akwrAVTHvLPx3ad20FsbXqczm07S43W/1eobVzheQPt0DdwLOoh4meatprqaXpg3h75840zKWPyueMrrB5S+5F3i/PLhTTwMqvriS0U6yiupeuJkem7Vcrr73q+LBa5y6TYR8+5tziPmvUU3xLx7m0Mq9kLMO2LeeR45wTti3lUcbfY2EPPur76y1gPxvMs2LhHK2cJ7cogaRUhMx5MnKXTuUMrI7dgG3jesXUVl4ru1AsSPFPaiz5WX0y1XXZ/w8L756EYtPGbhztfppIDv6eLflj47aBpD+4AbKT+zc5uh1eF91l3favUde+TTl7wnYP79VrHyXOhXD/6Avr67mOrHjBMPiJpAdYPaZsoxGgO8ezuiAO+Ad28zR+1egHfAO+Bd7THlxRrg3Ytq6vcJBN550erxk+WWvTHnf1ffZbUW7eA9hZKoQeQu71hxRsD7EMrIzLaE92EjR9OrGz6lxV260JhTJ+l7186gv//+13TXN74nstKU0tJFC2nGzNlao+Pd87720KpmaP8nFZ9qyvk+o9dtdMGpETTzC3dTVmq2rfh28G7cIfnUKc0bn75kEaUvXUK//tId9K0nnqAMcYeDN35IVA175i+dQDXjL9eeaAt4j36+A94B79HPougtAN4B74D36I+jaC0A3qNVUM3+MYd3XvzZOT+X/vzb76vpQRxYsYp5z57zO8r74fdEhplbRKaZ5yO2skzkg598cA8drK+jsRmZ9NeuhZSdJJLZJsj2cckyekvEtP9rx+t0qKKE0kLpmoedQ2OmiVeyT33J2buT0pZ/SA3LllPaJx9RSOTJN24N4gFZtSJWXoP5yy6nmlEXEonQG2xqFEDMuxodvVhBzLsX1dTtgwWr6rR0a8kpbMatPZR3pwBi3t3p5VfpmMO7XZ53vzoYC7tW8N7p3q9Q5ssv0KmfP0YV93zDsRlLq87Q46eO0Srhrb8hO4e+nZNP54qFrPG8sXf9f1c/Rq9ufYHqGuq0pl4jcrR/Z+xDdH6X1l5vP/phXrDK8J7+7w8pbeXHlLbiY0rZvrVVtdpTX4ePoFoB8dpCWPFe1/8cP5p2VtgEvAc3zID34LTnmgHvwekPeA9Oe64Z8B6s/nrtgHcF42AF712vHE+p69bQsX++LeKxIz9sSG/CvzWAP04fV1fS57I60LfzOtN5qfEH8PvK9tBvV/0iDO0pySk049zP07fGfI/65cUOhp2yzSSXHm6C+U8Y5j+i1I0b2oy25p0XT7vlEBstNaXw1NcXtiykVTA92q0JwHtwQwt4D057wHuw2gPeg9Uf8B6s/oHBO4fNXDlhNH3zKzPiQ4EoW2EX854qnqRaL57j3CG/C6VnZUWMeecsMX37D9Syx2SMv4I2PPsU/eSGm+im6hq6RCzYvP3zX9ZaGXTMe1aPXPrDmt/QvKI/UXV9FaWHMuhbeffTJeddQWNHTbBUUjZOVSbm3VyB2yes/uulv9C4UBr127aVUovWaw+MapXNprkCLXZ+9FjhpR9JteKpuAcHDqT313wSXnvgdcpEyugTTzbNbUHMe4siyDbjdaZGv5/suST6mpwtxNrzbncMOre0/ZVwgndkm/F3zBHz7q++stZj7nl3eqKpbMPjpZwlvIvGpYrUh/WhEHXo0o3S09Ol4V1PFfnX/7iNTh89QretX0MzbptNA4UHPih4f/X1eVScsZf+ceRFbSFql8yudOPgW+jGQbfQ6U3HqUBkyhk8dHjcw7s520xSbS2lbN1MKVs+o9QtTe8p/F68q1VfDhUU0Ou33Uaz9uynuiHnUa141Q0ZSnXnDHI1DQHvruSiePwRBry7G0OVpQHvWLDK8wnwrvKocm8L8O5eMz/2iDm8c8x7pK09ZJvhpaYpDO9icWSHzl09wfuN936X7t62kUavXkF/umIK3ZPTic5//126aOxlVNCzD8Uizzt7159d9zva+8k2+rRhNW0R/xjYvzfux+HwmKWL30pYeLeah0lizQEDfKoO8uL9+JHD9Ma4i+mrc+aEd2kU6xHqRJpPDeaHMsyLd/G38SFSZvuAd3enMMC7vV4yYTNWz3lwNwLxVRrwDngHvAd/TALegx8DbkHM4T0+uq22FeaY9w6PP0a5j/yETn/tPir72aOeKzteX0+PiUWsL4gnsdY1NlJBKIXuzOlIM7JzqZvw6vu5LSp+ix5e9r1wyscr+00T0P5wTBaiyvbLKeZd1o5TOX7iq9Ezn7q1yUMfOrC/1a6NWdnCK98E9EYPfXuNoUfMu9PM8e97GXj3r3ZYjnXYDBRvUcDJ8w6t/FUAMe/+6itrHfAuq1SEcmZ473TXLMp87WU6+fgf6Mwds6OuYX1NFf3wxBFaI7zCvKUkJdEVGVl0q4D4KzOzKF1hKsalexfRP0T2GH7xNqH3ZLp5yBfo5nO/EHU/VBuIFbzbtTtlswi32Shi59evpVSxNiG1aAOx5968aYthL7iQas8fQfV9+mphN/ye6BvgPbgRBLwHpz3XDHgPTn/Ae3Dac82A92D112sPBN457v2hR//USoFHHriTpl8zPj5UcdkKM7x3vWKcltnkyKLlWjpCVdvbZ07Tq2fKaVFlBVULTzxv+cIDf2NWDt0i0kuOTMvwXNWWY58JYP87vbLlb1R65jANzh+qQfstQ26nblk9PNv1c8eg4b1N3+rqtHCb1LVrKHXTBu2JsAz4VkDPaSvrBg3WUlXWDhuuPR1W+9vhKbF+6unWNuDdrWLqygPe1WnpxRLg3YtqavYBvKvR0asVwLtX5dTuF3N4f+q5V2nOXxbQ/Kd/TMOHDtB6U7R5F82892d0zx03JFwWGqsFq8kC4lJEyEtdZibl5Hb0FPMe6Qmr6T160j/Wr6biA3tp7shRmoZDDpbQBJGtJn3SNcIrn0kXpmfQNrHYteJ0GY0bPzk8a1YsX0LZHXJp+Kgx2mcnq07QKwLaD6zcQR9WL6WS9IMasN8sXiO6XqCViRSr3d5i3q0Or6hibQ1Av/DQPhqxaSOdt+xD4qfFWm2NYtx0iNfeOZa+GeytHjDlRxy9uV3INtOiCBasqv0BcmMtquPQTUUSZWMN78g20zIoTvAej2tlJKZUwhRBzHt8DFXM4X3C9G/SrddPagPpDPUvvfE+LXv9qfhQRrIVbeBd7JcswD0kPOP1AsRycvOUw7txwWqXyyfTyxXltHb7ZhooHlI0/6KLtZbzE1pv2ltMfauqacAllwuYz9Ti5I3w/uaOV+ll4Wnn+PaZ4l+oIIOmjr6Opva7tlXvAe+ltHTRQnWpIvsOoJTdO8VrF4V2iXeR3Yb/Du1qeqfmuyrGQWjo1Enz0tf3G0B1A8R7f/Eu/v5XyR4aNOJC6jfAXeYbyemtFQO8A97dzBe/ygLesWCV5xbg3a8jTM4u4F1OJ79LxRze7Z6wqofSJHy2GR3eKYnqRUYSv+GdU0vytnPnNvrksw20c8IVtKzyDG2qraZxu3ZQxzOV9LYIy+CNF7ze+NlGCiWdEnljltO/d7xMVXWVNKZgHN1YdwONv/BKGjxoWJs5B3hXDO8RQJvzzqdu2SSeDruNUnaIl3hKLMfS8wOnrLYXZ86kYQcP0cCU1Ca4791He9f+36cPcXhOtBvgHfAe7RxSsT/gHfAOeFdxJEVnA/AenX6q9o45vLc3zzsPhDHmPed/fkE5v/o5lX/n+1T+g4dVjZNrO6XC+/9x9RlaW1NNvOB1k0hdWXF6H9H+N8TrTaIK8f+sQurYdzqNOecWGp8/WIuZP1/kpGevfSJscRfz7qNooX17W3vr2VPPHnvhrU+qqrSsmdNZ1vfqI169Naiv79mr6Z1f4rM68Z1VKI5sNxDzLquU+nKIeVevqRuLsQ6bcdO29l7WyfPe3vsfdP8Q8x70CDTVH3N4b28x72Z473TvVyjz5Rfo5O+fpTO33R4Xo1xTX01/3fw3ekGEyHx26GNKTk6lrgLaq3t+jk7mN8W161uOeCrskLR0OjeUSueKd/7/4JQ06uJzakovQp1N8G6nD3vkU/bvI4b70IHmd/4/f7Z/LyWfPGkrbX1BYRPYGwFfg3wB9r17U2NObsRhAbx7mbVq9gG8q9HRqxXAu1flot8P8B69htFYALxHo566fWMO79z09pxtptuEMVqGkdJlq6lOPLwn6G3Jnne0uPYF21/RmjK571XaYlR+2BLnkd9aV0NbhXd+a20NbRMv/vuE+Ny4dUkOaSB/bqoA+tQMGiKe9nqueDHoB7kB3p3V12LpxVoILb6eX/vE/znOXvw/qeK0vQHxgLG63n2pfkBTbL0G+oU9qUG8+POG7t2pe0FHKj1RTQ0WMfrOLUOJaBQAvEejXvT7At6j19CrBcC7V+XU7Ad4V6NjtFYCgfdoGx1P+9stWE0WGV8aGhpiFvNevGs7bd+ykaZee5MmT1n1KXrqjV9S8eEdtLBxIeWm59EPLvlvOudEP8oRD3rSs83oWupx7QUC1LYIiOec8hvE+4aaShr172W0TnhotwiA0zcOreEQmys+XUU9CnvRsPNG0LnCQ8856I2bbJxqjQjreWHeHJp117ekh3fes0/S3fd+XcTtyz2w6s3X5tPosZdqT6h1s8n2wcmmH5lhorHJ8fUa3PPiWQb8HdvDcG+MsV96xRVa1yZ+8EHrLnbpQrUM9QUC6gXM87sWkiMAv75bDw30G/LynGSx/T4es0Yg24zn4Yx6R1XHYdQNEQZiDe/INtMyak7wHo/nDRVzLl5sIOY9PkYC8B7lOFjCu4D25IwMaqgPBt7/uf1l+uGH36HBlYMoPymfOp9fqIE7A7w5VaQZ3q2ylry18DVKGTCYigsLqUgAPcfQ7xDvvE1ft4aK8zvTOvHQIQb3geydT0mnEcJTP1B453uLlIibPnjXMVML4N3bRIwG3iPVyF750N6mOPvVu7dRUnk5jd8m4F6E6YRKDmjefJmNF8zWdxNgz0Av5ojmve/arenv5leD+N5qi8cfYcC7zKj7UwbwjgWrPLMA7/4cX7JWAe+ySvlbLmbwrse6W+Vyj/Sdv92P3rolvIswgmThlY41vBdtXE2vJL+ipX7k7eacW2laz+vpmiv/I9xRL/BuBYhl4gJlvfDKF73/Lu3u3IWWCc/8vrraNoL2EPD+HwLwP75qGvUXnvnBAu75vZ947ycypOiLYwHv3uaiX/BubI1ttpmak3Ri825KOnxIA3r21mux9wz3pYcomT+zyWdv7q0O9i1e/ELa07kzfVJVQTdMnEoN4nkJDeLvoDfAe3AjAHgHvAPegzv+9JoB78GPAbcgZvB+4+yHqHN+Lv35t9+37Pl/fudXdOx4Gf1z7iPxoYyLVujZZnIee4T4Vfajn9Hpb/2XCwvRF/3H1hfowQ++RRW1pyk7tQM9eMnP6I5hd1JKckr0xiUtHBQPJNotYuZ38bvwzPP/d4v/7xLvdRZx0Rki9KaPAPjeIr6a3/uIRbLa/wXY8/+dYuoR8y45MD4Vk1mwyk+X1WCeQb75PXSktOn/Lr343A320jewN589+ALmNc++CNVpzM6mBnEHiD/Xvf0cysMPvWqPG2Legx3VWIfNBNvb+KrdyfMeX61tf61BzHt8jGnM4N0uv7suQ6Lmeef26/De6T9vp8wFr9GJP/+NKm9oij33e2NYZ2hneOftyn7T6NErnqSCDj39rlrafkl9He0RAL9PgPxefgmw3ys+2ys89YfFu9XWUSyGbQL7JpBnoO/N782gnypCdADv0kPgS0EZeJetWPPaG4De6MXXQnjE98mHDxNfDLjaxHzRQF88LK0hr6MWuqODvnYhIGLy+b0+v4v2OX+fCBvgPdhRArwHpz/gPTjtuWbAe7D667UD3hWMgw7veqaZI4uWU+2oCxVYjmxizaGV9LV3Z9G+sj2at/3nl/+Gbh16h+/1qqqgurGhCeYFxGvv9eKdwb75s9Pie6uth3jYFIP9gHQRgiNymXdrSKbezd76nuJzbLFRQCW8S7dYzA3Ni3/VjvVYAAAgAElEQVT8mAjJOan9P0mE5iSXndJCdvgz/lsrU3aSeFGu242BvpFBX4d+ERbGgN+YlRX27GtefzEPuQxv2ru4UIjVBniPldLW9QDeg9Mf8B6c9oD3YLU31h4zeOeHM333ntto+jXjLXvPnvffzHmRlr3+VPyoI9ESq5j3JOE1Dokfcr9i3tM6Z9HfFz9Lu/dupxfr59PnBt5EMzrfQqEjjeFsM9z0onWrxYOZymjc+MnhnqiKedcNLl38FhWIbDODhzY9xdW8ycapmmPeOY2l5p3XvPS1mtd+jw75wovfKCp64K036Ykrr6YqAe68cc4Z9s73FUDfuxnwjd77jxa8hGwzEnPaXCTRn7CqAb7uvWcPPsM+v2sXAOL/7PE/flSUqXBciPvTn/yEHhYvu00P5dG8+MKbz0DfCvDF8xI4vEfz/osLhEa+UBChPbyolzdZz78MvFsd/x6GP252kT2XxKLBsYZ3ZJtpGVUneI/Hhe6xmJOxqgMx77FSOnI9MYP3B3/xDH22bY9tTLtTTHx8yNW2FZbwLn6gQ/wj7UO2mYwBHejlg/Np/45d4kFKQ6jn6HNo1vC76fTBk61SRXJLExneI403L5blbDdrn/8jld0ykzYLnXdyfH1tLVVHyDl+z8f/ppLzR1CHnj2pl1gL0IvDcrQ4+1QqiOA1VQUNfiwu9cNme4N3t+cOBvok4cFPETnxGeiTjx1t8vCXldFvunak76/8VPPua/H84iJAu3CUzL4j2xbN088Zq5rDfSgjXUu/qQG+iPFPTUmm5IIedIaEk0DcGdAuBkT8P18Y6BcMgHdZtd2XA7y710zVHoB3VUp6swN496ab6r1iBu/ccPa+82b2rvPnx0+Wi5SCc1X3z3d7sYL3RgGlz/3tcXqv/j1aXr6Ubu50G43Nuphuv+kerY/mPO/tGd71QbXK884Zb3YIiC8WML9T/J/f+W/+fPZHy+iDwUOpWOQnN2+c5rJX84LZ3qkC7A1wn3fyhFS6S6fJ5gdo+2HT3I9E97w7jYub752yzbBHP6mqSgvj4RAeqqrWMu/wxmE9vGllxHMN+KJAuzgQdwX4LgA1hwS5aU+ksivGjaOTPXvR1FWrtGLhuwHNO3Ee/sbmi1Z9HYBujy8Q9E2/Q6D/zQ/p0jcOL4oml7+bvqq6iHZTp11ZwLsKFb3ZALx7003VXoB3VUpGZyem8M5NZQ/8gnc/atXqiy8capuFJrruxWZvjnmvWbSEOt94DdWMu5SOvrlIacVbjn1Gzxf9keZtfJaSBGTOGna35m0fnD9UaT2JZszNglWOr99dW0f7RQjOQRGOs1+AEgP9wQbxLuCeP3PaCkQoTjfx6iruqnRLCVGBAPweyana/7XPxN+RPPhO9hPt+0Bi3hNNJI/tZU8/w7wW2iPgXr8DQHX1FDp4QPO8pxwsoZqqmqaLA3GRwBcLfNGgXTw03xHwWL3n3YxhP+G7Bs3WjN9xuBB/r23i2NFDi/hP/U4C/9+4rqDpu6YMQ0FvsYb3oPsbT/U7wXs8tbU9tgULVuNjVGMO7/HRbbWtYHiv+/sL1OmuWVqWGc42o2pbsO0Ven7jM7SiZDkN6XyegPav0peG3aXKfELbcQPvTh3lVJZNUN/kpd8v/t8E9gz5TdBvle7Syq4O+fmhZAH1HJrDcC+gP1kAfjP8c6hOom+A9+BGUCbmXeNiQzhP+G5Ac7ON3+mLfrWvxHoT7eKhedPvEPCfSeK4aP3dMW09QdCbHmZkbgevJbBKF2pXXvtcPKPDvJk/z8lKpTPV9VTTvcByobKWslQc623aE+OFzUGPix/1A979UFXeJuBdXis/SwLeFajL8N7w1O8o74HvUMWd99CpR38btdUjlaXC2/6M5nE/XnmMrh/0H5rH/ZKeE6K23V4MqIR3J00Y3I821NMRATb8OirAXvu/eD8qvKFHxHdHBdjwZ8fE/522dEqirsJj30UAfVfhsW/6f5NXnyG/C3vztb9TKE8sgI7HDfAe3KjIwntMW2gK+9HvGmhtsPiO7yZoFwQcQmS4U6DfSWj6rmVdAf/tKV1oTEXwVpldtiK7iwDbWgxrI7y1pPVedhczXmzzRZHd05Td2OO7TmniVZYvFnkryvAUXi/ipiERysY6+5SiZkuZAbxLyeR7IcB7lBL7FfO+6px19OnOFXRz8s2UMaaj5nH/6O33wtlStm0uooMl+2nilGlaDxDz7jyQb742PybZZnhBbakG8nXEOe6bIL+esj5YRDv79KP1PQq07ypsUmGae5LPITka1HNYjlhcq3nxkzUvfsWSt2nAuefT4HPOpVyfIB8x7y0j4hTz7jwL1ZSQgfezacGqvtbArC4vKLZ6NoBdeX09Qhs7IlSJQ5b0LSM1mWrqGijpQFN4U5vyvMDZIhRPD4dyOwuWXnGFtsvEDz5wu+tZV764Xz9ivWbNndsu+s5hYiqfLu36gtCkYmpKkrg510gNIuXbo1Mn08MPP9wudE60TgDeoxwx1fB+uPAonVp8kH4i/o3KvoC+kHo73f7Fr2mtNMIn4J3IasFqpOGMFbzbtcG8uJThnSGen0yrQ/4JBn8B+hyHz159vgjgiwG7beaqT2hdr960paBQK8KQn5Uk4vEF2GcnJ1E+x+kLT356EokFufxZiPIF5LNHP12sn5AJ3wG8t6gPeI/yhBnF7u15wapdtiLOeMTbykP7tfexPXo5KsjPNlAdyqQvtnasXLKA+W6L5G5asZA4r6WEkqi6tvV5UQ/v2tupIy0/ZwB9YfUaN2Yty2pZpiTWQ8lWFOR6FNk2uimnpc4FvLuRTFlZwHuUUqqC9179BtC76xbQY8d/qYH7Z0N30nfPf5BWCk/LjJmztVYC3lsPVqLDu+zU48W2pcJ7z/H4DPalAvaPNkN+l6WLaYd4KuhmAe8csy8bl2+sOzspWUA+Qz2Df9MDrzjqly8EOonPUtav0bz63S4cS7nis1xRJkWE/VzQrQOVnqgWHhjOuq9+i8d8zYB39eMsa7E9w7uTBsjz3qKQU8x7PJ43nMY3mu/11LbR2DDvG+miJTcrjSpr6qhW3Hn63doVgHeVwruwBXh3IZZdUY55z7hsHKWuW0Nenq5aUXuavrJwJi3bt4RSRGjE/0z6fUI9KVWBhJ5MxDLm3VMDA9hJB/3jAvLPCMDnxbcVAq5PMPSLC4Az4v8lAv7LxN9l4qLAK/Abu8bgzxcA7Mnn0B7eCkUsakj8nS1e7OnnjT3/vPHdAP48JC4A9Ow8fKGQLmxgk1NAJmxGzhJKeVEA2Wa8qKZmHyd4V1MLrNgpgJj3+JgbgHcF48DwnjV4AIUO7KfD67dRvcitLLsxuH/xjem0suQj6pbVnZ679kW6sMdY2d3P6nKAdzXDLwv8XNuJRhGrL4Id9X3UtKDFin4XgD9hsA81JlGeyNqTy3cAkhq19Jy88R0BvgBI5zsC4v/h8uKCwGhDdfvixR7gPdiRALwHpz/gPTjtuWbAe7D667UD3hWMw+mSUsru35sas7Lo8GfF1JiWJmX1s2NF9MSqR+lfO16jiwsuo2+PfYAu7z1Fal8UIgK8BzsLONvMrmNn6LSA+dPCk19BTe9nRBSN9rf4vILfhbf/tLgLUEHixZ8Jj7/2t3jnv7Wy2v8byDnbvlyf2eufJV4dBNwzzHcQLw4J6iDiZfnvbO3z5u+1//PFAInvQ9o+bcqKi4JkUT5eNsB7sCMBeA9Of8B7cNoD3oPV3lg74D3KsWgT8y5+4JPEj39IhAE01DdQjngQSbpIkZWRmS0+S6bKyko6dvQw9R07hD5dtZwWVv2LJmdeSRcMvZiyK7O07DF6XK05xhMx760H62yJeY80RdvbE1Z5ce6p5lSbW/cVU+malZQ37UbtIqBOeOEPNdRqcnAIULV458+Pi+OMNw4BqhcXELx/pEW+Xg95Dg+677VX6LkZt2px/3w3QN/0uwPhv8U5INcA+/qdAv17PXRI/1tfQKz/7RRGJAPvZ1O2Ga9j6nW/WMM7Yt5bRsoJ3s+2mHevc9jrfnjCqlfl1O4HeI9STy/wXlpaQus7FVGH0kwq71JJ49PG06ihY8OpHwHvcoMCeCdqb/BuHPlof4T1B2+xTX0NQJW4K3BEPHCrBfbFBYC4M3C8OaMPZ/nhTV8TYLTBn/9kwWv0E/EgtlhvZpjvKRYVp4qLgzqRso03fY2B3i7ONpQp0smGRC71qjEXh5urrzuwa3+k7EPGNQpW+/OaBW6nXxsWrBJdOPYyv+RNGLuA92CHCvAerP567YD3KMfBLbwfPnmQTh0/Tv+/vfuPlaq88zj+5fLrEvlxuejWahMoVeQWWQxthFgI9I9dMWsB3W0g2Xah0q6YLbHqbmil0V23WNmtVkJa0FqC1rhik/Zik1YbbaUiWjZaLTW4gRD8QYpW+XGp94Ll0j3P0DMchpl5fp57zpnznv5T7jy/zut5ZvzMmeeceezkY/J37VfLJ6ZfIScOHJfxH72I8P7QBlnypRuNZ4TwTng3XiwBCqqz/VvW3y3zlt9Uudi3J/p3/FDhX30LkPx3b+I+/pXnE3flUR8gjiXudKduC3o88by6RWjy3y7Dn7l3j3T09skTl051qR68jrqgWf02QaOHurXp2EGNn/+rnsPSteN52XvV/LpNqB8zS37bUVtoVPThotkPntV+G1JbP/ntyLjRw6Sn99QdN+KH7tsSH1DOvJ/WI7z7rCT/uoR3f8MQLRDeAygev/teGf6vN8n71/+LHFn93w1b3HPo/+SzP54n7/S+LVPO/Wt5eP6WykWqPNwE2PPu5haqFr+wGkpS3476LYAT0Zag+HGoLfqBoGgb3h/7Tn1ToH4noD8R/t9qcG/qNxMfOOr1Gn/zUO+5/uiCYTWORo+0LmLW6xSjhPo2ZEiT6ybiC7QbHY26hWvaj8oYE+ssjf7Obxtaufjc9RH/wur7xxqvxfMGD5X2/FyiYnSoY6OL8dUH2Lw/xo4aJsr+g+g++2OirYOTzzsn70NuyfER3gNM6wcrb5Vh//VN6fn6f8gfv/JvdVtUgV0FdxXgL7/gCnn4M91yztCRAXovbxOE92znnvCenb/JnvfsRle/Z124VxcvH4q+gWj0qGxvavKDOUeibzeONvnV4p6ovirT6KHaVmNo9Eh+O6IuXq79fYMQ35bkbc4YDwLNBG4Z0ynfmmh+dz00wwkQ3gNY/ukLy2Topo1yeO166f3HJWe1eLz/mHzmh3Pl1Xd/S3AP4B03QXgPiOnQFOHdAS1QlSKG90CHnotmXC5Yrf32pPZA1PPqgutGD/WbDWk/Tl30ne7j99EHNHXxueuj0S+sJtur3Zbm2tdA1otvwzuQfbr0pT64/jn6IKxW6rKRY+T2j576dW8eAytAePf0Ntnz/ocP3pY97++WEUNGyCfPmyk9Bw/Jp//mavntb/5XLp32Cdn92u/Y8378uPwPe96tVyMXrFqTeVXgF1a9+Lwqc8EqF6yqBcSed6+XkXdl9rx7EwZpgPDuyagL72rj3Vt9b0jfoD6Zct406Rg8tnKrSML7mfAfEN6dViLh3YnNuRLh3ZnOuyLhnfBOePd+GXk3QHj3JgzSAOHdk7FZeO+PvoLsGXRUDp84JOePvVAmjZtcvc874Z3w7rn0KtUJ7yEUzdsgvJtbhS5JeCe8E95Dv6rs2yO825ulUYPw7qva0yNy/vlycsQIORD9uqpEP+muHn0nemXV1ptl866HZMHFn5XVc74tY9s7fXujfkKAPe/ZLgf2vGfnz5737OxVzy573rMdcev0rts20zpHms8jSYZ3NcILxo3I50BbfFSEd98J3r1bZNIkOXFJl7zz3IvV1jb85l75z+dula5zL60E9xkf5sc1fKlr6xPeQ4vatUd4t/MKWZrwHlLTvi3Cu71ZqBqE91CSbu0Q3t3cQtcivPuKbtsmMnu2HJ81R97r/lmltV+8/qSs+tVNsr/nTblz7r3yuSnLfHuhfh0Bwnu2y4Lwnp0/4T07e9Uz4T07f8J7dvaqZ8J7tv5x74R3z3mo3fMu6v6/0c22Tvz5RPRDFEOls+NcGT58uLSPOEcGRz+q0tfXxwWrdcy5YNVtIbLn3c3NtRZ73l3l/Oux550972oV6cL77/e/IS/u2C5XX7PYf9HRwlkC7HnPx6IgvHvOw1nhPWqvP/rfyUEnpb2tXUaN7iC8P/VTuXbx0qbShHe3hUh4d3NzrUV4d5Xzr0d4J7wT3v1fR74tEN59BcPUJ7x7OjYK78OGDBOJfohi1OgxhHfCe2WVpRG002iz9iXx0o7nKn+afvmZ122kvW0mj2fQCO+eb5ge1QnvhHfCu8cLKFBVwnsgSM9mCO+egLJ8uch998nhdffJ6o+9Kd/a8Q2Zf/E/yPorH/JtmfoaAfa8Z7tE0g7v2R5dvntnz3u288Oe9+z8ddtmshtZOXpmz3s+5pnw7jsP11wj0t0tuzaul7/v/Y7sevd3ct+8h+Xqi671bZn6hPdcrwHCe3bTQ3jPzl71THjPzp/wnp296pnwnq1/3Dvh3XcePhVtJdi+Xb7z3Rvky++sl6s+Nl/un/eItA1q822Z+oT3XK8Bwnt200N4z86e8J6tPeE9W3/Ce7b+hPdA/vX2vLcNHhzdWWawnOw/yZ73d9+Rrex5r6y2NPanp9Fm7UuDPe+nRdjzHuiN06EZ9ryz510tG114z+O1Mg7LPbdV2POej6nhzLvBPCxYukr27NtfKXnRhAtly6bV1Vr1bhXZ1tZGeP+LkOl/cLnbjMFCrFOE8O7m5lqL8O4q51/P9L3Evyd9CwO9babRB2j9SFuvBOE92zklvGfrz5l3Q//rbl4j7x3sqQZ2FeTHdY6WjfesrLRQG94HRfd5H0R4r+qa/geX8G64IGuKEd7d3FxrEd5d5fzrmb6X+Pekb4HwrjdKqwThPS1Zs3YJ72ZOaZfizLtGePbCFXLL8kWycN6sSsnuJ7bJ3Rs2y7Pd607VjML60xNF/v3WT8v3rnpERg8fk/ac0f5fBLjbTLZLgT3v2fmz5z07e9XzQIf3bI82X73rwnu+Rtt6o2HPez7mlPDeZB527tori2+4Qx5df5tM7YoSevQ4629ReH9kqsgb69bKP136pXzMaklGQXjPdqIJ79n5E96zsye8Z2tPeM/Wn/CerX/cO+E9QHjfNLdDPvf0H2RI25B8zCqjQAABBBBAAAEEEGhJAcK7Z3ivt+ddXbA6ZMgQ6e/vl46ODmlvb5eRI0eK+ntvb68cOHBAronuD//888/LjBkz5JVXXpFLLrlEXn/9dVmwYEFlH/3tt99eKbdlyxa5/vrrK6N88MEHZc6cOTJhwgR5+eWXq+XVc6+99lqlnUWLFlWP6IUXXpAjR47IlVdeWf3bk08+KWPGjJGZM2eeceSbN2+WadOmyeTJk88SafacGt/48ePlsssuqytZewyNuI8dOyZr166VlStPXUtg8lizZo3ceOONFV+TR9LPpHxcxvQYdG02c9TVbfR8Gm3W9rV169bKn9TaG8jHvn37RPW9ZMmSgey2aV/xazM3A2oykHqv/yKMu9EYQ70Oi2iQ1WuwiFZ5fN8ooqPJmIv0fmhyPEUqQ3jXzFa9Pe+r7npAXn1mU6UmF6z+TD58wUdkUle0d6jOw/QiMy5YdXvb4IJVNzfXWlyw6irnX8/0vcS/J30LA73nnbvNnJ4T3bYZbhWpX78+Jbhg1UcvXF3Cu8aSu800vz/51qcJ76YvxzSCdhpt1h4P93k/LUJ4N13t4csR3rnPu1pVhPfwry2bFgnvNlrplSW8G9g2u8+7qn60909ytO+EQUsUCSnABashNe3b4oJVe7NQNbhgNZSkWzsDfebdbZStWUsX3lvzqPNzVFywmo+5ILwHmAfCewBEhyYI7w5oAasQ3gNiWjZFeLcEC1yc8B4Y1KI5wrsFVgpFCe8poDo0SXh3QKutQngPgOjQBOHdAS1gFcJ7QEzLpgjvlmCBixPeA4NaNEd4t8BKoSjhPQVUhyYJ7w5oySpcsMqed9MllMb+9DTarD0e9ryfFmHPu+lqD1+OPe/seVerShfeuWA1/Gsv2SJ73tP1NW2d8G4q1aAc4Z3wbrqE0gjaabRJeG88o4R309UevhzhnfBOeA//urJtkfBuK5ZOecK7pyvhnfBuuoTSCNpptEl4J7ybrumBLEd4J7wT3gfyFVe/L8J79nOgRkB4z8c8MAoEEEAAAQQQQAABBLQChHctEQUQQAABBBBAAAEEEMiHAOE9H/PAKBBAAAEEEEAAAQQQ0AoQ3rVEFEAAAQQQQAABBBBAIB8ChHePedD98qpH06WtamvarHz3E9tk1V0PnGX56jObSutre+C286Ha37lrryy+4Q55dP1tMrVrom2XpS0f0pq177+MbObjupvXyK9f2nVGp7zPmM9BSGvWvrl7o5I28/G1O++Xx3++nbXvz27VAuHdiut0YfVm/d7BHtmyaXXlj2qxj+scLRvvWenYItVsTXXl1Zv43Rs2y7Pd68B1END51mty9sIVcvDw0cpThHdz9NDWrH1z+3olbedDrfvk+4wKNNt27OS9x2AaQluz9g3QmxSxnQ+Vfb6xcln1RM267/9IHvvJL1n7ftOgrU141xLVL6DerG9ZvkgWzptVKcAbhiNkopqtqa48c+I3JzrfRq1z5t3ePbQ1a99+DpI1XOcjboPXgLl/aGvWvrl9oxMwPtmGte/nb1qb8G4qlShXb3GyYB0gPUxN5qDe16d8lW02Tya+hHczS12pNKxZ+zr1xs/7zEfcKmcfzfzTsGbtm9nXKxViPtSZ+9173+LMu/s0GNUkvBsxnVkoxAJ36Lalq9ia2pZXeLVfB7Y0qOfBufhy1tENfSCsWfvmc+MzH6qXuP7qr36x+s2see/lKjkQ1qx98zXlMx/JLZOcJDM3dy1JeHeQ81ngDt2VooqtqW355H9UeWPRLykXX8K73jX02S7Tb/zicqx9/RyFWPvLPz9fViy7Vt9ZyUsMhDVr33yR+cxH3Iv61mnDDx4X3mvM3V1KEt5d1KI69fbpqTubsGAdQR1Mbecg/jqVOTKbI1tfwruZa71SaVuz9u3mxmU+YmMu1M6XNWs//fmo7WHK3KXcsMCO3bo04d2a7FQF2yuyHbspVTWdqbqqXT3iO/zoytfeAYI7AtktJ51v7XwQ3u18k6VDW7P23efC5P29du1zkaS7t+3a11mz9t3nwmXtc6clP2/X2oR3V7mons29UD26KVXVZqb1wqKu/J59+6t+M6Z3cStPy9Wk801+mFL/P7nvUf27s2MUFy4Zmoe0TralumftG05CopjpfMRbDer1wL53M/eQ1qx9M/NmpUznQ7VR663+xrfb/nOga4HwrhPieQQQQAABBBBAAAEEciJAeM/JRDAMBBBAAAEEEEAAAQR0AoR3nRDPI4AAAggggAACCCCQEwHCe04mgmEggAACCCCAAAIIIKATILzrhHgeAQQQQAABBBBAAIGcCBDeczIRDAMBBBBAAAEEEEAAAZ0A4V0nxPMIIIAAAggggAACCOREgPCek4lgGAgggAACCCCAAAII6AQI7zohnkcAAQQQQAABBBBAICcChPecTATDQAABBBBAAAEEEEBAJ0B41wnxPAIIIIAAAggggAACOREgvOdkIhgGAggggAACCCCAAAI6AcK7TojnEUAAAQQQQAABBBDIiQDhPScTwTAQQAABBBBAAAEEENAJEN51QjyPAAIIIIAAAggggEBOBAjvOZkIhoEAAggggAACCCCAgE6A8K4T4nkEEEAAAQQQQAABBHIiQHjPyUQwDAQQQAABBBBAAAEEdAKEd50QzyOAAAIIIIAAAgggkBMBwntOJoJhIIBAdgLrvv8j2fCDx88awPLPz5cVy66V2QtXVJ57tnvdWWXUc50do2XLptWV53RtTZm7tOmBdnaMqvRz3c1r5Ncv7apbdvVXvygL582SBUtXyZ59+yX+d1y4+4ltsuquB+SiCRdWx1XbkMk4Zl0+VR7/+fZq1fl/e4V889Z/turX5Diym3l6RgABBIonQHgv3pwxYgQQCCgQh8tH198mU7smVltWIfypZ1+shl8VdmdM75KN96yslvnanffLth07q6HetK3akF0bvtXzqq33DvY0DN+qTBzea8cV/71ZeE8SxmG/3jjqPWfTr8lxBJxOmkIAAQRaXoDw3vJTzAEigEAzARXK4zPKzcrVhtidu/bK4hvuOOOst2lbIcP7uM7RlTP08YePeFwq0OvCv8k4GoV3034J77z+EEAAgbAChPewnrSGAAIFE1DbXi6e+JEzzqg3OgQVRHfvfatypl2dfVYBNnkm3qYt1UezM94moVeN4eOTxsvb7x6SD507trKlRX0boB7qb2mGd9N+TY6jYEuG4SKAAAKZChDeM+WncwQQyFogDtDxOOI9543Gldwr/uozm84oZtuWLryb7HlXIXrG9I9X9rir8ajxqbPw3/7eD1MP7yb9suc96xVO/wgg0GoChPdWm1GOBwEEnAXiLSdxA/W208SBO76YtVFnNm357HlX4T2+iFSNJf42wOaMt8ued9N+bcbhPHFURAABBEokQHgv0WRzqAggYC6gtp+oO63Unl2vt9dd12qjtnRn3nXbXuJtMyq8x3e5iT8I2IRmn/Cu69dmHDpHnkcAAQQQECG8swoQQKC0AiqIP/LjpypnrmsfcSitvQtNo/Du0lbI8K7Gr/bcx7eztAnNPuFd16/NOEq7EDlwBBBAwEKA8G6BRVEEEGgtgeTWluQZ9uQdW5IXpKqjbxbe1d1n1MO0rdDhPTk7NqHZN7w369dmHK21ujgaBBBAIB0Bwns6rrSKAAIFEqj3g0WN9rTrts3YtKUL76YXrNb75sAmNDcaR7zdJ57K5I80xXvea6e5tl8uWC3QC4GhIoBAIQQI74WYJgaJAGMB9mQAAALKSURBVAIIIIAAAggggAB73lkDCCCAAAIIIIAAAggURoAz74WZKgaKAAIIIIAAAgggUHYBwnvZVwDHjwACCCCAAAIIIFAYAcJ7YaaKgSKAAAIIIIAAAgiUXYDwXvYVwPEjgAACCCCAAAIIFEaA8F6YqWKgCCCAAAIIIIAAAmUXILyXfQVw/AgggAACCCCAAAKFESC8F2aqGCgCCCCAAAIIIIBA2QUI72VfARw/AggggAACCCCAQGEECO+FmSoGigACCCCAAAIIIFB2AcJ72VcAx48AAggggAACCCBQGAHCe2GmioEigAACCCCAAAIIlF2A8F72FcDxI4AAAggggAACCBRGgPBemKlioAgggAACCCCAAAJlFyC8l30FcPwIIIAAAggggAAChREgvBdmqhgoAggggAACCCCAQNkFCO9lXwEcPwIIIIAAAggggEBhBAjvhZkqBooAAggggAACCCBQdgHCe9lXAMePAAIIIIAAAgggUBgBwnthpoqBIoAAAggggAACCJRdgPBe9hXA8SOAAAIIIIAAAggURoDwXpipYqAIIIAAAggggAACZRcgvJd9BXD8CCCAAAIIIIAAAoURILwXZqoYKAIIIIAAAggggEDZBQjvZV8BHD8CCCCAAAIIIIBAYQQI74WZKgaKAAIIIIAAAgggUHYBwnvZVwDHjwACCCCAAAIIIFAYAcJ7YaaKgSKAAAIIIIAAAgiUXYDwXvYVwPEjgAACCCCAAAIIFEaA8F6YqWKgCCCAAAIIIIAAAmUXILyXfQVw/AgggAACCCCAAAKFESC8F2aqGCgCCCCAAAIIIIBA2QUI72VfARw/AggggAACCCCAQGEECO+FmSoGigACCCCAAAIIIFB2AcJ72VcAx48AAggggAACCCBQGAHCe2GmioEigAACCCCAAAIIlF2A8F72FcDxI4AAAggggAACCBRG4P8BpxA6jc+0Ts4AAAAASUVORK5CYII=",
"text/html": [
""
]
},
"metadata": {},
"output_type": "display_data"
}
],
"source": [
"dynamics.plot_history(colors=[\"darkturquoise\", \"green\", \"red\"],\n",
" title=\"Coupled reactions A <-> B and A <-> S (fast but disadvantaged energetically)\",\n",
" show_intervals=True)"
]
},
{
"cell_type": "markdown",
"id": "f4a147c6-cad1-47ca-bc1e-79b43a29e5c0",
"metadata": {},
"source": [
"### Notice how the \"alternate (shunt) path\" of the reaction, i.e. S (red) \n",
"### has a FAST START (fast kinetics),\n",
"### but EVENTUALLY PETERS OUT (energy dis-advantage)"
]
},
{
"cell_type": "code",
"execution_count": 17,
"id": "b139f5e4-625f-4a5e-8f57-8f00244dced4",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"0: A <-> B\n",
"Final concentrations: [A] = 5.887 ; [B] = 35.24\n",
"1. Ratio of reactant/product concentrations, adjusted for reaction orders: 5.98644\n",
" Formula used: [B] / [A]\n",
"2. Ratio of forward/reverse reaction rates: 6\n",
"Discrepancy between the two values: 0.226 %\n",
"Reaction IS in equilibrium (within 1% tolerance)\n",
"\n",
"1: A <-> S\n",
"Final concentrations: [A] = 5.887 ; [S] = 8.873\n",
"1. Ratio of reactant/product concentrations, adjusted for reaction orders: 1.50724\n",
" Formula used: [S] / [A]\n",
"2. Ratio of forward/reverse reaction rates: 1.5\n",
"Discrepancy between the two values: 0.4824 %\n",
"Reaction IS in equilibrium (within 1% tolerance)\n",
"\n"
]
},
{
"data": {
"text/plain": [
"True"
]
},
"execution_count": 17,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"# Verify that all the reactions have reached equilibrium\n",
"dynamics.is_in_equilibrium()"
]
},
{
"cell_type": "code",
"execution_count": null,
"id": "f87fe3c7-2c0d-4e3c-aff0-350bd04ea1a7",
"metadata": {},
"outputs": [],
"source": []
},
{
"cell_type": "markdown",
"id": "9e9260a7-5777-4eee-93b8-c46b8364e24f",
"metadata": {},
"source": [
"# Scenario 3 - downregulated by shunt: \n",
"### kinetically slow, \n",
"### but with thermodynamical advantage (i.e. energetically favored)"
]
},
{
"cell_type": "code",
"execution_count": 18,
"id": "04476dab-f845-4533-8f26-fd0eecf8304f",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"Number of reactions: 2 (at temp. 25 C)\n",
"0: A <-> B (kF = 30 / kR = 5 / delta_G = -4,441.7 / K = 6) | 1st order in all reactants & products\n",
"1: A <-> S (kF = 3 / kR = 0.1 / delta_G = -8,431.4 / K = 30) | 1st order in all reactants & products\n",
"Set of chemicals involved in the above reactions: {'S', 'A', 'B'}\n"
]
}
],
"source": [
"# Specify the chemicals (notice that we're starting with new objects)\n",
"new_chem_data = chem(names=[\"A\", \"B\", \"S\"])\n",
"\n",
"# Reaction A <-> B (as before)\n",
"new_chem_data.add_reaction(reactants=[\"A\"], products=[\"B\"],\n",
" forward_rate=30., reverse_rate=5.) \n",
"\n",
"# Reaction A <-> S (slow shunt, excellent thermodynamical energetic advantage)\n",
"new_chem_data.add_reaction(reactants=[\"A\"], products=[\"S\"],\n",
" forward_rate=3., reverse_rate=0.1)\n",
"\n",
"new_chem_data.describe_reactions()"
]
},
{
"cell_type": "code",
"execution_count": 19,
"id": "a45c9ebf-d06e-443f-a1a1-a5b2db6803b7",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"SYSTEM STATE at Time t = 0:\n",
"3 species:\n",
" Species 0 (A). Conc: 50.0\n",
" Species 1 (B). Conc: 0.0\n",
" Species 2 (S). Conc: 0.0\n",
"Set of chemicals involved in reactions: {'S', 'A', 'B'}\n"
]
}
],
"source": [
"dynamics = UniformCompartment(chem_data=new_chem_data, preset=\"small_rel_change\")\n",
"dynamics.set_conc(conc={\"A\": 50.}, snapshot=True)\n",
"dynamics.describe_state()"
]
},
{
"cell_type": "markdown",
"id": "e6bddc07-e9cf-4236-bd2b-0e1fe0567800",
"metadata": {
"tags": []
},
"source": [
"### Run the reaction"
]
},
{
"cell_type": "code",
"execution_count": 20,
"id": "3800cb7b-475e-4437-a982-af2760355795",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"454 total step(s) taken\n",
"Number of step re-do's because of negative concentrations: 0\n",
"Number of step re-do's because of elective soft aborts: 1\n",
"Norm usage: {'norm_A': 17, 'norm_B': 17, 'norm_C': 16, 'norm_D': 16}\n"
]
}
],
"source": [
"dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()\n",
"\n",
"# The changes of concentrations vary very rapidly early on; automated variable timesteps will take care of that\n",
"dynamics.single_compartment_react(initial_step=0.005, duration=7.0,\n",
" snapshots={\"initial_caption\": \"1st reaction step\",\n",
" \"final_caption\": \"last reaction step\"},\n",
" variable_steps=True)"
]
},
{
"cell_type": "code",
"execution_count": 21,
"id": "ec552dd4-4283-4d5c-9fe2-1eb796475848",
"metadata": {},
"outputs": [
{
"data": {
"application/vnd.plotly.v1+json": {
"config": {
"plotlyServerURL": "https://plot.ly"
},
"data": [
{
"hovertemplate": "Chemical=A
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "A",
"line": {
"color": "darkturquoise",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "A",
"showlegend": true,
"type": "scattergl",
"x": [
0,
0.00125,
0.0025,
0.0028125,
0.0031249999999999997,
0.0034374999999999996,
0.0037499999999999994,
0.004062499999999999,
0.0043749999999999995,
0.0046875,
0.005,
0.0053125,
0.005625000000000001,
0.005937500000000001,
0.006250000000000001,
0.0065625000000000015,
0.006875000000000002,
0.007187500000000002,
0.007500000000000002,
0.007812500000000002,
0.008125000000000002,
0.008437500000000002,
0.008750000000000003,
0.009062500000000003,
0.009375000000000003,
0.009687500000000003,
0.010000000000000004,
0.010312500000000004,
0.010625000000000004,
0.010937500000000005,
0.011250000000000005,
0.011562500000000005,
0.011875000000000005,
0.012187500000000006,
0.012500000000000006,
0.012812500000000006,
0.013125000000000006,
0.013437500000000007,
0.013750000000000007,
0.014062500000000007,
0.014375000000000008,
0.014687500000000008,
0.015000000000000008,
0.015312500000000008,
0.015625000000000007,
0.015937500000000007,
0.016250000000000007,
0.016562500000000008,
0.016875000000000008,
0.01718750000000001,
0.01750000000000001,
0.01781250000000001,
0.01812500000000001,
0.01843750000000001,
0.01875000000000001,
0.01906250000000001,
0.01937500000000001,
0.01968750000000001,
0.02000000000000001,
0.02031250000000001,
0.02062500000000001,
0.02093750000000001,
0.021250000000000012,
0.021562500000000012,
0.021875000000000012,
0.022187500000000013,
0.022500000000000013,
0.022812500000000013,
0.023125000000000014,
0.023437500000000014,
0.023750000000000014,
0.024062500000000014,
0.024375000000000015,
0.024687500000000015,
0.025000000000000015,
0.025312500000000016,
0.025625000000000016,
0.025937500000000016,
0.026250000000000016,
0.026562500000000017,
0.026875000000000017,
0.027187500000000017,
0.027500000000000017,
0.027812500000000018,
0.028125000000000018,
0.02843750000000002,
0.02875000000000002,
0.02906250000000002,
0.02937500000000002,
0.02968750000000002,
0.03000000000000002,
0.03031250000000002,
0.03062500000000002,
0.03093750000000002,
0.03125000000000002,
0.03156250000000002,
0.03187500000000002,
0.03218750000000002,
0.03250000000000002,
0.03281250000000002,
0.03328125000000002,
0.033750000000000016,
0.03421875000000001,
0.03468750000000001,
0.03515625000000001,
0.035625000000000004,
0.03609375,
0.0365625,
0.037031249999999995,
0.03749999999999999,
0.03796874999999999,
0.038437499999999986,
0.03890624999999998,
0.03937499999999998,
0.039843749999999976,
0.04031249999999997,
0.04078124999999997,
0.04124999999999997,
0.041718749999999964,
0.04218749999999996,
0.04265624999999996,
0.043124999999999955,
0.04359374999999995,
0.04406249999999995,
0.044531249999999946,
0.04499999999999994,
0.04546874999999994,
0.04593749999999994,
0.046406249999999934,
0.04687499999999993,
0.04734374999999993,
0.047812499999999925,
0.04828124999999992,
0.04874999999999992,
0.049218749999999915,
0.04968749999999991,
0.05015624999999991,
0.050624999999999906,
0.0510937499999999,
0.0515624999999999,
0.0520312499999999,
0.052499999999999894,
0.05296874999999989,
0.05343749999999989,
0.053906249999999885,
0.05437499999999988,
0.05484374999999988,
0.055312499999999876,
0.05578124999999987,
0.05624999999999987,
0.056718749999999866,
0.05718749999999986,
0.05765624999999986,
0.05812499999999986,
0.058593749999999854,
0.05929687499999985,
0.05999999999999985,
0.06070312499999985,
0.06140624999999985,
0.06210937499999985,
0.06281249999999985,
0.06351562499999985,
0.06421874999999985,
0.06492187499999985,
0.06562499999999985,
0.06632812499999985,
0.06703124999999985,
0.06773437499999985,
0.06843749999999985,
0.06914062499999984,
0.06984374999999984,
0.07054687499999984,
0.07124999999999984,
0.07195312499999984,
0.07265624999999984,
0.07335937499999984,
0.07406249999999984,
0.07476562499999984,
0.07546874999999983,
0.07617187499999983,
0.07722656249999983,
0.07828124999999983,
0.07933593749999983,
0.08039062499999983,
0.08144531249999983,
0.08249999999999982,
0.08355468749999982,
0.08460937499999982,
0.08566406249999982,
0.08671874999999982,
0.08777343749999982,
0.08882812499999981,
0.08988281249999981,
0.09093749999999981,
0.09199218749999981,
0.09357421874999981,
0.09515624999999982,
0.09673828124999982,
0.09832031249999983,
0.09990234374999983,
0.10148437499999984,
0.10306640624999984,
0.10464843749999984,
0.10623046874999985,
0.10860351562499986,
0.11097656249999986,
0.11334960937499987,
0.11572265624999988,
0.11809570312499988,
0.12046874999999989,
0.12402832031249988,
0.1275878906249999,
0.1311474609374999,
0.13470703124999991,
0.13826660156249992,
0.14182617187499993,
0.14538574218749994,
0.14894531249999995,
0.15250488281249996,
0.15606445312499997,
0.15962402343749998,
0.16318359375,
0.1667431640625,
0.17030273437500001,
0.17386230468750002,
0.17742187500000003,
0.18098144531250004,
0.18454101562500005,
0.18810058593750006,
0.19166015625000007,
0.19521972656250008,
0.1987792968750001,
0.2023388671875001,
0.20589843750000011,
0.20945800781250012,
0.21301757812500013,
0.21657714843750014,
0.22013671875000015,
0.22369628906250016,
0.22903564453125017,
0.23437500000000017,
0.23971435546875017,
0.24505371093750017,
0.25039306640625014,
0.2557324218750001,
0.2610717773437501,
0.26641113281250006,
0.27175048828125004,
0.27708984375,
0.28242919921875,
0.28776855468749996,
0.29310791015624993,
0.2984472656249999,
0.3037866210937499,
0.30912597656249985,
0.3144653320312498,
0.3198046874999998,
0.32514404296874977,
0.33048339843749974,
0.3358227539062497,
0.3411621093749997,
0.34650146484374966,
0.35184082031249964,
0.3571801757812496,
0.3625195312499996,
0.36785888671874956,
0.37319824218749953,
0.3785375976562495,
0.3838769531249995,
0.38921630859374945,
0.3945556640624994,
0.3998950195312494,
0.4079040527343744,
0.4159130859374994,
0.42392211914062444,
0.43193115234374946,
0.4399401855468745,
0.4479492187499995,
0.4559582519531245,
0.4639672851562495,
0.47197631835937454,
0.47998535156249955,
0.48799438476562457,
0.4960034179687496,
0.5040124511718745,
0.5120214843749995,
0.5200305175781245,
0.5280395507812494,
0.5360485839843744,
0.5440576171874993,
0.5520666503906243,
0.5600756835937493,
0.5680847167968742,
0.5760937499999992,
0.5841027832031241,
0.5921118164062491,
0.6001208496093741,
0.608129882812499,
0.616138916015624,
0.6241479492187489,
0.6321569824218739,
0.6401660156249989,
0.6521795654296864,
0.6641931152343739,
0.6762066650390613,
0.6882202148437488,
0.7002337646484363,
0.7122473144531238,
0.7242608642578113,
0.7362744140624988,
0.7482879638671863,
0.7603015136718738,
0.7723150634765613,
0.7843286132812488,
0.7963421630859363,
0.8083557128906238,
0.8203692626953113,
0.8323828124999988,
0.8443963623046863,
0.8564099121093738,
0.8684234619140613,
0.8804370117187488,
0.8924505615234363,
0.9044641113281238,
0.9164776611328113,
0.9284912109374988,
0.9405047607421863,
0.9525183105468737,
0.970538635253905,
0.9885589599609363,
1.0065792846679675,
1.0245996093749987,
1.04261993408203,
1.060640258789061,
1.0786605834960923,
1.0966809082031235,
1.1147012329101547,
1.1327215576171858,
1.150741882324217,
1.1687622070312482,
1.1867825317382794,
1.2048028564453106,
1.2228231811523418,
1.240843505859373,
1.2588638305664042,
1.2768841552734354,
1.2949044799804665,
1.3129248046874977,
1.330945129394529,
1.34896545410156,
1.375995941162107,
1.4030264282226539,
1.4300569152832008,
1.4570874023437477,
1.4841178894042946,
1.5111483764648415,
1.5381788635253884,
1.5652093505859352,
1.5922398376464821,
1.619270324707029,
1.646300811767576,
1.6733312988281228,
1.7003617858886697,
1.7273922729492166,
1.7544227600097635,
1.7814532470703104,
1.8084837341308573,
1.8355142211914042,
1.862544708251951,
1.889575195312498,
1.9166056823730448,
1.9436361694335917,
1.9706666564941386,
1.9976971435546855,
2.0382428741455056,
2.0787886047363258,
2.119334335327146,
2.159880065917966,
2.200425796508786,
2.2409715270996062,
2.2815172576904263,
2.3220629882812465,
2.3626087188720666,
2.4031544494628867,
2.443700180053707,
2.484245910644527,
2.524791641235347,
2.565337371826167,
2.6058831024169873,
2.6464288330078074,
2.6869745635986275,
2.7275202941894476,
2.7680660247802678,
2.808611755371088,
2.849157485961908,
2.889703216552728,
2.930248947143548,
2.9707946777343683,
3.0113404083251885,
3.0518861389160086,
3.0924318695068287,
3.132977600097649,
3.173523330688469,
3.214069061279289,
3.254614791870109,
3.2951605224609293,
3.3357062530517494,
3.3762519836425695,
3.4167977142333896,
3.4573434448242097,
3.49788917541503,
3.53843490600585,
3.57898063659667,
3.61952636718749,
3.6600720977783103,
3.7208906936645407,
3.781709289550771,
3.8425278854370015,
3.903346481323232,
3.9641650772094623,
4.024983673095693,
4.085802268981923,
4.1466208648681535,
4.207439460754384,
4.268258056640614,
4.329076652526845,
4.389895248413075,
4.4507138442993055,
4.511532440185536,
4.572351036071766,
4.633169631957997,
4.693988227844227,
4.7548068237304575,
4.815625419616688,
4.876444015502918,
4.967671909332264,
5.0588998031616095,
5.150127696990955,
5.241355590820301,
5.332583484649646,
5.423811378478992,
5.5150392723083375,
5.606267166137683,
5.697495059967029,
5.788722953796374,
5.87995084762572,
5.971178741455065,
6.108020582199084,
6.244862422943103,
6.381704263687122,
6.518546104431141,
6.65538794517516,
6.7922297859191785,
6.929071626663197,
7.134334387779226
],
"xaxis": "x",
"y": [
50,
47.9375,
45.9718203125,
45.503467606689455,
45.040610612130536,
44.583184741656176,
44.13112616716149,
43.68437181068289,
43.242859335582004,
42.80652713783324,
42.37531433741376,
41.94916076979458,
41.52800697753174,
41.11179420195624,
40.70046437496171,
40.29396011088851,
39.89222469850329,
39.495202093072756,
39.10283690853061,
38.71507440973658,
38.33186050482635,
37.953141737651485,
37.57886528030812,
37.20897892575354,
36.843431080509475,
36.48217075745115,
36.125147568681136,
35.77231171848688,
35.42361399638108,
35.079005770223745,
34.73843897942521,
34.40186612822897,
34.06924027907342,
33.7405150460317,
33.41564458832859,
33.094583603933565,
32.77728732322922,
32.46371150275405,
32.15381241901881,
31.847546862395507,
31.544872131078236,
31.24574602511499,
30.95012684050962,
30.657973363393076,
30.369244864263205,
30.08390109229219,
29.801902269700932,
29.52320908619953,
29.247782693493097,
28.97558469985216,
28.706577164746857,
28.440722593544198,
28.17798393226766,
27.918324562418356,
27.661708295857064,
27.408099369746434,
27.157462441552592,
26.909762584105536,
26.66496528071754,
26.42303642035896,
26.183942292890737,
25.9476495843529,
25.71412537230847,
25.483337121242084,
25.25525267801267,
25.029840267359607,
24.807068487461645,
24.58690630554808,
24.36932305356146,
24.15428842387128,
23.941772465038063,
23.731745577627223,
23.524178510072097,
23.319042354585637,
23.11630854312012,
22.915948843374363,
22.71793535484784,
22.52224050494119,
22.328837045102567,
22.137698047019235,
21.948796898853967,
21.762107301525656,
21.577603265033638,
21.395259104825218,
21.21504943820588,
21.036949180791698,
20.860933543003426,
20.686978026601807,
20.51505842126358,
20.345150801197747,
20.177231521801605,
20.011277216356067,
19.84726479275982,
19.68517143030189,
19.524974576472104,
19.366651943809064,
19.210181506785144,
19.055541498728118,
18.902710408778947,
18.751666978885332,
18.52775181180322,
18.30776489839474,
18.091636992706473,
17.87930006950633,
17.670687302763703,
17.46573304450902,
17.26437280406592,
17.066543227649603,
16.872182078324794,
16.681228216317045,
16.493621579671107,
16.3093031652503,
16.1282150100708,
15.95030017296503,
15.775502716568253,
15.60376768962276,
15.435041109593989,
15.269269945593106,
15.106402101600649,
14.946386399985913,
14.789172565316864,
14.63471120845547,
14.482953810933408,
14.333852709603219,
14.18736108156003,
14.043432929329098,
13.902023066314472,
13.763087102504185,
13.626581430427443,
13.492463211359366,
13.36069036176894,
13.23122154000585,
13.104016133222029,
12.979034244523758,
12.856236680350252,
12.735584938074755,
12.617041193824212,
12.500568290513659,
12.386129726091557,
12.273689641992348,
12.163212811792574,
12.054664630066979,
11.94801110144106,
11.843218829836612,
11.740255007906866,
11.639087406657852,
11.539684365252757,
11.442014780995988,
11.346048099493826,
11.251754304988525,
11.159103910862813,
11.068067950311784,
10.97861796717924,
10.890726006955568,
10.80436460793432,
10.677077884819875,
10.553114595310111,
10.43238687191389,
10.314809170631266,
10.200298209512985,
10.088772908844662,
9.980154332912683,
9.874365633309992,
9.771331993741068,
9.670980576286421,
9.573240469088036,
9.478042635418173,
9.385319864094955,
9.29500672120912,
9.20703950312728,
9.12135619073792,
9.037896404907269,
8.956601363113075,
8.877413837225113,
8.80027811240211,
8.725139947075567,
8.65194653399174,
8.58064646228379,
8.51118968054687,
8.443527460889623,
8.34465481545959,
8.249609280014845,
8.158239102199522,
8.070398548855577,
7.985947667272658,
7.904752055907955,
7.8266826442003925,
7.751615481118442,
7.679431532095141,
7.6100164840176365,
7.543260557951775,
7.479058329294929,
7.41730855506243,
7.35791400802463,
7.300781317422895,
7.218340565293883,
7.140593489868632,
7.067260995140461,
6.998080590395847,
6.93280540224666,
6.871203245443679,
6.8130557489740635,
6.7581575341535585,
6.706315441619872,
6.632863985666714,
6.565495622342111,
6.503668012642315,
6.446887218369929,
6.394703382568518,
6.346706795460069,
6.280433069491154,
6.221976425326633,
6.170292248126131,
6.124475760963174,
6.083743304922147,
6.047416125202848,
6.014906327756855,
5.9857047158896135,
5.9593702551598104,
5.935520948598144,
5.913825933447956,
5.893998635904289,
5.875790842218578,
5.858987563496375,
5.843402587937505,
5.828874628491694,
5.815263986222215,
5.8024496603404305,
5.790326845116018,
5.778804761872363,
5.767804781209725,
5.7572587966037965,
5.747107815728363,
5.737300740355653,
5.72779330958973,
5.71854718456777,
5.709529155691101,
5.700710455983126,
5.69206616636706,
5.679328967653936,
5.666893548309878,
5.654705235267859,
5.642720318410605,
5.63090384921902,
5.619227881458357,
5.6076700651396525,
5.5962125228177175,
5.584840951531689,
5.573543905078432,
5.562312220407454,
5.5511385591972875,
5.540017041484575,
5.528942952861405,
5.517912510468176,
5.506922675975676,
5.495971006120795,
5.485055533255019,
5.4741746698790115,
5.463327132346845,
5.452511879890534,
5.441728065888519,
5.43097499891948,
5.420252111636559,
5.409558935891619,
5.398895082854533,
5.388260227124469,
5.377654094031582,
5.367076449488452,
5.35652709187929,
5.346005845577714,
5.335512555766072,
5.325047084294959,
5.30939041741361,
5.293795804349249,
5.278262889122162,
5.262791350740035,
5.247380892944181,
5.232031237043137,
5.216742116903688,
5.201513275449505,
5.1863444622133095,
5.171235431625219,
5.156185941815565,
5.1411957537771995,
5.1262646307790405,
5.111392337955163,
5.096578642016583,
5.0818233110487725,
5.067126114369088,
5.052486822426079,
5.037905206728054,
5.023381039792104,
5.008914095107427,
4.994504147108639,
4.980150971156085,
4.965854343521027,
4.951614041374258,
4.937429842777111,
4.923301526674139,
4.909228872886978,
4.895211662109029,
4.881249675900722,
4.860389207077437,
4.839652016371997,
4.819037375245639,
4.798544559468648,
4.77817284909325,
4.757921528427404,
4.7377898860091605,
4.71777721458139,
4.697882811066787,
4.678105976543086,
4.658446016218462,
4.638902239407098,
4.619473959504905,
4.6001604939653955,
4.580961164275698,
4.561875295932717,
4.542902218419438,
4.524041265181368,
4.50529177360312,
4.486653084985133,
4.468124544520532,
4.449705501272121,
4.431395308149517,
4.413193321886414,
4.3950989030179874,
4.377111415858427,
4.350289634788691,
4.323705614785403,
4.297357248221014,
4.271242446150989,
4.245359138148196,
4.219705272138757,
4.194278814239356,
4.1690777485959885,
4.144100077224143,
4.119343819850396,
4.094807013755411,
4.070487713618334,
4.046383991362562,
4.022493936002886,
3.9988156534939776,
3.9753472665802323,
3.952086914646935,
3.9290327535727476,
3.906182955583503,
3.8835357091072957,
3.861089218630859,
3.8388417045572134,
3.805766252318255,
3.773130594967105,
3.7409288846761366,
3.7091553513746263,
3.6778043017148407,
3.6468701180518726,
3.616347257437043,
3.586230250624686,
3.556513701092141,
3.5271922840727745,
3.498260745601862,
3.469713901575156,
3.441546636819968,
3.4137539041786074,
3.386330723604006,
3.359272181267365,
3.33257342867767,
3.3062296818129147,
3.2802362202628728,
3.254588386383272,
3.229281584461213,
3.2043112798916815,
3.1796729983650205,
3.1553623250651954,
3.119381193285491,
3.0841177078300124,
3.04955755518995,
3.0156867073404845,
2.9824914160467824,
2.949958207283568,
2.9180738757659923,
2.8868254795895862,
2.856200334977119,
2.826186011130227,
2.796770325183742,
2.767941337260635,
2.739687345625607,
2.7119968819353266,
2.684858706583422,
2.658261804138292,
2.6321953788719346,
2.6066488503779373,
2.581611849276871,
2.557074213007345,
2.533025981701005,
2.5094573941398024,
2.486358883793905,
2.4637210749386185,
2.441534778848772,
2.4197909900689942,
2.3984808827583857,
2.377595807108089,
2.3571272858303303,
2.3370670107174556,
2.3174068392696316,
2.298138791389781,
2.2792550461444514,
2.260747938589283,
2.2426099566577986,
2.2248337381122383,
2.2074120675552167,
2.190337873500982,
2.173604225505085,
2.1572043313513047,
2.141131534294669,
2.117503198391883,
2.094581765839664,
2.072346087778903,
2.050775648073815,
2.0298505443823824,
2.0095514697930943,
1.98985969501111,
1.9707570510773238,
1.9522259126044894,
1.9342491815148066,
1.9168102712641417,
1.8998930915380803,
1.8834820334060334,
1.8675619549192304,
1.8521181671399392,
1.8371364205882013,
1.8226028920946353,
1.8085041720458177,
1.794827252012225,
1.7815595127450656,
1.7622533124282058,
1.7438135051831811,
1.7262012103958675,
1.7093792922693096,
1.6933122815412252,
1.6779663006505876,
1.6633089924179272,
1.6493094515482993,
1.6359381601267982,
1.6231669237756743,
1.610968816104998,
1.5993181124600018,
1.5826263318503955,
1.567058107952035,
1.5525380365322703,
1.5389946180371774,
1.526366021799498,
1.5145744448654386,
1.5036308059963888,
1.4879832957000314
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=B
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "B",
"line": {
"color": "green",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "B",
"showlegend": true,
"type": "scattergl",
"x": [
0,
0.00125,
0.0025,
0.0028125,
0.0031249999999999997,
0.0034374999999999996,
0.0037499999999999994,
0.004062499999999999,
0.0043749999999999995,
0.0046875,
0.005,
0.0053125,
0.005625000000000001,
0.005937500000000001,
0.006250000000000001,
0.0065625000000000015,
0.006875000000000002,
0.007187500000000002,
0.007500000000000002,
0.007812500000000002,
0.008125000000000002,
0.008437500000000002,
0.008750000000000003,
0.009062500000000003,
0.009375000000000003,
0.009687500000000003,
0.010000000000000004,
0.010312500000000004,
0.010625000000000004,
0.010937500000000005,
0.011250000000000005,
0.011562500000000005,
0.011875000000000005,
0.012187500000000006,
0.012500000000000006,
0.012812500000000006,
0.013125000000000006,
0.013437500000000007,
0.013750000000000007,
0.014062500000000007,
0.014375000000000008,
0.014687500000000008,
0.015000000000000008,
0.015312500000000008,
0.015625000000000007,
0.015937500000000007,
0.016250000000000007,
0.016562500000000008,
0.016875000000000008,
0.01718750000000001,
0.01750000000000001,
0.01781250000000001,
0.01812500000000001,
0.01843750000000001,
0.01875000000000001,
0.01906250000000001,
0.01937500000000001,
0.01968750000000001,
0.02000000000000001,
0.02031250000000001,
0.02062500000000001,
0.02093750000000001,
0.021250000000000012,
0.021562500000000012,
0.021875000000000012,
0.022187500000000013,
0.022500000000000013,
0.022812500000000013,
0.023125000000000014,
0.023437500000000014,
0.023750000000000014,
0.024062500000000014,
0.024375000000000015,
0.024687500000000015,
0.025000000000000015,
0.025312500000000016,
0.025625000000000016,
0.025937500000000016,
0.026250000000000016,
0.026562500000000017,
0.026875000000000017,
0.027187500000000017,
0.027500000000000017,
0.027812500000000018,
0.028125000000000018,
0.02843750000000002,
0.02875000000000002,
0.02906250000000002,
0.02937500000000002,
0.02968750000000002,
0.03000000000000002,
0.03031250000000002,
0.03062500000000002,
0.03093750000000002,
0.03125000000000002,
0.03156250000000002,
0.03187500000000002,
0.03218750000000002,
0.03250000000000002,
0.03281250000000002,
0.03328125000000002,
0.033750000000000016,
0.03421875000000001,
0.03468750000000001,
0.03515625000000001,
0.035625000000000004,
0.03609375,
0.0365625,
0.037031249999999995,
0.03749999999999999,
0.03796874999999999,
0.038437499999999986,
0.03890624999999998,
0.03937499999999998,
0.039843749999999976,
0.04031249999999997,
0.04078124999999997,
0.04124999999999997,
0.041718749999999964,
0.04218749999999996,
0.04265624999999996,
0.043124999999999955,
0.04359374999999995,
0.04406249999999995,
0.044531249999999946,
0.04499999999999994,
0.04546874999999994,
0.04593749999999994,
0.046406249999999934,
0.04687499999999993,
0.04734374999999993,
0.047812499999999925,
0.04828124999999992,
0.04874999999999992,
0.049218749999999915,
0.04968749999999991,
0.05015624999999991,
0.050624999999999906,
0.0510937499999999,
0.0515624999999999,
0.0520312499999999,
0.052499999999999894,
0.05296874999999989,
0.05343749999999989,
0.053906249999999885,
0.05437499999999988,
0.05484374999999988,
0.055312499999999876,
0.05578124999999987,
0.05624999999999987,
0.056718749999999866,
0.05718749999999986,
0.05765624999999986,
0.05812499999999986,
0.058593749999999854,
0.05929687499999985,
0.05999999999999985,
0.06070312499999985,
0.06140624999999985,
0.06210937499999985,
0.06281249999999985,
0.06351562499999985,
0.06421874999999985,
0.06492187499999985,
0.06562499999999985,
0.06632812499999985,
0.06703124999999985,
0.06773437499999985,
0.06843749999999985,
0.06914062499999984,
0.06984374999999984,
0.07054687499999984,
0.07124999999999984,
0.07195312499999984,
0.07265624999999984,
0.07335937499999984,
0.07406249999999984,
0.07476562499999984,
0.07546874999999983,
0.07617187499999983,
0.07722656249999983,
0.07828124999999983,
0.07933593749999983,
0.08039062499999983,
0.08144531249999983,
0.08249999999999982,
0.08355468749999982,
0.08460937499999982,
0.08566406249999982,
0.08671874999999982,
0.08777343749999982,
0.08882812499999981,
0.08988281249999981,
0.09093749999999981,
0.09199218749999981,
0.09357421874999981,
0.09515624999999982,
0.09673828124999982,
0.09832031249999983,
0.09990234374999983,
0.10148437499999984,
0.10306640624999984,
0.10464843749999984,
0.10623046874999985,
0.10860351562499986,
0.11097656249999986,
0.11334960937499987,
0.11572265624999988,
0.11809570312499988,
0.12046874999999989,
0.12402832031249988,
0.1275878906249999,
0.1311474609374999,
0.13470703124999991,
0.13826660156249992,
0.14182617187499993,
0.14538574218749994,
0.14894531249999995,
0.15250488281249996,
0.15606445312499997,
0.15962402343749998,
0.16318359375,
0.1667431640625,
0.17030273437500001,
0.17386230468750002,
0.17742187500000003,
0.18098144531250004,
0.18454101562500005,
0.18810058593750006,
0.19166015625000007,
0.19521972656250008,
0.1987792968750001,
0.2023388671875001,
0.20589843750000011,
0.20945800781250012,
0.21301757812500013,
0.21657714843750014,
0.22013671875000015,
0.22369628906250016,
0.22903564453125017,
0.23437500000000017,
0.23971435546875017,
0.24505371093750017,
0.25039306640625014,
0.2557324218750001,
0.2610717773437501,
0.26641113281250006,
0.27175048828125004,
0.27708984375,
0.28242919921875,
0.28776855468749996,
0.29310791015624993,
0.2984472656249999,
0.3037866210937499,
0.30912597656249985,
0.3144653320312498,
0.3198046874999998,
0.32514404296874977,
0.33048339843749974,
0.3358227539062497,
0.3411621093749997,
0.34650146484374966,
0.35184082031249964,
0.3571801757812496,
0.3625195312499996,
0.36785888671874956,
0.37319824218749953,
0.3785375976562495,
0.3838769531249995,
0.38921630859374945,
0.3945556640624994,
0.3998950195312494,
0.4079040527343744,
0.4159130859374994,
0.42392211914062444,
0.43193115234374946,
0.4399401855468745,
0.4479492187499995,
0.4559582519531245,
0.4639672851562495,
0.47197631835937454,
0.47998535156249955,
0.48799438476562457,
0.4960034179687496,
0.5040124511718745,
0.5120214843749995,
0.5200305175781245,
0.5280395507812494,
0.5360485839843744,
0.5440576171874993,
0.5520666503906243,
0.5600756835937493,
0.5680847167968742,
0.5760937499999992,
0.5841027832031241,
0.5921118164062491,
0.6001208496093741,
0.608129882812499,
0.616138916015624,
0.6241479492187489,
0.6321569824218739,
0.6401660156249989,
0.6521795654296864,
0.6641931152343739,
0.6762066650390613,
0.6882202148437488,
0.7002337646484363,
0.7122473144531238,
0.7242608642578113,
0.7362744140624988,
0.7482879638671863,
0.7603015136718738,
0.7723150634765613,
0.7843286132812488,
0.7963421630859363,
0.8083557128906238,
0.8203692626953113,
0.8323828124999988,
0.8443963623046863,
0.8564099121093738,
0.8684234619140613,
0.8804370117187488,
0.8924505615234363,
0.9044641113281238,
0.9164776611328113,
0.9284912109374988,
0.9405047607421863,
0.9525183105468737,
0.970538635253905,
0.9885589599609363,
1.0065792846679675,
1.0245996093749987,
1.04261993408203,
1.060640258789061,
1.0786605834960923,
1.0966809082031235,
1.1147012329101547,
1.1327215576171858,
1.150741882324217,
1.1687622070312482,
1.1867825317382794,
1.2048028564453106,
1.2228231811523418,
1.240843505859373,
1.2588638305664042,
1.2768841552734354,
1.2949044799804665,
1.3129248046874977,
1.330945129394529,
1.34896545410156,
1.375995941162107,
1.4030264282226539,
1.4300569152832008,
1.4570874023437477,
1.4841178894042946,
1.5111483764648415,
1.5381788635253884,
1.5652093505859352,
1.5922398376464821,
1.619270324707029,
1.646300811767576,
1.6733312988281228,
1.7003617858886697,
1.7273922729492166,
1.7544227600097635,
1.7814532470703104,
1.8084837341308573,
1.8355142211914042,
1.862544708251951,
1.889575195312498,
1.9166056823730448,
1.9436361694335917,
1.9706666564941386,
1.9976971435546855,
2.0382428741455056,
2.0787886047363258,
2.119334335327146,
2.159880065917966,
2.200425796508786,
2.2409715270996062,
2.2815172576904263,
2.3220629882812465,
2.3626087188720666,
2.4031544494628867,
2.443700180053707,
2.484245910644527,
2.524791641235347,
2.565337371826167,
2.6058831024169873,
2.6464288330078074,
2.6869745635986275,
2.7275202941894476,
2.7680660247802678,
2.808611755371088,
2.849157485961908,
2.889703216552728,
2.930248947143548,
2.9707946777343683,
3.0113404083251885,
3.0518861389160086,
3.0924318695068287,
3.132977600097649,
3.173523330688469,
3.214069061279289,
3.254614791870109,
3.2951605224609293,
3.3357062530517494,
3.3762519836425695,
3.4167977142333896,
3.4573434448242097,
3.49788917541503,
3.53843490600585,
3.57898063659667,
3.61952636718749,
3.6600720977783103,
3.7208906936645407,
3.781709289550771,
3.8425278854370015,
3.903346481323232,
3.9641650772094623,
4.024983673095693,
4.085802268981923,
4.1466208648681535,
4.207439460754384,
4.268258056640614,
4.329076652526845,
4.389895248413075,
4.4507138442993055,
4.511532440185536,
4.572351036071766,
4.633169631957997,
4.693988227844227,
4.7548068237304575,
4.815625419616688,
4.876444015502918,
4.967671909332264,
5.0588998031616095,
5.150127696990955,
5.241355590820301,
5.332583484649646,
5.423811378478992,
5.5150392723083375,
5.606267166137683,
5.697495059967029,
5.788722953796374,
5.87995084762572,
5.971178741455065,
6.108020582199084,
6.244862422943103,
6.381704263687122,
6.518546104431141,
6.65538794517516,
6.7922297859191785,
6.929071626663197,
7.134334387779226
],
"xaxis": "x",
"y": [
0,
1.875,
3.6609375,
4.086203100585938,
4.506413417053986,
4.921627870578563,
5.33190518398381,
5.737303389950974,
6.137879839129328,
6.53369120815177,
6.9247935075562195,
7.311242089613917,
7.69309165606572,
8.070396265767478,
8.443209342245556,
8.811583681163563,
9.175571457701325,
9.535224233847135,
9.890592965604306,
10.241728010113023,
10.588679132688503,
10.931495513776424,
11.270225755826631,
11.60491789008604,
11.935619383311721,
12.262377144405074,
12.585237530968046,
12.904246355782293,
13.219448893212197,
13.530889885532625,
13.838613549182329,
14.142663580943843,
14.443083164050766,
14.73991497422325,
15.033201185632574,
15.322983476795605,
15.60930303639999,
15.892200569060888,
16.17171630101005,
16.447889985718025,
16.7207609094503,
16.99036789675814,
17.25674931590491,
17.519943084228586,
17.77998667344129,
18.036917114866505,
18.290771004614765,
18.541584508698502,
18.78939336808678,
19.034232903700644,
19.276138021349727,
19.51514321661087,
19.75128257964939,
19.9845897999837,
20.215098171193898,
20.442840595575067,
20.667849588735855,
20.89015728414301,
21.109795437612526,
21.326795431747982,
21.54118828032674,
21.75300463263458,
21.962274777749396,
22.169028648774553,
22.37329582702249,
22.575105546149135,
22.774486696239773,
22.97146782784685,
23.166077155980354,
23.358342564051274,
23.548291607768736,
23.735951518991328,
23.921349209533158,
24.104511274925187,
24.285463998132357,
24.464233353227026,
24.640845009019245,
24.81532433264435,
24.98769639310842,
25.157985964792022,
25.32621753091284,
25.492415286947544,
25.65660314401349,
25.818804732210662,
25.97904340392432,
26.137342237088866,
26.293724038413337,
26.448211346568975,
26.600826435339354,
26.75159131673348,
26.900527744062316,
27.047657214979107,
27.19300097448404,
27.33658001789353,
27.478415093774654,
27.618526706845056,
27.756935120838822,
27.893660361338622,
28.028722218574607,
28.162140250190387,
28.359830550869578,
28.55390870786946,
28.744438428219066,
28.931482295862864,
29.115101791459367,
29.29535731183075,
29.472308189069555,
29.6460127093086,
29.81652813115998,
29.983910703829018,
30.148215684908877,
30.309497357861495,
30.46780904919034,
30.62320314531042,
30.77573110912092,
30.92544349628566,
31.07238997122656,
31.216619322835164,
31.35817947990717,
31.497117526304898,
31.633479715852424,
31.767311486968165,
31.89865747703949,
32.02756153654393,
32.15406674292145,
32.278215414202165,
32.40004912239382,
32.51960870663326,
32.63693428610605,
32.75206527273837,
32.86504038366513,
32.97589765347829,
33.084674446259285,
33.1914074673993,
33.2961327752112,
33.39888579233672,
33.49970131695261,
33.59861353377916,
33.69565602489471,
33.79086178035953,
33.88426320865233,
33.97589214692288,
34.065779871063846,
34.15395710560505,
34.240454033433366,
34.325300305341194,
34.40852504940668,
34.490156880208495,
34.570223907878265,
34.64875374699331,
34.72577352531269,
34.80130989235925,
34.87538902785029,
34.94803664997972,
35.01927802355414,
35.1240679402012,
35.22580450048185,
35.32456854252967,
35.42043876682677,
35.51349179273015,
35.60380221350325,
35.691442649892345,
35.776483802286194,
35.85899450149641,
35.93904175819507,
36.01669081104495,
36.0920051735572,
36.16504667970976,
36.23587552835966,
36.30455032648077,
36.37112813125833,
36.435664491070256,
36.49821348538485,
36.55882776360346,
36.61755858287601,
36.674455844916565,
36.729568131845404,
36.782942741083275,
36.83462571932295,
36.88466189560249,
36.95731142245461,
37.02644944427066,
37.09221557090486,
37.1547438744482,
37.21416310888267,
37.27059692102297,
37.32416405309122,
37.37497853725665,
37.42314988245885,
37.46878325382077,
37.51197964494532,
37.55283604337787,
37.59144558950572,
37.62789772915483,
37.6622783601339,
37.71086577938162,
37.755156149167064,
37.79540622611957,
37.83185748972529,
37.86473705127457,
37.894258508729514,
37.92062275072937,
37.94401871275987,
37.964624088332634,
37.991596952610806,
38.013020665092135,
38.029394131681855,
38.04117172877335,
38.04876727732274,
38.05255766225416,
38.05305036214574,
38.04645711341405,
38.033738794290514,
38.015727630039194,
37.99314441558984,
37.96661343260069,
37.93667536959462,
37.903798512494376,
37.86838843709625,
37.83079640402559,
37.791326629871136,
37.750242584942555,
37.70777244795579,
37.66411383050716,
37.61943786908854,
37.573892769310056,
37.52760687566249,
37.48069133033481,
37.43324237609933,
37.385343350912834,
37.337066415503024,
37.28847404968539,
37.23962034837007,
37.190552144074026,
37.14130997916401,
37.0919289479466,
37.042439426028785,
36.992867702039966,
36.94323652478614,
36.86873010334279,
36.79417251029523,
36.71961345098013,
36.645092546193645,
36.57064135746877,
36.49628500567028,
36.42204346457037,
36.34793259466982,
36.27396496942411,
36.2001505355602,
36.126497140799074,
36.053010955609366,
35.97969681027085,
35.90655846425398,
35.83359882150647,
35.760820102509015,
35.68822398178106,
35.61581169777424,
35.543584140698314,
35.471541922710585,
35.39968543401048,
35.32801488766945,
35.25653035545822,
35.185231796479144,
35.11411908004842,
35.043192003982796,
34.97245030921359,
34.90189369146551,
34.83152181058962,
34.7613342980216,
34.69133076274166,
34.621510796037,
34.55187397530738,
34.44769281300491,
34.343921759695455,
34.240559302228725,
34.13760390251718,
34.03505400694111,
33.9329080529134,
33.83116447345951,
33.729821700410575,
33.62887816662732,
33.5283323075468,
33.42818256225593,
33.32842737423439,
33.229065191866475,
33.1300944687916,
33.03151366414195,
32.93332124270145,
32.83551567500969,
32.73809543742738,
32.641059012175084,
32.54440488735316,
32.448131556948695,
32.35223752083333,
32.25672128475478,
32.161581360323964,
32.066816264999034,
31.972424522067396,
31.87840466062623,
31.784755215562054,
31.69147472752962,
31.59856174293039,
31.459741349370894,
31.321741335936334,
31.18455685447352,
31.048183085476808,
30.912615237920313,
30.777848549090447,
30.643878284419035,
30.51069973731726,
30.37830822901043,
30.24669910837369,
30.11586775176867,
29.98580956288105,
29.856519972559127,
29.727994438653273,
29.600228445856377,
29.47321750554522,
29.346957155622782,
29.221442960361475,
29.09667051024732,
28.97263542182503,
28.849333337544,
28.72675992560524,
28.60491087980918,
28.483781919404386,
28.363368788937176,
28.243667258102118,
28.065176053339048,
27.88826707978895,
27.71292631179933,
27.539139848047768,
27.366893910439806,
27.196174843016582,
27.026969110872177,
26.85926329908053,
26.693044111631895,
26.52829837037869,
26.365013013990726,
26.203175096919683,
26.042771788372754,
25.8837903712954,
25.726218241363124,
25.57004290598218,
25.41525198329913,
25.26183320121919,
25.109774396433288,
24.959063513453724,
24.809688603658394,
24.66163782434348,
24.441530244505074,
24.224349372597413,
24.010056293016962,
23.798612607609908,
23.589980428791772,
23.384122372758522,
23.18100155278794,
22.98058157263007,
22.782826519985544,
22.58770096007064,
22.39516992926788,
22.205198928861066,
22.01775391885363,
21.83280131186916,
21.65030796713305,
21.47024118453414,
21.292568698765365,
21.117258673542274,
20.944279695898445,
20.77360077055674,
20.60519131437541,
20.439021150868058,
20.27506050479644,
20.113279996835193,
19.873835959043653,
19.6391676502267,
19.40917981811809,
19.183779110265046,
18.962874036136338,
18.74637492998612,
18.53419391445846,
18.326244864917754,
18.122443374490608,
17.922706719804935,
17.726953827412395,
17.535105240880554,
17.34708308854137,
17.16281105188297,
16.9822143345718,
16.805219632092715,
16.63175510199451,
16.46175033472897,
16.295136325071493,
16.131845444111782,
15.971811411803131,
15.814969270059247,
15.661255356387658,
15.51060727804899,
15.362963886731658,
15.218265253731651,
15.076452645627379,
14.937468500439671,
14.801256404267269,
14.667761068388339,
14.53692830681867,
14.4087050143175,
14.283039144832008,
14.159879690371731,
14.039176660304333,
13.920881061064323,
13.804944876266482,
13.691321047215936,
13.579963453806943,
13.470826895802668,
13.363867074488333,
13.206627324791262,
13.054091812682511,
12.90611979861416,
12.76257475362927,
12.623324233391052,
12.48823975598081,
12.35719668335184,
12.23007410632998,
12.106754733054638,
11.987124780757446,
11.871073870778547,
11.758494926723886,
11.649284075669197,
11.543340552319988,
11.440566606038496,
11.340867410652598,
11.244150976962484,
11.150328067865615,
11.059312116020049,
10.971019143972221,
10.842541958023167,
10.719830367687743,
10.60262563369859,
10.490680628084286,
10.383759313078851,
10.281636243474766,
10.184096091165577,
10.090933191372997,
10.001951108392555,
9.916962222881523,
9.83578733267357,
9.758255283648136,
9.647176239654135,
9.543574480383842,
9.446946465866878,
9.356823629195162,
9.272764477334546,
9.194375760178556,
9.121213891980563,
9.019168479129675
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=S
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "S",
"line": {
"color": "red",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "S",
"showlegend": true,
"type": "scattergl",
"x": [
0,
0.00125,
0.0025,
0.0028125,
0.0031249999999999997,
0.0034374999999999996,
0.0037499999999999994,
0.004062499999999999,
0.0043749999999999995,
0.0046875,
0.005,
0.0053125,
0.005625000000000001,
0.005937500000000001,
0.006250000000000001,
0.0065625000000000015,
0.006875000000000002,
0.007187500000000002,
0.007500000000000002,
0.007812500000000002,
0.008125000000000002,
0.008437500000000002,
0.008750000000000003,
0.009062500000000003,
0.009375000000000003,
0.009687500000000003,
0.010000000000000004,
0.010312500000000004,
0.010625000000000004,
0.010937500000000005,
0.011250000000000005,
0.011562500000000005,
0.011875000000000005,
0.012187500000000006,
0.012500000000000006,
0.012812500000000006,
0.013125000000000006,
0.013437500000000007,
0.013750000000000007,
0.014062500000000007,
0.014375000000000008,
0.014687500000000008,
0.015000000000000008,
0.015312500000000008,
0.015625000000000007,
0.015937500000000007,
0.016250000000000007,
0.016562500000000008,
0.016875000000000008,
0.01718750000000001,
0.01750000000000001,
0.01781250000000001,
0.01812500000000001,
0.01843750000000001,
0.01875000000000001,
0.01906250000000001,
0.01937500000000001,
0.01968750000000001,
0.02000000000000001,
0.02031250000000001,
0.02062500000000001,
0.02093750000000001,
0.021250000000000012,
0.021562500000000012,
0.021875000000000012,
0.022187500000000013,
0.022500000000000013,
0.022812500000000013,
0.023125000000000014,
0.023437500000000014,
0.023750000000000014,
0.024062500000000014,
0.024375000000000015,
0.024687500000000015,
0.025000000000000015,
0.025312500000000016,
0.025625000000000016,
0.025937500000000016,
0.026250000000000016,
0.026562500000000017,
0.026875000000000017,
0.027187500000000017,
0.027500000000000017,
0.027812500000000018,
0.028125000000000018,
0.02843750000000002,
0.02875000000000002,
0.02906250000000002,
0.02937500000000002,
0.02968750000000002,
0.03000000000000002,
0.03031250000000002,
0.03062500000000002,
0.03093750000000002,
0.03125000000000002,
0.03156250000000002,
0.03187500000000002,
0.03218750000000002,
0.03250000000000002,
0.03281250000000002,
0.03328125000000002,
0.033750000000000016,
0.03421875000000001,
0.03468750000000001,
0.03515625000000001,
0.035625000000000004,
0.03609375,
0.0365625,
0.037031249999999995,
0.03749999999999999,
0.03796874999999999,
0.038437499999999986,
0.03890624999999998,
0.03937499999999998,
0.039843749999999976,
0.04031249999999997,
0.04078124999999997,
0.04124999999999997,
0.041718749999999964,
0.04218749999999996,
0.04265624999999996,
0.043124999999999955,
0.04359374999999995,
0.04406249999999995,
0.044531249999999946,
0.04499999999999994,
0.04546874999999994,
0.04593749999999994,
0.046406249999999934,
0.04687499999999993,
0.04734374999999993,
0.047812499999999925,
0.04828124999999992,
0.04874999999999992,
0.049218749999999915,
0.04968749999999991,
0.05015624999999991,
0.050624999999999906,
0.0510937499999999,
0.0515624999999999,
0.0520312499999999,
0.052499999999999894,
0.05296874999999989,
0.05343749999999989,
0.053906249999999885,
0.05437499999999988,
0.05484374999999988,
0.055312499999999876,
0.05578124999999987,
0.05624999999999987,
0.056718749999999866,
0.05718749999999986,
0.05765624999999986,
0.05812499999999986,
0.058593749999999854,
0.05929687499999985,
0.05999999999999985,
0.06070312499999985,
0.06140624999999985,
0.06210937499999985,
0.06281249999999985,
0.06351562499999985,
0.06421874999999985,
0.06492187499999985,
0.06562499999999985,
0.06632812499999985,
0.06703124999999985,
0.06773437499999985,
0.06843749999999985,
0.06914062499999984,
0.06984374999999984,
0.07054687499999984,
0.07124999999999984,
0.07195312499999984,
0.07265624999999984,
0.07335937499999984,
0.07406249999999984,
0.07476562499999984,
0.07546874999999983,
0.07617187499999983,
0.07722656249999983,
0.07828124999999983,
0.07933593749999983,
0.08039062499999983,
0.08144531249999983,
0.08249999999999982,
0.08355468749999982,
0.08460937499999982,
0.08566406249999982,
0.08671874999999982,
0.08777343749999982,
0.08882812499999981,
0.08988281249999981,
0.09093749999999981,
0.09199218749999981,
0.09357421874999981,
0.09515624999999982,
0.09673828124999982,
0.09832031249999983,
0.09990234374999983,
0.10148437499999984,
0.10306640624999984,
0.10464843749999984,
0.10623046874999985,
0.10860351562499986,
0.11097656249999986,
0.11334960937499987,
0.11572265624999988,
0.11809570312499988,
0.12046874999999989,
0.12402832031249988,
0.1275878906249999,
0.1311474609374999,
0.13470703124999991,
0.13826660156249992,
0.14182617187499993,
0.14538574218749994,
0.14894531249999995,
0.15250488281249996,
0.15606445312499997,
0.15962402343749998,
0.16318359375,
0.1667431640625,
0.17030273437500001,
0.17386230468750002,
0.17742187500000003,
0.18098144531250004,
0.18454101562500005,
0.18810058593750006,
0.19166015625000007,
0.19521972656250008,
0.1987792968750001,
0.2023388671875001,
0.20589843750000011,
0.20945800781250012,
0.21301757812500013,
0.21657714843750014,
0.22013671875000015,
0.22369628906250016,
0.22903564453125017,
0.23437500000000017,
0.23971435546875017,
0.24505371093750017,
0.25039306640625014,
0.2557324218750001,
0.2610717773437501,
0.26641113281250006,
0.27175048828125004,
0.27708984375,
0.28242919921875,
0.28776855468749996,
0.29310791015624993,
0.2984472656249999,
0.3037866210937499,
0.30912597656249985,
0.3144653320312498,
0.3198046874999998,
0.32514404296874977,
0.33048339843749974,
0.3358227539062497,
0.3411621093749997,
0.34650146484374966,
0.35184082031249964,
0.3571801757812496,
0.3625195312499996,
0.36785888671874956,
0.37319824218749953,
0.3785375976562495,
0.3838769531249995,
0.38921630859374945,
0.3945556640624994,
0.3998950195312494,
0.4079040527343744,
0.4159130859374994,
0.42392211914062444,
0.43193115234374946,
0.4399401855468745,
0.4479492187499995,
0.4559582519531245,
0.4639672851562495,
0.47197631835937454,
0.47998535156249955,
0.48799438476562457,
0.4960034179687496,
0.5040124511718745,
0.5120214843749995,
0.5200305175781245,
0.5280395507812494,
0.5360485839843744,
0.5440576171874993,
0.5520666503906243,
0.5600756835937493,
0.5680847167968742,
0.5760937499999992,
0.5841027832031241,
0.5921118164062491,
0.6001208496093741,
0.608129882812499,
0.616138916015624,
0.6241479492187489,
0.6321569824218739,
0.6401660156249989,
0.6521795654296864,
0.6641931152343739,
0.6762066650390613,
0.6882202148437488,
0.7002337646484363,
0.7122473144531238,
0.7242608642578113,
0.7362744140624988,
0.7482879638671863,
0.7603015136718738,
0.7723150634765613,
0.7843286132812488,
0.7963421630859363,
0.8083557128906238,
0.8203692626953113,
0.8323828124999988,
0.8443963623046863,
0.8564099121093738,
0.8684234619140613,
0.8804370117187488,
0.8924505615234363,
0.9044641113281238,
0.9164776611328113,
0.9284912109374988,
0.9405047607421863,
0.9525183105468737,
0.970538635253905,
0.9885589599609363,
1.0065792846679675,
1.0245996093749987,
1.04261993408203,
1.060640258789061,
1.0786605834960923,
1.0966809082031235,
1.1147012329101547,
1.1327215576171858,
1.150741882324217,
1.1687622070312482,
1.1867825317382794,
1.2048028564453106,
1.2228231811523418,
1.240843505859373,
1.2588638305664042,
1.2768841552734354,
1.2949044799804665,
1.3129248046874977,
1.330945129394529,
1.34896545410156,
1.375995941162107,
1.4030264282226539,
1.4300569152832008,
1.4570874023437477,
1.4841178894042946,
1.5111483764648415,
1.5381788635253884,
1.5652093505859352,
1.5922398376464821,
1.619270324707029,
1.646300811767576,
1.6733312988281228,
1.7003617858886697,
1.7273922729492166,
1.7544227600097635,
1.7814532470703104,
1.8084837341308573,
1.8355142211914042,
1.862544708251951,
1.889575195312498,
1.9166056823730448,
1.9436361694335917,
1.9706666564941386,
1.9976971435546855,
2.0382428741455056,
2.0787886047363258,
2.119334335327146,
2.159880065917966,
2.200425796508786,
2.2409715270996062,
2.2815172576904263,
2.3220629882812465,
2.3626087188720666,
2.4031544494628867,
2.443700180053707,
2.484245910644527,
2.524791641235347,
2.565337371826167,
2.6058831024169873,
2.6464288330078074,
2.6869745635986275,
2.7275202941894476,
2.7680660247802678,
2.808611755371088,
2.849157485961908,
2.889703216552728,
2.930248947143548,
2.9707946777343683,
3.0113404083251885,
3.0518861389160086,
3.0924318695068287,
3.132977600097649,
3.173523330688469,
3.214069061279289,
3.254614791870109,
3.2951605224609293,
3.3357062530517494,
3.3762519836425695,
3.4167977142333896,
3.4573434448242097,
3.49788917541503,
3.53843490600585,
3.57898063659667,
3.61952636718749,
3.6600720977783103,
3.7208906936645407,
3.781709289550771,
3.8425278854370015,
3.903346481323232,
3.9641650772094623,
4.024983673095693,
4.085802268981923,
4.1466208648681535,
4.207439460754384,
4.268258056640614,
4.329076652526845,
4.389895248413075,
4.4507138442993055,
4.511532440185536,
4.572351036071766,
4.633169631957997,
4.693988227844227,
4.7548068237304575,
4.815625419616688,
4.876444015502918,
4.967671909332264,
5.0588998031616095,
5.150127696990955,
5.241355590820301,
5.332583484649646,
5.423811378478992,
5.5150392723083375,
5.606267166137683,
5.697495059967029,
5.788722953796374,
5.87995084762572,
5.971178741455065,
6.108020582199084,
6.244862422943103,
6.381704263687122,
6.518546104431141,
6.65538794517516,
6.7922297859191785,
6.929071626663197,
7.134334387779226
],
"xaxis": "x",
"y": [
0,
0.1875,
0.3672421875,
0.4103292927246094,
0.45297597081548313,
0.49518738776526755,
0.5369686488547025,
0.5783247993661397,
0.6192608252886747,
0.6597816540149926,
0.6998921550300232,
0.7395971405915039,
0.7789013664025428,
0.8178095322762787,
0.856326282792729,
0.8944562079479184,
0.9322038437953779,
0.9695736730801061,
1.006570125865078,
1.0431975801503923,
1.0794603624851407,
1.1153627485720876,
1.150908963865243,
1.186103184160411,
1.2209495361788,
1.2554520981437722,
1.2896149003508157,
1.3234419257308183,
1.3569371104067207,
1.3901043442436278,
1.422947471392455,
1.455470290827185,
1.4876765568758112,
1.5195699797450402,
1.5511542260388278,
1.5824329192708222,
1.6134096403707827,
1.6440879281850485,
1.6744712799711245,
1.7045631518864557,
1.734366959471455,
1.7638860781268575,
1.7931238435854613,
1.822083552378327,
1.850768462295496,
1.8791817928412962,
1.9073267256842938,
1.9352064051019608,
1.9628239384201134,
1.9901823964471874,
2.0172848139034096,
2.0441341898449252,
2.0707334880829404,
2.0970856375979388,
2.1231935329490312,
2.1490600346784925,
2.174687969711546,
2.2000801317514482,
2.22523928166993,
2.25016814789305,
2.274869426782515,
2.2993457830125132,
2.323599849942125,
2.3476342299833535,
2.371451494964831,
2.39505418649125,
2.418444816298572,
2.4416258666050576,
2.464599790458178,
2.4873690120774397,
2.5099359271931916,
2.53230290338144,
2.5544722803947346,
2.576446370489165,
2.598227458747511,
2.6198178033986004,
2.6412196361329077,
2.6624351624144484,
2.6834665617890052,
2.704315988188733,
2.7249855702331827,
2.7454774115267884,
2.7657935909528586,
2.78593616296411,
2.8059071578697914,
2.825708582119426,
2.8453424185832272,
2.8648106268292124,
2.884115143397063,
2.9032578820687664,
2.9222407341360745,
2.941065568664822,
2.959734232756135,
2.978248551804574,
2.996610329753238,
3.0148213493458758,
3.0328833723760296,
3.0507981399332538,
3.0685673726464384,
3.0861927709242734,
3.112417637327194,
3.1383263937357926,
3.163924579074454,
3.189217634630803,
3.2142109057769233,
3.2389096436602265,
3.2633190068645206,
3.2874440630417916,
3.311289790515219,
3.334861079853933,
3.3581627354200108,
3.3811994768882006,
3.403975940738855,
3.4264966817245446,
3.448766174310821,
3.4707888140915744,
3.492568919179446,
3.514110731571726,
3.5354184184921738,
3.556496073709183,
3.577347718830708,
3.5979773045763643,
3.618388712027103,
3.6385857538528517,
3.658572175518519,
3.6783516564687355,
3.6979278112917076,
3.717304190862558,
3.736484283466508,
3.755471515902259,
3.7742692545659255,
3.7928808065158552,
3.811309420518683,
3.8295582880769397,
3.8476305444385477,
3.8655292695885195,
3.8832574892231753,
3.900818175707183,
3.9182142490137317,
3.9354485776481254,
3.9525239795551,
3.9694432230101415,
3.9862090274950948,
4.002824064558332,
4.019290958659764,
4.035612288000945,
4.051790585340558,
4.067828338795507,
4.083727992627901,
4.09949194801816,
4.115122563824487,
4.1306221573289585,
4.145993004970459,
4.161237343064697,
4.176357368511522,
4.19885417497891,
4.221080904208024,
4.2430445855564285,
4.26475206254195,
4.286209997756853,
4.3074248776520765,
4.328403017194961,
4.349150564403802,
4.369673504762505,
4.389977665518499,
4.410068719866996,
4.429952191024613,
4.449633456195267,
4.469117750431203,
4.488410170391926,
4.50751567800373,
4.526439104022458,
4.545185151502058,
4.56375839917141,
4.582163304721865,
4.60040420800785,
4.618485334162838,
4.636410796632918,
4.65418460013016,
4.671810643507866,
4.69803376208578,
4.723941275714475,
4.749545326895599,
4.774857576696206,
4.799889223844652,
4.824651023069055,
4.849153302708363,
4.873405981624883,
4.897418585445984,
4.9212002621615705,
4.944759797102883,
4.968105627327176,
4.991245855431829,
5.014188262820515,
5.036940322443186,
5.070793655324475,
5.104250360964285,
5.137332778739954,
5.170061919878847,
5.202457546478753,
5.234538245826789,
5.266321500296547,
5.297823753086552,
5.329060470047477,
5.375539061722464,
5.421483712565737,
5.466937855675816,
5.511941052856708,
5.556529340108732,
5.600735542285761,
5.6665165683630985,
5.731566461259307,
5.795968957583348,
5.859796608997625,
5.9231122794880084,
5.985970442196456,
6.048418302648519,
6.110496771616005,
6.172241307743934,
6.233682647376258,
6.294847436680899,
6.355758779153149,
6.416436709825621,
6.476898605996451,
6.537159542973938,
6.597232602198237,
6.657129138115283,
6.716859009324753,
6.776430778784639,
6.835851887214792,
6.895128803287238,
6.954267153710803,
7.01327183590156,
7.0721471155703135,
7.130896711246255,
7.189523867485624,
7.248031418280107,
7.306421841976903,
7.3646973088468,
7.4519409290032685,
7.538933941394881,
7.625681313752006,
7.712187135395744,
7.798454793312201,
7.884487112871353,
7.970286470289965,
8.055854882512447,
8.141194079044189,
8.226305559361357,
8.311190638793455,
8.395850485193332,
8.480286148244558,
8.564498582884594,
8.648488668025333,
8.732257221515285,
8.815805012098124,
8.899132768970714,
8.982241189422648,
9.065130944942544,
9.147802686098961,
9.230257046442008,
9.312494645622275,
9.39451609188427,
9.476321984059934,
9.557912913162648,
9.639289463661916,
9.720452214502885,
9.801401739921902,
9.882138610099082,
9.962663391680604,
10.0429766481969,
10.123078940397633,
10.24291676958145,
10.362282435955265,
10.481177808649084,
10.599604746742758,
10.71756510011468,
10.835060710043434,
10.95209340963677,
11.068665024139886,
11.184777371159337,
11.300432260827948,
11.415631495928473,
11.53037687198838,
11.64467017735445,
11.758513193253208,
11.871907693841438,
11.984855446249743,
12.097358210621188,
12.209417740146502,
12.321035781096823,
12.432214072854697,
12.542954347943843,
12.653258332057995,
12.763127744089093,
12.872564296154973,
12.981569693626673,
13.090145635155459,
13.198293812699594,
13.306015911550931,
13.413313610361314,
13.520188581168851,
13.679869443551631,
13.838606647691629,
13.9964057702808,
14.153272355054504,
14.309211912986395,
14.464229922482108,
14.618331829571762,
14.771523048101308,
14.923808959922743,
15.075194915083184,
15.225686232012832,
15.375288197711814,
15.52400606793593,
15.671845067381296,
15.81881038986789,
15.964907198522027,
16.110140625957744,
16.25451577445712,
16.39803771614952,
16.5407114931898,
16.68254211793543,
16.8235345731226,
16.963693812041264,
17.10302475870916,
17.241532308044796,
17.379221326039417,
17.584534311872225,
17.78802730542561,
17.989716439979624,
18.18961770580121,
18.387746951411966,
18.58411988484463,
18.778752074888434,
18.97165895232345,
19.16285581114393,
19.352357809770886,
19.540179972253835,
19.726337189461955,
19.910844220264657,
20.09371569270169,
20.274966105142873,
20.45460982743756,
20.63266110205391,
20.809134045208037,
20.984042647983184,
21.157400777438955,
21.329222177710722,
21.499520471099284,
21.752703503176647,
22.00252003243546,
22.24901482230688,
22.492232041015445,
22.732215269493366,
22.969007509189584,
23.202651189774997,
23.433188176745226,
23.660659778922295,
23.885106755856565,
24.106569325130238,
24.32508716956376,
24.540699444326382,
24.75344478395221,
24.963361309262925,
25.17048663419848,
25.374857872556948,
25.576511644644793,
25.775484083838663,
25.971810843059966,
26.165527101163352,
26.356667569240237,
26.54526649683852,
26.73135767809959,
27.006782847670838,
27.27671464194327,
27.54126262669194,
27.800534182394447,
28.054634547816857,
28.30366686273029,
28.547732209775525,
28.78692965549264,
29.021356290532253,
29.251107269064818,
29.476275847403844,
29.696953421858794,
29.913229565833007,
30.12519206618169,
30.332926958844762,
30.536518563768976,
30.736049519133537,
30.931600814893077,
31.123251825651618,
31.311080342880853,
31.495162606495846,
31.67557333580093,
31.852385759818418,
32.02567164701237,
32.19550133441955,
32.36194375619934,
32.52506647161422,
32.68493569245222,
32.841616309902385,
32.99517192089419,
33.14566485391168,
33.293156194292706,
33.43770580902353,
33.57937237103897,
33.718213383037856,
33.854285200823426,
33.987643056178285,
34.11834107928307,
34.24643232068796,
34.37196877284602,
34.49500139121699,
34.67586947681684,
34.851326421477815,
35.021534113606926,
35.1866495982969,
35.346825222226556,
35.502208774226084,
35.65294362163704,
35.799168842592685,
35.94101935434086,
36.07862603772774,
36.2121158579573,
36.341611981738026,
36.46723389092476,
36.58909749276077,
36.70731522682156,
36.82199616875919,
36.93324613094287,
37.041167760088555,
37.145860631967714,
37.2474213432827,
37.39520472954862,
37.53635612712907,
37.671173155905535,
37.7999400796464,
37.92292840537991,
38.04039745587463,
38.15259491641648,
38.25975735707869,
38.362110731480634,
38.45987085334279,
38.553243851221424,
38.642426603891856,
38.770197428495464,
38.88936741166412,
39.000515497600844,
39.10418175276765,
39.20086950086595,
39.291049794955995,
39.37515530202304,
39.492848225170285
],
"yaxis": "y"
}
],
"layout": {
"autosize": true,
"legend": {
"title": {
"text": "Chemical"
},
"tracegroupgap": 0
},
"template": {
"data": {
"bar": [
{
"error_x": {
"color": "#2a3f5f"
},
"error_y": {
"color": "#2a3f5f"
},
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "bar"
}
],
"barpolar": [
{
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "barpolar"
}
],
"carpet": [
{
"aaxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"baxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"type": "carpet"
}
],
"choropleth": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "choropleth"
}
],
"contour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "contour"
}
],
"contourcarpet": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "contourcarpet"
}
],
"heatmap": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmap"
}
],
"heatmapgl": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmapgl"
}
],
"histogram": [
{
"marker": {
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "histogram"
}
],
"histogram2d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2d"
}
],
"histogram2dcontour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2dcontour"
}
],
"mesh3d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "mesh3d"
}
],
"parcoords": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "parcoords"
}
],
"pie": [
{
"automargin": true,
"type": "pie"
}
],
"scatter": [
{
"fillpattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
},
"type": "scatter"
}
],
"scatter3d": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatter3d"
}
],
"scattercarpet": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattercarpet"
}
],
"scattergeo": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergeo"
}
],
"scattergl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergl"
}
],
"scattermapbox": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattermapbox"
}
],
"scatterpolar": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolar"
}
],
"scatterpolargl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolargl"
}
],
"scatterternary": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterternary"
}
],
"surface": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "surface"
}
],
"table": [
{
"cells": {
"fill": {
"color": "#EBF0F8"
},
"line": {
"color": "white"
}
},
"header": {
"fill": {
"color": "#C8D4E3"
},
"line": {
"color": "white"
}
},
"type": "table"
}
]
},
"layout": {
"annotationdefaults": {
"arrowcolor": "#2a3f5f",
"arrowhead": 0,
"arrowwidth": 1
},
"autotypenumbers": "strict",
"coloraxis": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"colorscale": {
"diverging": [
[
0,
"#8e0152"
],
[
0.1,
"#c51b7d"
],
[
0.2,
"#de77ae"
],
[
0.3,
"#f1b6da"
],
[
0.4,
"#fde0ef"
],
[
0.5,
"#f7f7f7"
],
[
0.6,
"#e6f5d0"
],
[
0.7,
"#b8e186"
],
[
0.8,
"#7fbc41"
],
[
0.9,
"#4d9221"
],
[
1,
"#276419"
]
],
"sequential": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"sequentialminus": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
]
},
"colorway": [
"#636efa",
"#EF553B",
"#00cc96",
"#ab63fa",
"#FFA15A",
"#19d3f3",
"#FF6692",
"#B6E880",
"#FF97FF",
"#FECB52"
],
"font": {
"color": "#2a3f5f"
},
"geo": {
"bgcolor": "white",
"lakecolor": "white",
"landcolor": "#E5ECF6",
"showlakes": true,
"showland": true,
"subunitcolor": "white"
},
"hoverlabel": {
"align": "left"
},
"hovermode": "closest",
"mapbox": {
"style": "light"
},
"paper_bgcolor": "white",
"plot_bgcolor": "#E5ECF6",
"polar": {
"angularaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"radialaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"scene": {
"xaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"yaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"zaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
}
},
"shapedefaults": {
"line": {
"color": "#2a3f5f"
}
},
"ternary": {
"aaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"baxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"caxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"title": {
"x": 0.05
},
"xaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
},
"yaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
}
}
},
"title": {
"text": "Coupled reactions A <-> B and A <-> S (slow but with energetic advantage)"
},
"xaxis": {
"anchor": "y",
"autorange": true,
"domain": [
0,
1
],
"range": [
0,
7.134334387779226
],
"title": {
"text": "SYSTEM TIME"
},
"type": "linear"
},
"yaxis": {
"anchor": "x",
"autorange": true,
"domain": [
0,
1
],
"range": [
-2.7777777777777777,
52.77777777777778
],
"title": {
"text": "Concentration"
},
"type": "linear"
}
}
},
"image/png": "iVBORw0KGgoAAAANSUhEUgAAAu8AAAFoCAYAAADn3hWsAAAgAElEQVR4Xu29C9gdRZmo++XyJxDIn5BAkEAgBnWMEfAygmBA2fpo9AwYPDMBt6JRQGEL+xlgjyjxMMgIXuZAnB2Vi4BBxrMB2e4AezRy9AgSMeKMAiHGC4ZwCV4gMRcC5H7W10n91N/pXqu7v1q1Vv/r7efJk2Struqqt2p1v139VfWwnY1N2CAAAQhAAAIQgAAEIACBricwDHnv+jaigBCAAAQgAAEIQAACEEgIIO90BAhAAAIQgAAEIAABCNSEAPJek4aimBCAAAQgAAEIQAACEEDe6QMQgAAEIAABCEAAAhCoCQHkvSYNRTEhAAEIQAACEIAABCCAvNMHIAABCEAAAhCAAAQgUBMCyHtNGopiQgACEIAABCAAAQhAAHmnD0AAAhCAAAQgAAEIQKAmBJD3mjQUxYQABCAAAQhAAAIQgADyTh+AAAQgAAEIQAACEIBATQgg7zVpKIoJAQhAAAIQgAAEIAAB5J0+AAEIQAACEIAABCAAgZoQQN5r0lAUEwIQgAAEIAABCEAAAsg7fQACEIAABCAAAQhAAAI1IYC816ShKCYEIAABCEAAAhCAAASQd/oABCAAAQhAAAIQgAAEakIAea9JQ1FMCEAAAhCAAAQgAAEIIO/0AQhAAAIQgAAEIAABCNSEAPJek4aimBCAAAQgAAEIQAACEEDe6QMQgAAEIAABCEAAAhCoCQHkvSYNRTEhAAEIQAACEIAABCCAvNMHIAABCEAAAhCAAAQgUBMCyHtNGopiQgACEIAABCAAAQhAAHmnD0AAAhCAAAQgAAEIQKAmBJD3mjQUxYQABCAAAQhAAAIQgADyTh+AAAQgAAEIQAACEIBATQgg7zVpKIoJAQhAAAIQgAAEIAAB5J0+AAEIQAACEIAABCAAgZoQQN5r0lAUEwIQgAAEIAABCEAAAsg7fQACEIAABCAAAQhAAAI1IYC816ShKCYEIAABCEAAAhCAAASQd/oABCAAAQhAAAIQgAAEakIAea9JQ1FMCEAAAhCAAAQgAAEIIO/0AQhAAAIQgAAEIAABCNSEAPJek4aimBCAAAQgAAEIQAACEEDe6QMQgAAEIAABCEAAAhCoCQHkPVBDffPb35cvfvV/yPzPfkLe+dY3Bcn1v112tXzv//uZLL9nYZD8yAQCQ4EAv4s4reg4v/s/HSP/9yXnFD7oB879nPzxz2vlh7ddVThNu3Z8+5wL5GWTJsi3vvKZdh2ia/L956tvkYW3LpbbrrtUZrxqateUi4LYCLTDLWwl6r7UVc9V3VeT4iWqhbzffe/P5fx//OoetbroE++XD/3du4rXto17tuMHVmdJaVb2GW+bm1xQu+Hi3qpLuHbV/ULemLU6bszvtT3ytm68caz6u3D1LCujMdvCeqzlv10lcz526R7ZlK2z6/dzT50l/3DOaaWKNdTlPaYkp8+V7lqYde2LWa5SHaJLd87rp45x2d9Mu6rZDrewlFX7ZLew8evhOHWTF1o4t0rb9fKuIyc6ipNuENdQr3vtK7piVKUdP7CqktKq0WN8P1TkXU/wDz7yaIIs9AnLXWzLCpJFrLLaXk/GWb8j99vrNoGv8rvwb8KUQcg6OWGuckMa8kY2rz9VOVdaRqyR93BnWOQ9HMt0Tsh7NbbdKu9aG3e9Dnl+r0ap/am6Wt7dRbrZY0BtrG54JIq8D+6sVQSr/d293BGclKlcP7j80UTiQ5wUfJGsOkrg51FW/tMU8uS9HX26XAtk712lb7kL9YcbT+o0vK0q9yx2+pllEME9Eagi/648rq82u8FUbkXCX6ztjryH6OXZeTDyHo5tN/XTZrWy/h7DEduVUzfLu3/NLvvEMDSndufXtfJe5GKUBcdd2N13WRfVvB9tlhS4jnrk9GnJRd9t6Ytk3g8s6zF2VvhF1n56MdenDkWE0ZXznW/964EQI18GipTDjdylueaFi/ij0i6N7nv3vf+exOrn5ZM3qpfOL0tEXFqVMD+UKquds8Ktyoiu/xj6Px76jVn6Qkh7mmkIic+Td1f/IuFC6ZFtV868J2aa502NeSLuqYbu347fheabPqHn1bfMydYJt0Xa824Eqkh8yItWK6lJn2O1Hv4AS1769PklXc+8c2iWLLQqo2Obd75In1vypDjNNe8c2exm0FIv/1yZFz7q6uKfr/7rZ/57cu1wW5k4+CLXCv9amQ67y7peZZU9XSa/Tf08/f3Sx9Lfn55D0nm1Ol7Wtcv1Y/1bQ8+yrhV517xm89zKXlfdE0/Xds4D3Pmx2aBm+pydzkvzzDq/OPb//XP/dVDYXXrfrBBLt09e/8y75ma5mrZl1v7pejS7jhc9N5Q533fjvl0r71Xil7SBdfNjqbM+Kyvvmqd/sned1O9AWSforPCGrHpl5afHLPMIqJlMFC2H/vD/9MxfBo3O5QlclgDpvrrpHW+z0dEsec/KL+sz9yP2pSlLXLI4l5FRrUe6nFWlrx3Snj6ZWI6RV68y9dXjf78xN8V/CpbVBnlzCLL6S4jfhXJKxwIXeaKXd7Juh7TnSXzZGwNXtjKillVPzSfv4ph1TkqfY7POr1np0p81+x1rOX0pLDr61+x8kXVOT0t4VpnKxpZb6pU+BxUZeXcS6iatlpGZotcKX7z8dsk6tztePtusc7EvxukbgLzBPJfG7/Nljpc1sTrvRrjVNS/vnBHiuqoTkZ28N7tRV/6vf+0rB67h+n/l7t9c5DmRinOWrKdvdPN+e9o3dYDT968y5/CsvufqmnW9z3vKWPb32Y1iXqRMXSvvZUWr1ci3fzEqK+9ZnSQtG1nHz+vkmvaXj/xuoJPnjUSXCQ9odjErWo5mwuLzK1KuMvKe92PLulA1G7X3T8R5x9c8dWu1IlDWsctKnz+CFSpMo9WPOk+Om6XLGk1x+5d5UpF1jPTFJO93Wqati/Q/vyx5AlSmbk4Sygp1q/Yq0i5F51qEmNjv+mxef807l+hv2D2mTp9fmwlnOr90W2lb/+HPa5LR1bS8FPlN5Z0v0uecoiPvWTeDRdq4ar2qyHv65q1M2EXRa0XebzDNtZloaj/Rzd3wN7vJaPU0x9U5xPGy8ih7zinSJ9I3yXnHyGq/rJth14dbPSl1+RV5WtYsGqFICJ5ySLdzCFfLqoNjXpRDkTbq5n2GjLyXkcUQ8p7ugOn/Fxkh0dGFZiebMieMZnfDGl7SbHUCf5SjyGoVRSazhWgP/eG0uri7H1e6Td1FpEoIguZZZiQ47wdeVd6zZLrVCdmVoaq8Z0lp1qhHq5NZVtn9vIvKe6jfRd7vMGv0qVndqsh71iP2MjcMjmVReU/fsPhhE/pdkWO3uvC5kewyj62bjYTl/W7dOUkZuBCrgyZNTEYUXR8qEk6Yd65K94t2y7tjULZeMeW96DUr7/yYdWPT7MYhPYjVTN6b3TT6y2OGOF7WuafINa/Z+cNyXc2qU1ZbtbrBSZfPv6aEcCKXf6tQnTw3yOLeymuyroutzmGtrmF1+b7r5b3I6Iq7u8tbW7jIY928E1Je52kl775E5XUGvfNd/YdnktjtrHqGkPei5dBHrE420hfmNIMij6zLyHuzE2P6UWXRkXf/QuLzLypCzUajy94QWEJaip5ILMdoFh5T9AmYa+8033R7FZX3ZiJR5neRFZ/tMy16U+TSdHPYTLO+4i6orUJqilz4si7OzUSgWXulRxD94+v5XOdF6GN4X9jdaHyRhQq6Rd6r1iumvJe5VhQdec+L9/b7qruhyRPIZjfy6RvDdh2vyDUv7/dnva7mnTP9vpH1xMy/YfB/93kj+Vn+VGbk3bFPDwSl27VoO+fd8Pics7ypyDms6HW1m/frWnnPi3HLg1lGFkPcZaZPGnkj763kIJSkVLlDHXQC3b0+dNaIWtHRbz+/EO2h+RU9dpG4TncSbXVD2Gz0pqjMZvVTi2Dn9Xs/zyIjq1n5NJN3l38rZnn9r9PynleusueXNLd2SLzLs+zNYZELTNF+2ypsJn0s/wKbJ2FlRt7db177ss6/0U1H291x9HyqcbWzTjy60PrzefJe9ElpiJh3/8avbL1iynsZ6Skq783CG9J9qajU+enyrsOtblKbDfiFHHlvduNRdFAs73rk1/27P1yavKDLfxqV97trl7wXHVgr087p0KIi57oy/bhIft26T9fKu/tx+bGOWRD1JOI/Sk3LctaPJ4S8p/PI+kEUvVtvFmdY9A2rzY5VpBxl4lKbibmLfU0/JvbbLv0jb1fMezomr9mJ1C9fVjyh+94qfZpPCIkPIe2+VOTFcheRvlYTqPw3XBYdec+6cXPlLTry3urGo1k7Fz1hh5D4ENKuv18dOct7aV1RZr48p5da03ZWSUh/nv79WmLe3Xlf/06f+13sbKtrQrNzTV4fKhM2U0ZI0+eVsvVKnyub3VyVkbW8/l3kWqFpi8p70XNuM5nW74rOXShzvHTYTvocX3Selz/fI821zHU1z0uaDSa59tI5dP5E1WZtZJX39Dwm/zqS9WQ7LzQufYNVxtWanZ+ZsFr06tXm/dwj2iIvaXL7pme/axH9GdBZndd/vJ5e1SAtNllCk5Wn+yw9IqqdVJfycmXKys9//FcktrPZSbdoObJGYLPibvNiof275GYnrawTsabNmumeZl/m7j69hFgZEW0WXhNC+rRPuvKUHTHPa8+qP8W8kXd3nCKhRlkx5O73WCXm3efj35CX+V20aqdWcl+Up/s9VBkxz+r3RY/r7+dPVk1fFMv2s1YjY1m/Sf8cm5XetUV6klzWuxPy5qv4N6xFzolO+vLO/+lrSp4oa3r/N1pGEP02qlKvrPNdUZF1x24mf+m+VvRaUVTe/d9xmrf2WQ2LKjJhNeu3mtfnHedWx8vjktW+Ra55zW6I0r+ZrOuqq4/f1/w6Zj3F970lb6Kyz8H/DVWNec9r+6zzrfus6LVdByCyfmt5LxHUpS3dqkqOf5Gn8FXOsd2WpqtH3tMnnzS8Vp1Z9281oujy1A7+8IqVyfrkaXl366z6xy86oz8vbistbOlYPfd4NcTIu5a7aDnSsd7KollIhD8pLi156Tq59mom4P66383WeU/Hu2b9YLNic1s9Ti0i+KGkr1tOBs3i+1uFy/h1SPNW1nqTWnXk3b/wu+MU/V0UfULSLGSoW9qnTDny2rKo7Oqxmsle1oo2rWJcXfnT54O8m50siXF5lG0vd1OZnrybF87o89PyuXWv0+drX4K0bEV+J1XqlXWuTJ/Ls9Z594WmjLwXvVaUkXfNM6vfpLm1kq40c+13r5vxij3CRYoeT/dLz4nRc5Zueeu8p89xRQY2NL+i19U0J63ju976piRUrNnkzDzPSTPT8rp31lSVd62Pz8H/HTtZd79XN9k8K54+zUT3zZv/l3Utb7bIQtkBsTLn2G7Ztxby3klYRR8jdrKMHBsCEIBASAJ5N9ghj0FeELASyAt9seZL+vgEQsSqt3raGr9W7Tsi8t6CLfLevs5HzhCAQHcSaDZK3J0lplRDmUA61FTr2iw0cyizGAp1U8nWt6SnXx7lP6UtW8+h9kS8Vf2Rd+S9VR/hewhAoAcJ5C3/2YMoqHKHCeSFfbZaza3DxebwOQTS4TW6W9EQpKwse/Fchbzz84IABCAAAQhAAAIQgEBNCCDvNWkoigkBCEAAAhCAAAQgAAHknT4AAQhAAAIQgAAEIACBmhBA3mvSUBQTAhCAAAQgAAEIQAACyDt9AAIQgAAEIAABCEAAAjUhgLzXpKEoJgQgAAEIQAACEIAABJB3+gAEIAABCEAAAhCAAARqQgB5r0lDUUwIQAACEIAABCAAAQgg7/QBCEAAAhCAAAQgAAEI1IQA8l6ThqKYEIAABCAAAQhAAAIQQN7pAxCAAAQgAAEIQAACEKgJAeS9Jg1FMSEAAQhAAAIQgAAEIIC80wcgAAEIQAACEIAABCBQEwLIe00aimJCAAIQgAAEIAABCEAAeacPQAACEIAABCAAAQhAoCYEkPeaNBTFhAAEIAABCEAAAhCAAPJOH4AABCAAAQhAAAIQgEBNCCDvNWkoigkBCEAAAhCAAAQgAAHknT4AAQhAAAIQgAAEIACBmhBA3mvSUBQTAhCAAAQgAAEIQAACyDt9AAIQgAAEIAABCEAAAjUhgLzXpKEoJgQgAAEIQAACEIAABJB3+gAEIAABCEAAAhCAAARqQgB5r0lDUUwIQAACEIAABCAAAQgg7/QBCEAAAhCAAAQgAAEI1IQA8l6ThqKYEIAABCAAAQhAAAIQQN7pAxCAAAQgAAEIQAACEKgJAeS9Jg1FMSEAAQhAAAIQgAAEIIC80wcgAAEIQAACEIAABCBQEwLIe00aimJCAAIQgAAEIAABCEAAeacPQAACEIAABCAAAQhAoCYEkPeaNBTFhAAEIAABCEAAAhCAAPJOH4AABCAAAQhAAAIQgEBNCCDvNWkoigkBCEAAAhCAAAQgAAHknT4AAQhAAAIQgAAEIACBmhBA3mvSUBQTAhCAAAQgAAEIQAACyDt9AAIQgAAEIAABCEAAAjUhgLzXpKEoJgQgAAEIQAACEIAABJB3+gAEIAABCEAAAhCAAARqQgB5r0lDUUwIQAACEIAABCAAAQgg7/QBCEAAAhCAAAQgAAEI1IQA8l6ThqKYEIAABCAAAQhAAAIQQN7pAxCAAAQgAAEIQAACEKgJAeS9Jg1FMSEAAQhAAAIQgAAEIIC80wcgAAEIQAACEIAABCBQEwLIe00aimJCAAIQgAAEIAABCEAAeacPQAACEIAABCAAAQhAoCYEkPcADfX0mhcC5NKbWYzbp0+2bd8pm17c1psAAtR6Yv9oee6FrbJ5644AufVmFgeMGy3rNm2VrdtgWLUHvGzCXvLnv2yWHTt3Vs2ip9MNHzZMJu03Wv649sWe5mCpfN/I4TK+cU15Zv1mSzY9nXavUSNkzOgRsnbjlkIcJk/cu9B+7BSWAPIegCfyXh0i8l6dnUuJvNsZIu92hsi7jSHybuOnqZF3O0Pk3c4wRg7IewDKyHt1iMh7dXbIu52dywF5t7NE3m0MkXcbP+Tdzk9zQN7DcGx3Lsh7AMLIe3WIyHt1dsi7nR3yHo4h8m5jibzb+CHvdn7IexiGMXJB3gNQRt6rQ0Teq7ND3u3skPdwDJF3G0vk3cYPebfzQ97DMIyRC/IegDLyXh0i8l6dHfJuZ4e8h2OIvNtYIu82fsi7nR/yHoZhjFyQ9wCUkffqEJH36uyQdzs75D0cQ+TdxhJ5t/FD3u38kPcwDGPkgrwHoIy8V4eIvFdnh7zb2SHv4Rgi7zaWyLuNH/Ju59et8v7eufNk4oR+ufGqi8JUMmIuy1aslNPOuUxuufoSOWL6tGBHRt6NKI/69W/lewdMMebSu8mRd3vbs1SknSGrzdgZIu82hsi7jR/ybufXKXn/6AVflJ/9YsWgCkwYP1buW7Qg+awT8r5o8RKZ94Xr5fJPnSmzZ82sDBd5r4yuvQmH/fJhWX3oK9t7kCGcO/Jub1zk3c4QebczRN5tDJF3Gz/k3c6vE/I+421zxRd1VwsV+gP3308+f/HHOiLvYWiKIO+hSAbOB3m3AUXebfw0NfJuZ4i82xki7zaGyLuNH/Ju5xdb3lXQf7fyqYER9rwauJF3/d6N0OcJvz+C74eqHD/7PJl59BGy5IFlsnbdxuRQZ59+skw5eFIywu42lyZLutNPCDT9eWe8T7KeHCy/Z2GSJfIepl8GzwV5tyFF3m38kHc7P80BebdzRN5tDJF3Gz/k3c4vtrzrqPvJ7zwuGV1vtqm8P7pqdSLbKsu6qYy/ctohA3HwKtBr1m6QOxZenny/4IbvyDU33ylOonV/lXYn5+77dHiOptU80tKdvtHQ7+d//dvJ8fW788/6u4GYdi1vXj5hWkmEmHcjSeTdBhB5t/FD3u38kPcwDJF3G0fk3cYPebfziynvTo6LxJRnxbx/+orr5Fe/fTxTtB0JFfY5J52YCL8beXc3Clkj4pqnjsxrrL3/veank06LlNXdONx214/2yIcJq2H6aJBcVN6XHjRVpvT1Bcmv1zJB3u0tTtiMnSEj73aGyLuNIfJu44e82/nVVd7d5NIsAm60Pk/efSHX0fgs6f79408noTVuFD/rOG5k3/9O9ydsJky/DJ4L8m5Dirzb+DHybufHyHsYhsi7jSPybuOHvNv5xZR3PVaZsJn0UpH+yLuT91ZyrTHv6ZH3EPKu9TjmDdMHQnj8kB3kPUy/DJ4L8m5Dirzb+CHvdn7IexiGyLuNI/Ju44e82/nFlvdWE1ZV0PNWm8kKm2kW1mIZeVcueWEzWTcOyHuYvtjWXJB3G17k3cYPebfzQ97DMETebRyRdxs/5N3OL7a8u9H39MoxTojdZNZWMe+aj1vxxR99V8E/5g2vSdZpt8i7xqprGdau2zCwMo6bsKoTVdNiryPxuhE2E6ZPtiUX5N2GFXm38UPe7fyQ9zAMkXcbR+Tdxg95t/PrhLz74u3XwB9FLyLvefn4q81UDZtxE03dqjeunK6MepNw5933DxRf4+zdSjeEzYTpl8FzQd5tSJF3Gz/k3c4PeQ/DEHm3cUTebfyQdzu/Tsl7mJL3Vi4sFWlsb+TdBhB5t/FD3u38kPcwDJF3G0fk3cYPebfzQ97DMIyRC/JupIy82wAi7zZ+yLudH/IehiHybuOIvNv4Ie92fsh7GIYxckHejZRV3m8/8GA5dvQYY069mRx5t7c767zbGbLOu50h8m5jiLzb+CHvdn7IexiGMXJB3o2UkXcbQOTdxo+Rdzs/Rt7DMETebRyRdxs/5N3OD3kPwzBGLsi7kTLybgOIvNv4Ie92fsh7GIbIu40j8m7jh7zb+SHvYRjGyAV5N1JG3m0AkXcbP+Tdzg95D8MQebdxRN5t/JB3Oz/kPQzDGLkg70bKyLsNIPJu44e82/kh72EYIu82jsi7jR/ybueHvIdhGCMX5N1IGXm3AUTebfyQdzs/5D0MQ+TdxhF5t/FD3u38kPcwDGPkgrwbKSPvNoDIu40f8m7nh7yHYYi82zgi7zZ+yLudH/IehmGMXJB3I2WV9xsOmCyz9t7HmFNvJkfe7e3OUpF2hiwVaWeIvNsYIu82fsi7nR/yPpjhjLfNlVdMPVjuWHh5GLgBc0HejTBV3udPOFDm7NtvzKk3kyPv9nZH3u0MkXc7Q+TdxhB5t/FD3u38kPeXGC644Tvyg/v+Q9au2yBf+/z5csT0aWEAB8oFeTeCRN5tAJF3Gz9NjbzbGSLvdobIu40h8m7jh7zb+SHvLzF879x58o7j3yi/XP47OXD//eTzF38sDOBAuSDvRpDIuw0g8m7jh7zb+WkOyLudI/JuY4i82/gh73Z+nZD3VVu2yKrNW8IUvkQuU0ePkqmjRmWmWLZipZx2zmVyy9WXyO8ff1quvOZWuW/RghK5t39X5N3IGHm3AUTebfyQdzs/5D0MQ+TdxhF5t/FD3u38OiHvX/7zM3L+6j+EKXyJXP5+0gEy/+CDMlO4kBkX666x7yry3RQ6g7yXaOysXZF3G0Dk3cYPebfzQ97DMETebRyRdxs/5N3OrxPyvnDNWrlp7bowhS+Ry4cnjJe5EydkpnAhM+ed8b7k+49e8MWuC51B3ks0NvJuhJWRHHm3MyXm3c6QsBk7Q+TdxhB5t/FD3u38OiHvYUodLhcXMpPOccL4sV0VOoO8G9tcR94vHX+AnNU/3phTbyZH3u3tjrzbGSLvdobIu40h8m7jh7zb+SHvIumQGUdVQ2cu/9SZMnvWzDCgjbkg71aADXm/cNwEuWDcRGNOvZkcebe3O/JuZ4i82xki7zaGyLuNH/Ju54e8ixw/+zyZc9KJ4kJmHFUNndHtxqsuCgPamAvy7gH89BXXyZ1337/HxASNf3p01epkz/SC/TryjrxX74XIe3V2LiXybmeIvNsZIu82hsi7jR/ybueHvIdhGCMX5H035UWLl8g3bvleIun+rGK921qzdsPAG7ZU5CdO6B+4+0Lebd0Uebfx09TIu50h8m5niLzbGCLvNn7Iu50f8h6GYYxckPfdlN1SQG5tT7ckkD5CufDsUwfinFTy/TU/kXdbN0XebfyQdzs/zQF5t3NE3m0MkXcbP+Tdzg95D8MwRi7Ie4OyjqZ/5LR3y+GHTR5YmF/l3V+o38l8+jOV9/82fmLyh608gbFj+mT79p3y/OZt5ROTIiGw39hRsunFbbJl6w6IVCQwsX+UbHh+m2zdBsOKCOWA8aNlzfotsmPnzqpZ9HQ6lfeJ40bJM+s29zQHS+X7Rg6X/jEjZc2G+C/9sZS7m9KOHjVC9h41XNY9t7VQsQ7cb69C+7FTWAI9L+8a5/6nZ/+ShMGkxbyovF/ysklyyYEHhm2ZHslt+DARvdRzva/e4HrR39kAiDLZGCKd1flpyhGNH/P2HfRCC0UYWujtSqvnQ37L1Tk2LskyrARD7bNs8Ql0RN41FGXtuo2ZtV1+z8JoFNIhMFXlfc4+/TJ/IvJepeEIm6lCbXAaYt7tDAmbsTMkbMbGkLAZGz9NrSPv4/fpk2fWD82nF8PXr5dh6we/0Gjkk4/vAW7EE4M/G9ZIN3zD+j33e/KJPT7re+pxGdGQ962NJ+L+lnscRt7sHbdCDtHlPT3hs0KZgyVReZ/3hesz8zv79JOTpYKyYt41jbvJ0LAZ5L16kyDv1dm5lMi7nSHybmeIvNsYIu82fu2Wd1+IVYSHpyTa/z5Llkd4oqxpdR9/0898wVZJV1nv+g1570gTRZf3blvo3qeeFSZTZLUZ5L1630Xeq7ND3u3sXA7Iu50l8m5jiLyX4+ePQjuZHjFiuOy710h5bsXvBjIbJM0N6falOT1C7Y8up78rV7r27L1j3DjZOW7wCyG3TTlsj4NtP3TwZzsb6Xb0j9tzvymH7vHZiGnTZHQj5n3j84Nj3vOOM3ni3u2pLLk2JYC8e6kH38cAACAASURBVHiy5F2/brXOO/Je/VeGvFdnh7zb2SHv4Rgi7zaWQ03efbn2pTgZld49au2L9aB/e2Ef/oh0J4XaF2IV4R0pifa/z5Ll7Z4oa1rdx9/0M1+wVdJV1mNvezUmrI4ZPULWbiw26Rd5j91Cu44XXd5VhN9x/Bv3eHtVZ6pvP2qrsJknNzwuT218XPpHj5cpYw9r/B3/x2ivZftyQN7tbAmbsTNk5N3OEHm3Mey0vDsxzhp99sNAnGT7oR+dEGx/FNrJtM6dHDlimLww+aUR5UHS3JBuX5rTI9T+6HL6O1vr1ic18l6Ptoou7+lJovXAlF/KZvJ+1QOXy5WNP/527MEnyIVHXyz6N5sI8m7vBci7nSHybmeIvNsYVpF3N7rtx2A7CR+Q7N2hIlmy3c646iy5VkLJqPTuUWtfrAf92wv78EekWwn1UJ+wauthxVL3ury7CIw0rcs/debA+36KkWzvXtHlXWPem20xV5sJgVbl/di9xsjtkw4elN3XH/yKXLrkk8lnKuobNq+T5c8+PLAPEr8LBfJu74XIu50h8m5niLyXZ+jkW0e7dXm+/bZtkvWrn5G0eDshdyPc7ZBuJ8Z+SIj7zA8DcZLth36UEezylIqnQN6Ls8rbE3lfOeh9P8pJlxRf8sAyuW/RAjvgQDlEl/dA5e6abLLkfcPm9XLMN1/dEPb1Mv/t18qc6acn5dX/X//QV+TrjT/6byf2vTwSj7zbuzLybmeIvNsZ9qq8+wKuFFW0XZiJjoi7GG/3t34WKnbbjW77wu1Gtp1w63f+Z1pGJ9udiqu297bsHJB3O1nkfU95dysTdtPgMvJu7OtZ8u7CZXR0/fZTFu9xhCyJV8H/7Mwv9VxMPPJu7ICN5Mi7nSHybmdYd3kvIuF+XHiI0W8n3xprra+66Ttggryw91gZGN3eLd4Do+K7JzUONem2975dOSDvdpKx5X3VulWif2JvU8dPFf2T3vJWHdT99GWe3bJ1RN6z1lfvtniiog2UJe/Tv35QMrKu4t4stt1JvIuL18msZx11rpzZ+NMrE1uR96I9LX8/5N3OEHm3M+w2eVfRdpMv/dFwDUlx8d8uDMUyEu4LuFJU0c4b9W424l0l5t3eakMrB+Td3p6x5f3LS78s53//fHvBS+bw92/+e5n/rvm58p7+wr37p+Rh2rZ7dHlfcMN35Jqb75Rbrr5Ejpg+LamYu9PpNjhFqKfl/aerfyx/+79mJfK94qw/FMlCdEWa83/4cdG0umlaHYV34TaFMqnpTsi7veGQdztD5N3OsN3yroI9sJ63F5qiMu6PiFcV8SIS7kbAdaQ89Og38m7vg8i7nWFseV/44EK56aGb7AUvmcOHj/qwzH3d3Fx59x2VsJkGJn1j6ZyTTtxjqUiV+tvu+lFXTQgo0hfS8u4mqp71unPl0oaAl9lU3q984IoBie+FSa3Ie5kekr0v8m5niLzbGVaR97SQJ6Piu2PCXYz4iMZkzipvmlTRdkv/+aPhGpLiJlu6kfBWq5jY6bTOAXlvzajVHsh7K0Ktv48t761LFHePvPf96GIrvtDHLdWeR4s+8p73htVuvLMp0jgq7zNGjZa7X7ZrXdkzvnuqLF5516CJqkXy8fe5bcXNctXPr0hG5HUbyhKPvJftHXvuj7zbGSLvdoa+vLuQFRVwJ+huhFxDVaoIuQq2m5iZlvF2jojbyRTLAXkvxqnZXsi7nSHyvueEVRcx0tMTVofiyPuUkX2ydPLU5Ffj4t2XfmiFTOnf87XFRX9aeZNaL3jTxaZ8ix4/1n7Iu5008m5niLwXY9hMykdv2iA7Hnus1Ci5hqpsb4Sg6Ai4CngyKu5N0kw+T74f+i+3Q96L9UHk3c6pWQ7I+y55T2/dJO5atugj70Mx5t3Ju46Uv/mb00vFu7f6GWZJ/IVHzxsyk1qR91Y9oPX3yHtrRq32QN53LXGYrDe+e7R85FO7Ysn1//p5mVhyF7KiEzfdiLmKuT9C3g2hKq36RczvkXc7bUbe7Qx7Xd7tBOPkEF3etVpDbbUZJ+8aLqNhM3lLRFqaVG8MNJRGQ2p0Gyor0yDvll6xKy3ybmc41OVdY8Y1VEUF3IWx9D3ycGkxzxopd1K+35RJ8uz4ybKtv9/eID2YA/Jub3Tk3c4QebczjJFDR+Q9RsViHUNj3p28u8mqukqMvpypHZu+pfUf7/vkoJVp6ry8JPJu7yXIu51h3eVd5bzvkYcSGVcp10mfI5c1/i4RX65ivvW1RyXLHCYj54fsGinX/+vEz1Yj5VUmrNpbbujkgLzb2xJ5tzNE3u0MY+SAvBspq7z3Dx8uKw45XC5d8klJBL6xyoyuNtPOLb0yTV1H4pF3ey9B3u0Mu13em8m5ynqrzR8x33bEkUlc+dbXHllYzFvlr98j70Uo5e+DvNv4aWrk3c4QebczjJFDNHnXVWZ0HXdd473Z1m2TAlo1gsq7bqsPfeXASjM3vOdWmTXtpFZJg3xfd4lH3u3dAHm3M+wGeU9GzBsj5Tpi7uLNkzCXAksl5o2a75oEGmfCJ/Ju64fIu40f8m7npzkg72E4tjuXaPLe7op0Kn9f3nWyqsamW1eaqVKXuko88l6ltQenQd7tDGPJuwq6iztXQXehLa1Gz93IeSLju0NakhH0LlqNBXm39UPk3cYPebfzQ97DMIyRS3R5z1vnvc4vaXIj726ZSH2zqoaxdGKrm8Qj7/ZegrzbGYaU9/QIejKSXmC1FrdCi1sy0YW1aBx6HZZKRN5t/RB5t/FD3u38kPcwDGPk0jXyXueXNGlDrThwUrLGu0q7ynunt7pIPPJu7ynIu51hWXl3MejpEJdWI+gq5DparqPmOiHUjZ7r53XfkHdbCyLvNn7Iu50f8h6GYYxcukbeP33FdbLkgWVy36IFMeod7BgubGbp+FHJGu/6YiYNm+mWrdslHnm39xTk3c4wT979UfS+5cuSkBdd1UXlPW8byoLejDTybuuHyLuNH/Ju54e8h2EYI5co8p61rntW5S7/1Jkye9bMGPUOdoyXL/+1rNqyRS59cZlcevcH27LGe4jCdqvEI+/21kXe7QwnrVohz//m9zLsoYeSpRZ3SXr+Ki7+BFGVdf2j4S5DYQS9Kk3kvSq5XemQdxs/5N3OD3kPwzBGLlHk3a9IXsx7jMq24xhO3i9cv0SuvPcT0s413kOUv9skHnm3tyryXpyhL+ZlJN0Pc6lLDHpxKmH2RN5tHJF3Gz/k3c4PeX+Jobqqv71i6sFyx8LLw0AOkEt0eQ9Q5q7Kwsn7Wc/8m3z9pxcn67vrOu/dvmVJ/KxpJ8sFb7o4Cf2JtSHvdtLI+54M0zHpGpveLNxl51Gvk21j+2VLYwR983HHJ+ufI+nl+ibyXo5Xem/k3cYPebfzQ953MTx+9nky8+gj5PMXf2wA6nvnzkPew3Sx7sjFyfuslV+TxY9cG+UFTSFrnpZ4zfvYg0+QC4++OPm73Rvybifc6/Luj6CP+sl9TSXdhbvoSLov6RMPnSTrNm2Vrdt22BukR3NA3m0Nj7zb+CHvdn7Iu8iyFSvltHMuk1uuvkSOmD4tDNQ25BJ95N2ByatL3V7S9Prf/E4efP4FmbH8c7L899+W+W+/Ngmdqdum69Pf8PBX5dYVN8uGzbsm48WQeOTd3lN6Rd6zRtNH/+THmQB9SXerujQbSS+72oy91YZeDsi7rU2Rdxs/5N3OryPyvmqViP6JvU2dKqJ/MjYdeZ8wvr+rRtrTxYwu7+5xxDFveI1cec2tA6vL6COJdxz/RjnvjPfFbkLT8U58dKXcs/E56f/3C2TD0z+UmG9XNRU8J7GK+/UPfUW+3vjjJF7DaM448hNyauOmJPT69ci7vRWHorz7oj76/vuaTiD1J4tWnTiKvNv7IfJuY4i82/gh73Z+HZH3L39Z5PzzwxS+TC5///ci8+fnpkjHvJ99+sld5afR5d1NWD38sMnyXz49f0DedUUaX+bLtEEn93XyLvfMEdnwG7n7tKUyY//6r9nsJP62X/9r8tZY3VTczzrqXDmz8SeUxCPv9t5bd3mvIuoa8qKhL6Hi0pF3ez9E3m0MkXcbP+Tdzq8j8r5wochNN4UpfJlcPvxhkblzC6XQl4hec/Od0k0rInZM3nVJSBV5FyZT15c0Dcj7//tukReeTtZ4jznhs1DPM+6UFRc/a9pJDYn/hDkuHnk3Nk4jed3kXUNddAJpqxH1zW85YdDLjPT/7dqQdztZ5N3GEHm38UPe7fw6Iu9hit32XDRqZM5JJ3bN6Ht0edfwmNe86rBkFq//77q+pGlA3r/XWJ9+68bk7aqhRqXb3htLHkAl/vqHviqLV941kFLj4lXiVearbMh7FWqD03SzvLvJpCrqibDnxKg7Ud864wjZesRR0ddLR97t/RB5tzFE3m38kHc7P+RdRAeSv3HL9wbFu3fj4HJ0eU93Lz+uqNtn92b9NE557HFZtK4xwfPOo5KvV5/7fJhfUBfnkhUXXzWkBnm3N3S3yLsLf9EVX5qt+pK81EgFvSHqyaovbRxRL0oXeS9KKn8/5N3GEHm38UPe7fyQ910MdWD50VWrBwHttsVUOi7vYbpb53L5yONPysKnlov84N1JuIyGzfTKphKvq9N8/7H/LToq77Yyq9Qg7/be0il511H1vmUPyaj7lyR/Z72R1L11NHSMup3a4ByQdztR5N3GEHm38UPe7fyQ9zAMY+QSXd6H2htWe1ne/Q6at178u17+N01XqUHe7T/zWPLux6qPaoS/6Eh7enPhLyrrbuUXew3bnwPybmeMvNsYIu82fsi7nR/yHoZhjFyQdyPlRN4fWypy75xklRldbaaXt7yQGl1mUkU+/eIn5N3eW9oh7+kQmKxY9fSoejeEv1SlibxXJfdSOuTdxhB5t/FD3u38kPcwDGPkEl3e67qee15jnL/6afnyr74ncv+ZiZjefsriGO1Wi2MUGY1H3u1NGULeVdZ1NL3ZxFIXq/7iu/+mVqPqRQgj70UoNd8HebcxRN5t/JB3Oz/kPQzDGLlEl3d9w6q/vnuMSrbzGJ/945/k0p9dJ/LgJcmbVfUNq2yDCSx/9mH5fmOFGv/FTzrBVUfj3zd9dnLTs+nFbWCrSKCKvPuyPmrJjzPj1f0QmC2NSaX61tKhuiHv9pZF3m0MkXcbP+Tdzg95D8MwRi7R5T391qp0JbttRm+rRkDeWxF66XsNqbl/93KT/gTXmVPeKu847P9oyxtci5euvnsWkXcXBrPX9/63NJP1LW85XvRPqJcf1YUq8m5vKeTdxhB5t/FD3u38kPcwDGPkEl3eY1Qq5jESef/JlSKP/LOc9bpz5dKZX4p5+Noey43G67rx6zevS+qho/Ez9j9KLjz6YvPLn2oLpkLB8+Rd49Tdso1ZMes6su5kvc7x6hWQ7ZEEebdTRN5tDJF3Gz/k3c4PeQ/DMEYu0eU9b7UZff3sbXf9SO5btCBGvYMdI5H3ey4V+c01DemcJxc0/rAVJ6Ax7/esukcuX/JPg5abdOvGv6vx8iedCMyWT8DJ+45fPpiMquvoet8jD+2xGowLg9GY9V6X9TRN5N3+C0PebQyRdxs/5N3OD3kPwzBGLl0j7934BqsiDbBwzV/kI989T2Tlt5JRdx19ZytOwJ+w6laq0Te46si82zQmvtWSk8WPOHT2HPHE48kbS8f+/H4Zds89Mrzxf3/TCaZbZp4gumzjUI9Zt7Yq8m4lKIK82xgi7zZ+yLudH/IehmGMXLpG3j99xXWy5IFltRt5T+T9jrkiT96ZTFbVSatsxQnkrTajMfH68id9CZRKvS/yGlaj4TU6Ot9rm8p6Xty6Lt2YhMIcNzP5W//PVowA8l6MU7O9kHcbQ+Tdxg95t/ND3sMwjJFLFHl3o+qtKnT5p86U2bNmttqtq75H3m3N0WqpyLxJrm61mqy1420l6q7UbnQ9EfaMFyOppI84ZbZsevNbZNOrj+iuwteoNMi7vbGQdxtD5N3GD3m380PewzCMkUsUefcrMtTesLpo/QY55ea3i6z5d7nhPbfKrEaMNltxAq3k3c9JRV5H4nVE3l+tZqiJfLPRdRcK48etF1ltpniL9OaeyLu93ZF3G0Pk3cYPebfzQ97DMIyRS3R5j1GpmMe457nn5MSFJybyri9oSr9BNGZZ6nisMvLu1+/JDY/Lt3/9r3Jb44/+221T+g+TOa/+YKMdjq9NW7g113V0XcVdR9vdpmura7y6k/WsUBjk3d7zkXc7Q+TdxhB5t/FD3u38kPcwDGPkgrwbKSfy/vW/FtnwG7n7tKWsjFKSZ1V59w/TLD7+uIbE/11D5lXqu2lz4TB73/KtRNj9zcWuv3DaBwqtCoO821sWebczRN5tDJF3Gz/k3c4PeQ/DMEYuHZH342efJ2vXbcysX91e0pTI+9f+SuSFp2Xph1Z0nSTG6ESWY4SQ97qIfN8jD8te370r+aP/9jeNXdfRdV0dRkNjymzIexla2fsi73aGyLuNIfJu44e82/kh72EYxsglury/d+48mTihX2686qIY9Wv7MR584QV5/YJXIO8VSYeW9zIiHyO0RkfV3eh62XCYokiR96Kk8vdD3u0MkXcbQ+Tdxg95t/ND3sMwjJFLdHkfahNWV23eIi+/cn+RrRtlxVl/6MnlCy0dtZ3y7sqlE12XP/tQ5tKT7YiRzxP2suEwRbki70VJIe92Uvk5IO82usi7jR/ybueHvIdhGCMX5N1IOZH3L4xOcll97vPG3HoveQx5T1PNi5G3rFrTTNhffM9JAxNO29HCyLudKiPvdobIu40h8m7jh7zb+SHvYRjGyCW6vGvYzDuOf6Ocd8b7YtSv7cdA3m2IOyHvfombifysaSfLsZNnNpb/PDnziUqesLvlHJ8/7YOl49er0ETeq1AbnAZ5tzNE3m0MkXcbP+Tdzg95D8MwRi7R5V1f2HTlNbfW7k2qeY2xat0qefm/vFykb6ys/vifYrTZkDpGp+U9LfI/XX3fHstP6oi8vtFVXwh17qYjM2PYVdhV1nWUPfabTZF3+08CebczRN5tDJF3Gz/k3c4PeQ/DMEYu0eVdY96bbXVbbWZA3veeLKvPeDRGmw2pY3STvPtgde34xSvvSuLkdYT9ww+KvG2VyNR1L+218rBxMvGseR0Rdr+syLv9J4G82xki7zaGyLuNH/Ju54e8h2EYI5fo8h6jUjGPgbzbaHervOvKMPte99VkWUd/lZhV40UWvVrkX94sov/WUflW4TU2Qq1TI++tGbXaA3lvRaj198h7a0bN9kDebfyQdzs/5D0Mwxi5IO9Gyg/+8UF5/bWvb1jcX8nS9z8gU/r6jDn2VvJuknf34qR9rv3qoHXY3Soxmz7+iSSG3cXJ68i8/3ZXbTl9w66+GCrGMpSupyDv9t8M8m5niLzbGCLvNn7Iu50f8h6GYYxcOiLvOmn10VWrk/pd/qkzZfasmaLhNMe8YXrt1n+/Z9U9cuJNJ4pM/GtZ+rc/RN5L9tpOy/vw9esbo+t3DsSxu+IXXdbRD6/R5Sh1WUq3xRqVR95LdrqM3ZF3O0Pk3cYQebfxQ97t/JD3MAxj5BJd3v2XNOmbVi88+9RE3hfc8B257a4f1W4iK/Ju66adkne3UoyKuwq8255//+nJso468bTs5q8nnzUqr2vKz5p2UjLxVSfAqtyH2JB3O0Xk3c4QebcxRN5t/JB3Oz/kPQzDGLlEl3cdYb/l6kvkiOnTxJd3XYVm3heul7pNWF3060Vyyq2niLzsRLn9pG/LsaPHxGi3IXOMmPKuYTFjbvnX5I8fx775LSfIC6d9QPTvkCvFtBqVV4EPEWKDvNt/Dsi7nSHybmOIvNv4Ie92fsh7GIYxcoku7yrsX/v8+XvIe6dG3j96wRflZ79YMYh1+gbCD/N5xdSD5Y6Flw/sv/DBhfKROz4iMuVkuX3Wjch7yV4bQ97H/I+bM8NidHT9uY99IqiwN6u+xsrrUpT3N/7ov/3NX47yuENOaIzMH1mYJPJeGFXujsi7nSHybmOIvNv4Ie92fsh7GIYxcoku75++4jpZ8sCyJDzGjbwffthkOe2cy+Tkdx4nn7/4YzHqPXAMLYOWxW1++fQzlfs1azcMCLsf9qPfI++25mqXvPc98rDoxFMNj/FH2TUsxo2y20puS60hNvc3BH7p0/clS1KmJ76qzB/XmPz65snHJ6E2GnKTtyHvtrbQ1Mi7nSHybmOIvNv4Ie92fsh7GIYxcoku71opFyLjV/Ds00/uireuLluxMrmRyArtcWX3XzLly/sN77xBZu29T4x2GzLHCCnvbvJperUYFxbz4ntOlh3jwsSZh24AlfnFK++Unz69JBmVz5J5XZLyNROPkPTIPPJubw3k3c4QebcxRN5t/JB3Oz/kPQzDGLl0RN5jVKzqMfzwnbTIa57pzz5772fl0nsuFfmrs2X+sZfKnH37qx66J9OFkHc3yu5PPnWrxWz8h4ujhcWEbEAXL/+rNcsSqfdXsdHj+CPzxx321/KGScfJ5q07Qhahp/JC3u3NjbzbGCLvNn7Iu50f8h6GYYxcosu7izFPx5V3w1KRTszd8pVl5X3BzMvk/f3Ie5mOu89eI2X7jp3y4pbtZZIl+47+1s2y19VfkZHLHhpIu3XmCbL5P58umz9weun8ujnBE403vv7kyR8noTb6t/7f38aNHi+vPeBIecuUE+QtjZh5/bd+xlaMwLh9RslzL26T7du5ASpGbM+9JowdJX95bqvs3LmzahY9nW7YsGGy3759snbjlp7mYKn8iBHDZd/GNWX9JhhW5Thq5AgZPWq4bHx+a6Es9MkvW3wC0eVdY8znnHTiHiEynZqw6pA7UffDd8rK+2eO/0f5zKRJ8VuxxkccOWJY42IvicAX2YY9vkpG/NNlMvzee0X/rdvOw6bKjre+Vbb/X5ck/+6F7fH1q+THj98rD//5Ibnrt3eIvuk3vR114OvkhMPeKkdOOir5+7BxvcGmSvuPGjlMtm3fKQW7YZVDDPk0o/qGy9bG059iv+Qhj6N0BYc1UvQ1GG7hCVppdi7B8AZEvaZs2UYvrApx+HCREY0bya2N82GRbXSjz7LFJxBd3nWE3Y1s+9Xt5FKR7tguzt0vl7+cpX6eLqcfNnPh0fPkgnET47dijY9YNGwma8UYF8uuk1B7edORj9XrnpF7Vt2bTIB95JmH91jNRvnopFd9A6zGzevIfMi15uvOn7AZewsSNmNjSNiMjZ+m7hs5XMbv0yfPrN9sz6xHc9hr1AgZM3pE4SdAkyfu3aOkOlvt6PLebSPvKuP+BNR0c7RabUaXidRJq/K6y+Ss135YLh1/QGdbtGZHbybvukqMrhYz9p+vGFgxRiec6sTTusayt6N5siasuhdGqcir0Gu4TVbc/JSxhyUTYFXoVeybrWrTjrJ3S57Iu70lkHcbQ+Tdxg95t/PTHJD3MBzbnUt0edfwmGtuvnNgNRetYFbISrsr7h8361j+04Fm67z78j5n+ukyf+KBMYo+ZI6RJe8q7SrsOtruNp2AqsIe+kVKQwFk0dVmdBJsstZ8zoo2ysKfCKsir8tVhnoTbDezRt7trYO82xgi7zZ+yLudH/IehmGMXKLLu1Yqa6nIrFCaGACsx/DlfdarPyA37H+QNcueSu/Le1ZojL5IadPHP5FIO1s2gaLynk5dZHRe0/RCuA3ybv91Ie82hsi7jR/ybueHvIdhGCOXjsh7jIrFOoYv78e+6v1y+6SDYx16SBxH5X34TTfJqCv+aSA0pu7LPMZumKrynlVONzqvS1RqyM3yZx/aI9xmKAo98m7vtci7jSHybuOHvNv5Ie9hGMbIBXk3Uj7xphMbEwXvETnuejn2kLci7wV5utCYve//cWPVmF3LHhIaUxBeareQ8t5K6O9/qvECqY2PtxR6Ha2fsf+RtYmhR96r9T0/FfJuY4i82/gh73Z+yHsYhjFy6Yi866TVtes2ZtYvvf57DAiWY/jyPmPyTLn7ZYdashvyabPi2bcd/1Z5bs5/ll5fNaZq47db3rPKtfzZxqj8Mw+JjtA3E3oXQ68y382TYpH3qr3vpXTIu40h8m7jh7zb+SHvYRjGyCW6vOvkz4kT+uXGqy6KUb+2H8OX9ykvO06WTp7a9mPW8QC6aszet3xr0CRUlfXhcz+cxLNvarwgh60agU7Ie1ZJi4bcqNDrKjczDjiqa5atRN6r9T0/FfJuY4i82/gh73Z+yHsYhjFyiS7veeu8x6hsO47x+mtfLw/+8cEkbKZ/0jGy4pDD23GY2uaZNQlVpd0t9Vh0nffaAohQ8G6R9zyh11H6X+lI/e4/KvlZW3qUXuVeJT/GajfIu72jIu82hsi7jR/ybueHvIdhGCMX5N1I+eX/8vJdb7d8x/dExkyW1Ye+0pjj0Eiu0u6vz67x7M+f9sHkj/7bbci7vb27Wd6zaqer3GjcvIbbrH7uiaZhN5rej5/X0Jt2SD3ybu+HyLuNIfJu44e82/kh72EYxsglurxr2Mw7jn+jnHfG+2LUr+3HSMu7jrz36/uFe3TLknYdZc+LZ0fe7R2lbvKeV2MdkdfRef1bY+k1pj5vcmxoqUfe7f0QebcxRN5t/JB3Oz/kPQzDGLlEl/dWbzSNUemQx0jL+9KDpsqUvr6Qh6hFXmWl3VUKebc371CR92ZSry+XemrjEw2ZfyJ50VRe6I2T+kMa4TZukmyRlW+Qd3s/RN5tDJF3Gz/k3c4PeQ/DMEYu0eVdY96bbXVbbcbJ+5RZP5AnRx0gtx94sBw7ekyMtuuKY1SVduQ9XPMNdXlvJvVlRuo1dn7c6PFybOOtseMa//ZDcA6fNEnWbdoqW7ftCNcwPZYT8m5rcOTdxg95t/ND3sMwjJFLdHmPUamYx9jvi/vJuhfXyYz3/lyW7xzVM/JulXbkPVwv7VV5LyL1ihSY0wAAIABJREFUGlPvh+PkpRm/13jR0frX6Nr0Yw9t/PvQtsTWh2v17ssJebe1CfJu44e82/kh72EYxsgFeTdSHvbZYUkOs077vSx+/jm54YDJMmvvfYy5dm/ytLTrMo/PffLiZLnHKhthM1WoDU6DvBdj6CbKaiy9C8FpFVevOftLW7oR+/7GCH6dXkJVjJBtL+Tdxg95t/FD3u38kPcwDGPk0hF517j3eV+4flD9Lv/UmTJ71swYdQ56DCfvc97/mNy2aYPMn3CgzNm3P+gxuiGz0NLu6oS821sXebcz7Bv9gjzyx0flocayr2XEXo/sYupV6DUURwW/HSvi2GvZ3hyQdxtf5N3GD3m380PewzCMkUt0eV9ww3fkmpvvlFuuvkSOmD4tqeOyFSvltHMuk7NPP7l2q9A4eb/w9CflyvVr5cJxE+SCcRNjtF2UY7RL2pH3cM2HvNtZNpuwmjdiv2FLY8nLnDXrXYncqL0Kvv45eN9Dk7+TP5HWsLfTKZYD8l6MU95eyLuNH/Ju54e8h2EYI5fo8n787PNkzkkn7iHpKvW33fUjuW/Rghj1DnYMJ++XfuhpuXTdM3JW/3i5dPwBwfLvVEahYtpblZ+R91aEWn+PvLdm1GqPqqvNqNiv37xuYIlLF2OvUt9smUtf7nUSrYbg6Mi9i7dXuXcr5rQqe7d8j7zbWgJ5t/FD3u38kPcwDGPkEl3e896w6kJp6rbajMq7XmgveN/P5Py1f5I5+/TL/IkHxmi7thwjlrS7wiPv9mZE3u0Mq8p7qyP7o/brG6JfVu41fz2/9I/aFYqjITk6eu9Cc/Rz/b4bNuTd1grIu40f8m7nh7yHYRgjl+jyPhRH3vXiOf/U/5C//dNqOXavMXL7pINjtF3QY4z+yY9l/HkflxFP7Hp1vb4FtdnLlUIdHHm3k0Te7QzbJe+tSubkPhmpb/xRuVfJ14m0RcJyXP55gu9G8HV0X0N42rkh7za6yLuNH/Ju54e8h2EYI5fo8j4UY959eZ8xarTc/bJDY7RdkGOorI/95ytER9xjSrsrPPJub0bk3c6wU/JepOQq9SryKvTp0fsygu/WuXfhOP4ofgjJR96LtGb+Psi7jR/ybueHvIdhGCOX6PKulRpqq83ohe/b739Y3vyHVTJlZJ8snTw1RtuZjpEl7c+f9kHZ+Ml5pnzLJkbeyxLbc3/k3c6wm+W9SO2yBF9FXz9/auPjSVy+jvIX2ZpJfhKXvzuMJx2ug7wXoYu82yg1T903criM36dPnlm/uZ2HGdJ57zVqhIwZPULWbtxSqJ6TJ+5daD92CkugI/Ietgqdzc3FvC/90Ao5+InfJYVZfegrO1uoJkdXaR9zy7/K2C9dPrDX8+8/PQmR0VCZ2BvybieOvNsZ1l3eixDwJ9duaMi8LompI/d+mE4ZyddjOoHX0fxXHTBNRsm+SXy+vuTKF/0YYTtFGHTzPoy821sHebczRN7tDGPkgLwbKau860oRd5+2VKY/9XvZsGOHLD1oqkzp6zPmHD55ejJqJ6Xd1Q55t7cz8m5n2AvyXpSSk3wdsdeRexeqo3+r9C9/9uEkq1bLZKaP50b03SRbt7rO2IbsawhPr8s+8l60h+bvh7zbGSLvdoYxcogm7y7WPWst92bfxYBgOYbK+7EHnyC3n7JY3vnHJ2T5ls1y+4EHy7Gjx1iyDZo2a632dQuu7chIe7piyLu9qZF3O0PkvRpDF66jUr9h+9Py1F/WJEtkOtF335cVfVcaf2Tf/VuX0xyKwo+8V+uDfirk3c4QebczjJFDNHl/79x5MnFCv9x41UWZ9froBV+UNWs3yB0LXwrniAHAegxf3v/2z6vlpy8+LzccMFlm7b2PNWtzeg2RGfeZT8pe370rySvWCjJlCo68l6GVvS/ybmeIvNsZtop5dyP6bpJtOnRH/+9kv2z4jl96X/jdiH7yd2OE35d+/UzDfXTrhuU2kXd7H0Te7QyRdzvDGDlEk/e89d1dJeu8zrsbedeXNH19w7rkJU36sqZObZ1eQaZMvZH3MrSQdzut7ByQdzvZVvJe5Qj+yL4bvfdj9UMJv5bNhfXov1Xq88Tfyb6G+egNQah4fuS9Sg8ZnAZ5tzNE3u0MY+SAvBsp+yPvV61fI1euX9vRt6x2Y1x7M8TIu7EDNpIz8m5niLzbGbZD3quU6iXJfyl8R8N4NjYm5z7ZmKSrm0q/fqZx/bpVDevxy5eWfyf5yd+NUB/d3Mh/+gZA/3/YuKkyab/R8se1L1apNmkaBJB3ezdA3u0MY+QQTd715UwXnn2qzJ41M7NeOvJ+5TW3yn2LFsSod7BjqLzPmX66zH/7tbL4hU1yxjNPy6wx+8oN+x8U7BhFMkq/ZOnF95wk6z/3pa6Ia29WfuS9SOs23wd5tzNE3u0Mu0Xeq9bEhfVo+l3La+6aoOvE363M4z5L4vyT1XqKL8NZpGzNbgJc+I/moyv66KZPANxTAn0SoFuopwFFyttN+yDv9tZA3u0MY+QQTd4/fcV18qvfPp4b094qJj4GjCrH8OX9p5ufT96yGvNFTXUKkcnii7xX6XWD0yDvdobIu51h3eXdSiBP/nfdDOwa8U+P/LsbgOS7xso+7dj8eP6sGH/3VMC/IdB/u3QuPKgONwXIu70HIe92hjFyiCbvWhkdfdctPbqun69dt1GW37MwRp2DHsOXd10mUpeL7B8+XFYccnjQ42Rlpmu173PtV2T4+l0vX9EXLMV+yZK1ksi7lSBhM3aCIsi7nWKvy7uVoIt5//mq3wxklQ7rcU8BWt0IhH4akFU3/wmBjvir5O8S/Jf+ndwE7H5CoP92Twv03+6Jgfu3e2qQpGm8CKzKhrxXoTY4DfJuZxgjh6jyrhXSEfg7775/UN2OecP03FVoYkCwHMOXd83HrfV+90GHyoy+0Zasc9OmQ2S6Yb32qhVF3quSeykdI+92hsi7nSHybmPYrgmr/oh+Voy/eyqgpXdPBpJ/734S4D8diHFTkKbo3yQkot/kRmG/vcbLpLET5PkXtyXZ+HMMkrS7Q4zcMdyTCPf/Xg038pkj77bfcazU0eU9VsViHSct7265yHas9Z4VIqNx7RrfXtcNebe3HPJuZ4i82xki7zaG7ZJ3W6nyU/thQhr7r5Kvm5sr4FLm3Ry4uQO6n3+D4N84tKvsRfNN3zhourTwZz0l8J82ZN1EZN1IuM/8JxD6WewbCuS9aO/o7H7Iu5F/Wt7dcpEXjpsgF4ybaMz9peTpVWTqGCKTBQN5t3cR5N3OEHm3M0TebQzrJu+22hZP7d8kJKLf5EbhuW0bZMvO5wZG3t0kY3c0/4ZBP3NPItz3nXiyUJzE4D2zbhr8pxL+3nlhSOmbDE0zcsRwOXzCVNm0++mFn0/6yYV+938e+a6qVSCdgQDyboCnSVXeLzx6nlzQ+KNb6BVndLR9/9mzRP/Wrc4hMlmokXdjB2wkR97tDJF3O0Pk3cYQebfx09ShY97TNw5Zwp810dh/2qBp0jcRyWe7VzLya52+mdDvuv2GYuc/7rQ3HDmUJoC8l0Y2OEFa3pdv3Szv/MMTMmVknyydPNWUu05I1T+6dePbUU2V250YebdTRN7tDJF3O0Pk3cYQebfxa4e820vUvhyybhr8pxL+kfNWMkrfZGgaHXl/etMTsnnrjj0Kn3XDcf+Z97avkuScSwB5N3aOtLxrdm9+epU8uW2rVJ20mp6Quunsc+W5j32i69dsr4ISea9CbXAa5N3OEHm3M0TebQyRdxu/XpN3O63sHIh5bxfZsPki70aeWfJ+/to/y23PrZf5EybJnH13vTSjyJY1IfXZRYuHpLQ7Hsh7kZ7RfB/k3c4QebczRN5tDJF3Gz/k3c5Pc0Dew3Bsdy7Iu5FwlryruKvAH7vXGLl90sGFjjBUJ6S2qjzy3opQ6++R99aMWu2BvLci1Pp75L01o2Z7IO82fsi7nR/yHoZhjFyQdyPlLHnXlzUd8/Rjon9r3LvGv+dt6dH2zW85QdYtuHZIj7b7LJB3YwdsJEfe7QyRdztD5N3GEHm38UPe7fyQ9zAMY+SCvBspq7zPf/u1Mmf66YNycqEzGjaj4TNZmz/avmPcONn4D/NE49t7aUPe7a2NvNsZIu92hsi7jSHybuOHvNv5Ie9hGMbIBXk3Us6Td52wqhNXddPQGQ2hcVt6tH2oLf9YBinyXoZW9r7Iu50h8m5niLzbGCLvNn7Iu50f8h6GYYxckHcj5Tx512yvWr9Wrly/RvqHD5cb9z8oEXh/tH2oLv9YBinyXoYW8m6nlZ0D8m4ni7zbGCLvNn7Iu50f8h6GYYxckHcj5Wbyrlm78JmpTz8tt33jRnnT//x2csRei23Pw4y8GztgIzkj73aGyLudIfJuY4i82/gh73Z+yHsYhjFyQd6NlFvJu2a/6GdL5D1nzRUV+HVjx8pnP362/M8PfURmjBot/cOGNya0jpRDRvRJ/4gRMm74MDlkeJ+Ma/xbR+yH+oa821sYebczRN7tDJF3G0Pk3cYPebfzQ97DMIyRC/JupNxK3tNvST39xm/Kv+0/IVmJpsimK9WoxCd/N0R/XOPtZ/q3yr5uU/oa4t+Q/V3/zl/VpsixOrEP8m6njrzbGSLvdobIu40h8m7jh7zb+SHvYRjGyAV5N1JWeb/hPbfKrGknDcpp+Pr10v+ZTyYx7rpt/OS85I/bnty6VZZv2yIbtm+Xp7ZvbbyRdZts2LkjeTOriv36HdsLC75/YBX9ccN3jdo3E/5+3Uf0ZqCzI/zIu7EDNpIj73aGyLudIfJuY4i82/gh73Z+yHsYhjFyQd6NlFXebz9lsRx78AkDOelqMhM+dKr0PfKw6BKQ6xZcJy++Z7DcFz2sSv4GUalvyH1D9Nc3xH7jzu0Dsq//f6oh/Lqp+Ffd3Fr0hzRG+Mc1xN8f5R87TMN5Gp95YT16nBAj/ch71RZ7KR3ybmeIvNsZIu82hsi7jR/ybueHvIdhGCMX5N1IOS3vOuJ+wIlvFhV4XU3m2UWLo75wKRm119H8HVsbor9zD+FPJH9740Zg98h+1RF+H1uW+Cdy34jl180P8Un+74X5IO/GDthIjrzbGSLvdobIu40h8m7jh7zb+SHvYRjGyAV5N1JOy/vE986S0T/5cUfE3VIVHeHXrZn0a1hPqJF+v6yHNmL1d+6UJNRHw3ncyH/WDYAb/XdhP8k+NYz1t7RVOi3ybqeJvNsZIu82hsi7jR/ybueHvIdhGCMX5N1I2Zf3cY0Y932u+UrtxN2IQLLEP7kRaMTyJ8K/fdfkXH/EP/m/Icwnq8zuCYB+p+E/idiP2DX67yb6Jt/tnuybfN+Y8Jt8tvtpwK7P6jXxF3m39mAR5N3OEHm3MUTebfw0dd/I4TJ+nz55Zv1me2Y9msNeo0bImNEjZO3GLYUITJ64d6H92CksAeTdyNPJ+wl/GS8HvO3NSW5r7licrOPO1pqAhs2sfGGLPL+5EcrTiO3XcB4X7qOp9QZAN435182N/ruwn13fVY/1b1ZCN/lX93FPBfTf7oYg+ffu0CA3L8Dl524K/CcEyf5tuDFA3lv3s1Z7IO+tCLX+HnlvzajZHsi7jZ+mRt7tDJF3O8MYOSDvRspO3v/mv1yRhMtsOvtcWf+5Lxlz7Z3kIWPe3RMApafhP4nYb90l/W6ib/LZ7huB5N+N+P9kf+8GoF03A36r+jcG+rl7UqD/9m8O/CcG6RuEgRCiMaNk1LYdste2YQOHaMdNwlDulci7vXWRdxtD5N3GT1Mj73aGyLudYYwckHcjZZX3Xx5+rbzu9I8nK8v8+Re/Tv5mK0YgpLwXO2LxvdzkX03hngokwr/7hiCR/t1PBvzwIP+mwH9CEGJycPHSv7Rn+kbBf4qQvllI/r/7aYLLwQ8z0s/cTcPA917IkX7W6eVHqzBC3qtQG5wGebcxRN5t/DQ18m5niLzbGcbIAXk3UlZ5/83PT5BX/duP91jL3Zh1TyTvZnlvZwP4NwbJTcDuJwXpmwP/iUH6BsGFEPWNHCZrtzbCjRqrDLktxtODMnz8+QgD0r97XoL7v//EYeCz1I2Efp6+mdDPXJiSXyZ/HoP7PO/GAnkv05rZ+yLvNobIu40f8m7npzkg72E4tjsX5N1IeL9PD5PHviwy/kWRP/7+D4y6l+TZq/JeElPT3VvFvKdvFPynCOmbheRGYvfThJduBHaFFrnN3TS4//shR/pZp54wWJiOb7zDYGzjzcXpLf2Uwn2fdaOh3/lhTum8sm46BvLbPXF6z+Pvepla3tZNTzmQd0sPFEHebfw0NSPvdobIu51hjByQdyPlj5wyTL6xSJIJqjpRla0cAeS9HK+svVvJu/0I9hz8+QiaW/oGQj/zw5HcEdM3Esl+3pwFt5+bu+CXNH1TMbBvmyY42ymFzyHriYd/lLybE7ePv2xr/g3Erpe67bv3SNn04vbGsq+NdV8ztmY3L4PKtPtlcEVoZD1daZaum+eCIO9FWrz5Psi7nSHybmcYIwfk3Uh54euHydwHG6ONjUmqOlmVrRwB5L0cr6y96yDv9lq2N4fn9x4mG57fJlsbE3/9LesmI7mB8OY9+Pv7YU7pEmfddAzcUOyeOJ1O48+ZyCJQx6cc7W3JsLm3uvnJO5o/Ab1oiXS6+QF79cnordk3P0XzSc9ZKZrO3y89Ob5KHuk0WaFtIfL1V/RSee8fM1LWbCi2zGHZ43fTk66yZS+6P/JelFRn90PejfzXNS76GjLzp1+siPomVWOxuyY58m5vCuTdznCoxrynn3jscXOwe3nWPIL+sq35++yQjTu3txx5b3bzMuiGaffL4Iq0at7Tlby03TYXpEgd2QcCjkB68YF2kNGbyGHDhsmOnCdo6WM+ccT0dhSDPFsQQN4LdJH3zp0nj65anez5iqkHyx0LL38plXbyxuoyGu/OVp4A8l6eWToF8m5nOFTl3U6meA5DMeY9PV+kKA1/AnrRNCpMw/YeIY+vb4wGGbasULOy2aUnx5dNn7V/VmhbiHzTT6c0/KioeJY9Pk+69iS28/VHlsXI/gEIIO8tIH70gi/KmrUbBoRdRX7ihH658aqLdqVsnChefM9JsvabtwZojt7LAnm3tznybmeIvNsZDkV5t1MpngMx78VZ5e051GPeq95MliE7uvGG1b1HDZd1zxV7+eExL+svkz37BiKAvLcAefzs8+TCs0+V2bNmJnsuWrxErrzmVrlv0YIBeSfevXpvRN6rs3MpkXc7Q+TdzhB5tzFE3m38NPVQl3c7odY5EPPemlE37IG8N2mFZStWymnnXCa3XH2JHDF9WrLnHp81Rt511F1H39nKE0DeyzNLp0De7QyRdztD5N3GEHm38UPe7fw0B+Q9DMd254K8B5B3eewxkalT291W5A8BCEAAAhCAAAQg0OMEkHejvK+dMU0mLF/Z492I6kMAAhCAAAQgAAEIxCCAvLegnBXzPu8L18vyexYOpHx6zQsx2mpIHoOwGXuzEjZjZ0jYjJ0hYTM2hoTN2PhpamLe7QwJm7EzjJED8t6CcsvVZhrpkffqXRV5r87OpUTe7QyRdztD5N3GEHm38UPe7fw0B+Q9DMd254K8FyDcdJ135L0AwfxdkHcTviQx8m5niLzbGSLvNobIu40f8m7nh7yHYRgjF+Q9AGVG3qtDRN6rs2Pk3c7O5YC821ki7zaGyLuNH/Ju54e8h2EYIxfkPQBl5L06ROS9Ojvk3c4OeQ/HEHm3sUTebfyQdzs/5D0Mwxi5IO8BKCPv1SEi79XZIe92dsh7OIbIu40l8m7jh7zb+SHvYRjGyAV5D0AZea8OEXmvzg55t7ND3sMxRN5tLJF3Gz/k3c4PeQ/DMEYuyHsAysh7dYjIe3V2yLudHfIejiHybmOJvNv4Ie92fsh7GIYxckHeY1DmGBCAAAQgAAEIQAACEAhAAHkPAJEsIAABCEAAAhCAAAQgEIMA8h6DMseAAAQgAAEIQAACEIBAAALIewCIZAEBCEAAAhCAAAQgAIEYBJD3ipRbvXW1YrY9l2zZipVy2jmXyS1XXyJHTJ/Wc/W3VPijF3xRfvaLFYOyWH7PQkuWPZf201dcJ3fefT8MA7W848nvuTjQRYuXyLwvXL9HAn7LxRm6PWe8be5AorNPP1nOO+N95TPpwRTuOpxVdfphd3YI5L1Cu6g0rVm7Qe5YeHmSWkV+4oR+ufGqiyrk1rtJjp99nqxdtzEBwMW+fD9QfvctWjCQUMVpyQPLBn1WPtfeSqG/3c9ddMbAjeOCG74jt931IxhW6AYqod+45Xvy6KrV/J5L8FNuV15zK32uBLP0rk4+L//UmTJ71kxDTiR1BPRc+Mvlv8NrurRLIO8VGkal6cKzTx04SXDyrQBxdxJG3quzy7uAcSNUnSn9sTo7HfXUvseTtHIMuX6U45W1t96Ev+P4NzLSbkc5kIN6ztc+fz5PxAMyDZkV8l6SZtbFnQt+SYje7rCrzi6dklFjO0t9qva7lU8xCloSpcrTR057txx+2GTkvSS7rLAZQhXKQdQbxwnjxw48ydXUDGKUY+jvzah7dXaxUiLvJUkj7yWBtdgdeQ/Dk8fGNo5+CBfiVI6lhmv96dm/JI/X+T2XY5e1dzos057j0M4h69zn5l7wW67W9oy6V+MWMxXyXpI28l4SGPIeFlhGbq5PMkHLjlpHnK65+U7hol+MZTrkA3kvxq3ZXo4hfbAYy7w+p6PxxMAXY+jv5d+Ml09NilgEkPcKpLNi3nW1AE625WFysS/PzE/hHrnziNjG0U/tYrdZ/ag107yVUjQlN5Ot+WXt4ZhyPSnOL0vUkffi/Dj/VWPVyVTIewX6rDZTAVpOEuS9OksmulVn51KyYo+doZ8Dv+fyPNN9kNXLyjNMz1Vh5a3yDDUFo+7VuHUiFfJekTrrvFcE5yXz44z1Y51w5C99aD/C0M2h2bq8PCou3u7+79ilYsSzOL/0nsh7eXbpPnjMG6azPF95jMmSzbpMKdeSCvAaSXiKW41bp1Ih750iz3EhAAEIQAACEIAABCBQkgDyXhIYu0MAAhCAAAQgAAEIQKBTBJD3TpHnuBCAAAQgAAEIQAACEChJAHkvCYzdIQABCEAAAhCAAAQg0CkCyHunyHNcCEAAAhCAAAQgAAEIlCSAvJcExu4QgAAEIAABCEAAAhDoFAHkvVPkOS4EIAABCEAAAhCAAARKEkDeSwJjdwhAAAIQgAAEIAABCHSKAPLeKfIcFwIQgAAEIAABCEAAAiUJIO8lgbE7BCAAAQhAAAIQgAAEOkUAee8UeY4LAQhAAAIQgAAEIACBkgSQ95LA2B0CEIAABCAAAQhAAAKdIoC8d4o8x4UABCAAAQhAAAIQgEBJAsh7SWDsDgEIQAACEIAABCAAgU4RQN47RZ7jQgACEIAABCAAAQhAoCQB5L0kMHaHAAQgAAEIQAACEIBApwgg750iz3EhAAEIQAACEIAABCBQkgDyXhIYu0MAAhCAAAQgAAEIQKBTBJD3TpHnuBCAAAQgAAEIQAACEChJAHkvCYzdIQCBoUdgwQ3fkWtuvnOPip19+sly3hnvk+Nnn5d8d9+iBXvso99NGN8vdyy8PPmuVV4z3ja3KcAJ48cmx/noBV+Un/1iRea+l3/qTJk9a6a8d+48eXTVanH/dzsvWrxE5n3hennF1IMHypXOqEg5Zh59hNx59/0DSU9+53Hy+Ys/Vuq4Reox9HoUNYIABCDQPgLIe/vYkjMEIFADAk4ub7n6Ejli+rSBEquE/+C+/xiQX5XdY94wXW686qKBfT59xXWy5IFlA1JfNK+0ZKflW7/XvNas3ZAr37qPk/d0udznzeTdbxon+1nlyPquzHGL1KMG3YQiQgACEOgaAsh71zQFBYEABDpBQKXcjSg3O35aYpetWCmnnXPZoFHvonmFlPeJE/qTEXp38+HKpULfSv6LlCNP3oseF3nvRK/mmBCAwFAmgLwP5dalbhCAQEsCGvbyymmHDBpRz0ukIvq7lU8lI+06+qwC64/El8lLj9FsxLuI9GoZXvOqw+RPz/5FDtx/vySkRZ8G6KaftVPeix63SD1aNhI7QAACEIDAAAHknc4AAQj0NAEn0A6CiznPg+LHii+/Z+Gg3crm1Urei8S8q0Qf84bXJDHuWh4tn47Cz//6t9su70WOS8x7T/+8qDwEINAGAsh7G6CSJQQgUE8CLuTElT4rnMYJt5vMmlfTMnlZYt5V3t0kUi2LexpQZsS7Ssx70eOWKUc9ew2lhgAEIBCXAPIelzdHgwAEakJAw090pZX06HpWrHurKuXl1WrkvVXYiwubUXl3q9y4G4Ey0myR91bHLVOOVhz5HgIQgAAERJB3egEEINCzBFTE/5//9YNk5Dq9OSlNr0KTJ+9V8gop71p+jbl3y1mWkWaLvLc6bply9GxHpOIQgAAEShBA3kvAYlcIQGBoEfBDW/wRdn/FFn9Cqta+mbzr6jO6Fc0rtLz7rVNGmq3y3uy4ZcoxtHoXtYEABCDQHgLIe3u4kisEIFAjAlkvLMqLaW8VNlMmr1byXnTCataTgzLSnFcOF+7jmtJ/SZOLeU83c/q4TFit0Q+BokIAArUggLzXopkoJAQgAAEIQAACEIAABIh5pw9AAAIQgAAEIAABCECgNgQYea9NU1FQCEAAAhCAAAQgAIFeJ4C893oPoP4QgAAEIAABCEAAArUhgLzXpqkoKAQgAAEIQAACEIBArxNA3nu9B1B/CEAAAhCAAAQgAIHaEEDea9NUFBQCEIAABCAAAQhAoNcJIO+93gOoPwQgAAEIQAACEIBAbQgg77VpKgoKAQhAAAIQgAAEINDrBJD3Xu8B1B8CEIAABCAAAQhAoDYEkPfaNBUFhQAEIAABCEAAAhDodQLIe6/3AOoPAQhAAAIQgAAEIFAbAsh7bZqKgkIAAhADvlkRAAACA0lEQVSAAAQgAAEI9DoB5L3XewD1hwAEIAABCEAAAhCoDQHkvTZNRUEhAAEIQAACEIAABHqdAPLe6z2A+kMAAhCAAAQgAAEI1IYA8l6bpqKgEIAABCAAAQhAAAK9TgB57/UeQP0hAAEIQAACEIAABGpDAHmvTVNRUAhAAAIQgAAEIACBXieAvPd6D6D+EIAABCAAAQhAAAK1IYC816apKCgEIAABCEAAAhCAQK8TQN57vQdQfwhAAAIQgAAEIACB2hBA3mvTVBQUAhCAAAQgAAEIQKDXCSDvvd4DqD8EIAABCEAAAhCAQG0IIO+1aSoKCgEIQAACEIAABCDQ6wSQ917vAdQfAhCAAAQgAAEIQKA2BJD32jQVBYUABCAAAQhAAAIQ6HUCyHuv9wDqDwEIQAACEIAABCBQGwLIe22aioJCAAIQgAAEIAABCPQ6AeS913sA9YcABCAAAQhAAAIQqA0B5L02TUVBIQABCEAAAhCAAAR6nQDy3us9gPpDAAIQgAAEIAABCNSGAPJem6aioBCAAAQgAAEIQAACvU4Aee/1HkD9IQABCEAAAhCAAARqQwB5r01TUVAIQAACEIAABCAAgV4ngLz3eg+g/hCAAAQgAAEIQAACtSGAvNemqSgoBCAAAQhAAAIQgECvE0Dee70HUH8IQAACEIAABCAAgdoQ+P8B5xmUPXosbuQAAAAASUVORK5CYII=",
"text/html": [
""
]
},
"metadata": {},
"output_type": "display_data"
}
],
"source": [
"dynamics.plot_history(colors=[\"darkturquoise\", \"green\", \"red\"],\n",
" title=\"Coupled reactions A <-> B and A <-> S (slow but with energetic advantage)\")"
]
},
{
"cell_type": "markdown",
"id": "05dbe681-2fa9-478c-b4eb-8c322e173c0c",
"metadata": {},
"source": [
"### Notice how the \"alternate (shunt) path\" of the reaction, i.e. S (red) \n",
"### has a SLOW START (slow kinetics),\n",
"### but EVENTUALLY DOMINATES (energy advantage)"
]
},
{
"cell_type": "code",
"execution_count": 22,
"id": "f0d7d7ed-158c-4cf4-8d95-ef057ca612d6",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"From time 0 to 0.0025, in 2 steps of 0.00125\n",
"From time 0.0025 to 0.03281, in 97 steps of 0.000313\n",
"From time 0.03281 to 0.05859, in 55 steps of 0.000469\n",
"From time 0.05859 to 0.07617, in 25 steps of 0.000703\n",
"From time 0.07617 to 0.09199, in 15 steps of 0.00105\n",
"From time 0.09199 to 0.1062, in 9 steps of 0.00158\n",
"From time 0.1062 to 0.1205, in 6 steps of 0.00237\n",
"From time 0.1205 to 0.2237, in 29 steps of 0.00356\n",
"From time 0.2237 to 0.3999, in 33 steps of 0.00534\n",
"From time 0.3999 to 0.6402, in 30 steps of 0.00801\n",
"From time 0.6402 to 0.9525, in 26 steps of 0.012\n",
"From time 0.9525 to 1.349, in 22 steps of 0.018\n",
"From time 1.349 to 1.998, in 24 steps of 0.027\n",
"From time 1.998 to 3.66, in 41 steps of 0.0405\n",
"From time 3.66 to 4.876, in 20 steps of 0.0608\n",
"From time 4.876 to 5.971, in 12 steps of 0.0912\n",
"From time 5.971 to 6.929, in 7 steps of 0.137\n",
"From time 6.929 to 7.134, in 1 step of 0.205\n",
"(454 steps total)\n"
]
}
],
"source": [
"dynamics.explain_time_advance()"
]
},
{
"cell_type": "markdown",
"id": "4f451620-86fd-4e3c-a060-6b506f78c13d",
"metadata": {},
"source": [
"### Check the final equilibrium"
]
},
{
"cell_type": "code",
"execution_count": 23,
"id": "96424a3f-31da-46bf-81c0-0a85b9bc476d",
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"0: A <-> B\n",
"Final concentrations: [A] = 1.488 ; [B] = 9.019\n",
"1. Ratio of reactant/product concentrations, adjusted for reaction orders: 6.06134\n",
" Formula used: [B] / [A]\n",
"2. Ratio of forward/reverse reaction rates: 6\n",
"Discrepancy between the two values: 1.022 %\n",
"Reaction IS in equilibrium (within 12% tolerance)\n",
"\n",
"1: A <-> S\n",
"Final concentrations: [A] = 1.488 ; [S] = 39.49\n",
"1. Ratio of reactant/product concentrations, adjusted for reaction orders: 26.5412\n",
" Formula used: [S] / [A]\n",
"2. Ratio of forward/reverse reaction rates: 30\n",
"Discrepancy between the two values: 11.53 %\n",
"Reaction IS in equilibrium (within 12% tolerance)\n",
"\n"
]
},
{
"data": {
"text/plain": [
"True"
]
},
"execution_count": 23,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"# Verify that all the reactions are close to equilibrium\n",
"dynamics.is_in_equilibrium(tolerance=12)"
]
},
{
"cell_type": "markdown",
"id": "fbeba7fe-7a4d-4223-aaa1-6b5b9b3cc1b8",
"metadata": {},
"source": [
"### Please note the **much-longer** timescale from the earlier plots\n",
"If we look at early time interval, this is what it looks like:"
]
},
{
"cell_type": "code",
"execution_count": 24,
"id": "aba771b5-330c-4df0-a666-7b1133aaac41",
"metadata": {},
"outputs": [
{
"data": {
"application/vnd.plotly.v1+json": {
"config": {
"plotlyServerURL": "https://plot.ly"
},
"data": [
{
"hovertemplate": "Chemical=A
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "A",
"line": {
"color": "darkturquoise",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "A",
"showlegend": true,
"type": "scattergl",
"x": [
0,
0.00125,
0.0025,
0.0028125,
0.0031249999999999997,
0.0034374999999999996,
0.0037499999999999994,
0.004062499999999999,
0.0043749999999999995,
0.0046875,
0.005,
0.0053125,
0.005625000000000001,
0.005937500000000001,
0.006250000000000001,
0.0065625000000000015,
0.006875000000000002,
0.007187500000000002,
0.007500000000000002,
0.007812500000000002,
0.008125000000000002,
0.008437500000000002,
0.008750000000000003,
0.009062500000000003,
0.009375000000000003,
0.009687500000000003,
0.010000000000000004,
0.010312500000000004,
0.010625000000000004,
0.010937500000000005,
0.011250000000000005,
0.011562500000000005,
0.011875000000000005,
0.012187500000000006,
0.012500000000000006,
0.012812500000000006,
0.013125000000000006,
0.013437500000000007,
0.013750000000000007,
0.014062500000000007,
0.014375000000000008,
0.014687500000000008,
0.015000000000000008,
0.015312500000000008,
0.015625000000000007,
0.015937500000000007,
0.016250000000000007,
0.016562500000000008,
0.016875000000000008,
0.01718750000000001,
0.01750000000000001,
0.01781250000000001,
0.01812500000000001,
0.01843750000000001,
0.01875000000000001,
0.01906250000000001,
0.01937500000000001,
0.01968750000000001,
0.02000000000000001,
0.02031250000000001,
0.02062500000000001,
0.02093750000000001,
0.021250000000000012,
0.021562500000000012,
0.021875000000000012,
0.022187500000000013,
0.022500000000000013,
0.022812500000000013,
0.023125000000000014,
0.023437500000000014,
0.023750000000000014,
0.024062500000000014,
0.024375000000000015,
0.024687500000000015,
0.025000000000000015,
0.025312500000000016,
0.025625000000000016,
0.025937500000000016,
0.026250000000000016,
0.026562500000000017,
0.026875000000000017,
0.027187500000000017,
0.027500000000000017,
0.027812500000000018,
0.028125000000000018,
0.02843750000000002,
0.02875000000000002,
0.02906250000000002,
0.02937500000000002,
0.02968750000000002,
0.03000000000000002,
0.03031250000000002,
0.03062500000000002,
0.03093750000000002,
0.03125000000000002,
0.03156250000000002,
0.03187500000000002,
0.03218750000000002,
0.03250000000000002,
0.03281250000000002,
0.03328125000000002,
0.033750000000000016,
0.03421875000000001,
0.03468750000000001,
0.03515625000000001,
0.035625000000000004,
0.03609375,
0.0365625,
0.037031249999999995,
0.03749999999999999,
0.03796874999999999,
0.038437499999999986,
0.03890624999999998,
0.03937499999999998,
0.039843749999999976,
0.04031249999999997,
0.04078124999999997,
0.04124999999999997,
0.041718749999999964,
0.04218749999999996,
0.04265624999999996,
0.043124999999999955,
0.04359374999999995,
0.04406249999999995,
0.044531249999999946,
0.04499999999999994,
0.04546874999999994,
0.04593749999999994,
0.046406249999999934,
0.04687499999999993,
0.04734374999999993,
0.047812499999999925,
0.04828124999999992,
0.04874999999999992,
0.049218749999999915,
0.04968749999999991,
0.05015624999999991,
0.050624999999999906,
0.0510937499999999,
0.0515624999999999,
0.0520312499999999,
0.052499999999999894,
0.05296874999999989,
0.05343749999999989,
0.053906249999999885,
0.05437499999999988,
0.05484374999999988,
0.055312499999999876,
0.05578124999999987,
0.05624999999999987,
0.056718749999999866,
0.05718749999999986,
0.05765624999999986,
0.05812499999999986,
0.058593749999999854,
0.05929687499999985,
0.05999999999999985,
0.06070312499999985,
0.06140624999999985,
0.06210937499999985,
0.06281249999999985,
0.06351562499999985,
0.06421874999999985,
0.06492187499999985,
0.06562499999999985,
0.06632812499999985,
0.06703124999999985,
0.06773437499999985,
0.06843749999999985,
0.06914062499999984,
0.06984374999999984,
0.07054687499999984,
0.07124999999999984,
0.07195312499999984,
0.07265624999999984,
0.07335937499999984,
0.07406249999999984,
0.07476562499999984,
0.07546874999999983,
0.07617187499999983,
0.07722656249999983,
0.07828124999999983,
0.07933593749999983,
0.08039062499999983,
0.08144531249999983,
0.08249999999999982,
0.08355468749999982,
0.08460937499999982,
0.08566406249999982,
0.08671874999999982,
0.08777343749999982,
0.08882812499999981,
0.08988281249999981,
0.09093749999999981,
0.09199218749999981,
0.09357421874999981,
0.09515624999999982,
0.09673828124999982,
0.09832031249999983,
0.09990234374999983,
0.10148437499999984,
0.10306640624999984,
0.10464843749999984,
0.10623046874999985,
0.10860351562499986,
0.11097656249999986,
0.11334960937499987,
0.11572265624999988,
0.11809570312499988,
0.12046874999999989,
0.12402832031249988,
0.1275878906249999,
0.1311474609374999,
0.13470703124999991,
0.13826660156249992,
0.14182617187499993,
0.14538574218749994,
0.14894531249999995,
0.15250488281249996,
0.15606445312499997,
0.15962402343749998,
0.16318359375,
0.1667431640625,
0.17030273437500001,
0.17386230468750002,
0.17742187500000003,
0.18098144531250004,
0.18454101562500005,
0.18810058593750006,
0.19166015625000007,
0.19521972656250008,
0.1987792968750001,
0.2023388671875001,
0.20589843750000011,
0.20945800781250012,
0.21301757812500013,
0.21657714843750014,
0.22013671875000015,
0.22369628906250016,
0.22903564453125017,
0.23437500000000017,
0.23971435546875017,
0.24505371093750017,
0.25039306640625014,
0.2557324218750001,
0.2610717773437501,
0.26641113281250006,
0.27175048828125004,
0.27708984375,
0.28242919921875,
0.28776855468749996,
0.29310791015624993,
0.2984472656249999,
0.3037866210937499,
0.30912597656249985,
0.3144653320312498,
0.3198046874999998,
0.32514404296874977,
0.33048339843749974,
0.3358227539062497,
0.3411621093749997,
0.34650146484374966,
0.35184082031249964,
0.3571801757812496,
0.3625195312499996,
0.36785888671874956,
0.37319824218749953,
0.3785375976562495,
0.3838769531249995,
0.38921630859374945,
0.3945556640624994,
0.3998950195312494,
0.4079040527343744,
0.4159130859374994,
0.42392211914062444,
0.43193115234374946,
0.4399401855468745,
0.4479492187499995,
0.4559582519531245,
0.4639672851562495,
0.47197631835937454,
0.47998535156249955,
0.48799438476562457,
0.4960034179687496,
0.5040124511718745,
0.5120214843749995,
0.5200305175781245,
0.5280395507812494,
0.5360485839843744,
0.5440576171874993,
0.5520666503906243,
0.5600756835937493,
0.5680847167968742,
0.5760937499999992,
0.5841027832031241,
0.5921118164062491,
0.6001208496093741,
0.608129882812499,
0.616138916015624,
0.6241479492187489,
0.6321569824218739,
0.6401660156249989,
0.6521795654296864,
0.6641931152343739,
0.6762066650390613,
0.6882202148437488,
0.7002337646484363,
0.7122473144531238,
0.7242608642578113,
0.7362744140624988,
0.7482879638671863,
0.7603015136718738,
0.7723150634765613,
0.7843286132812488,
0.7963421630859363,
0.8083557128906238,
0.8203692626953113,
0.8323828124999988,
0.8443963623046863,
0.8564099121093738,
0.8684234619140613,
0.8804370117187488,
0.8924505615234363,
0.9044641113281238,
0.9164776611328113,
0.9284912109374988,
0.9405047607421863,
0.9525183105468737,
0.970538635253905,
0.9885589599609363,
1.0065792846679675,
1.0245996093749987,
1.04261993408203,
1.060640258789061,
1.0786605834960923,
1.0966809082031235,
1.1147012329101547,
1.1327215576171858,
1.150741882324217,
1.1687622070312482,
1.1867825317382794,
1.2048028564453106,
1.2228231811523418,
1.240843505859373,
1.2588638305664042,
1.2768841552734354,
1.2949044799804665,
1.3129248046874977,
1.330945129394529,
1.34896545410156,
1.375995941162107,
1.4030264282226539,
1.4300569152832008,
1.4570874023437477,
1.4841178894042946,
1.5111483764648415,
1.5381788635253884,
1.5652093505859352,
1.5922398376464821,
1.619270324707029,
1.646300811767576,
1.6733312988281228,
1.7003617858886697,
1.7273922729492166,
1.7544227600097635,
1.7814532470703104,
1.8084837341308573,
1.8355142211914042,
1.862544708251951,
1.889575195312498,
1.9166056823730448,
1.9436361694335917,
1.9706666564941386,
1.9976971435546855,
2.0382428741455056,
2.0787886047363258,
2.119334335327146,
2.159880065917966,
2.200425796508786,
2.2409715270996062,
2.2815172576904263,
2.3220629882812465,
2.3626087188720666,
2.4031544494628867,
2.443700180053707,
2.484245910644527,
2.524791641235347,
2.565337371826167,
2.6058831024169873,
2.6464288330078074,
2.6869745635986275,
2.7275202941894476,
2.7680660247802678,
2.808611755371088,
2.849157485961908,
2.889703216552728,
2.930248947143548,
2.9707946777343683,
3.0113404083251885,
3.0518861389160086,
3.0924318695068287,
3.132977600097649,
3.173523330688469,
3.214069061279289,
3.254614791870109,
3.2951605224609293,
3.3357062530517494,
3.3762519836425695,
3.4167977142333896,
3.4573434448242097,
3.49788917541503,
3.53843490600585,
3.57898063659667,
3.61952636718749,
3.6600720977783103,
3.7208906936645407,
3.781709289550771,
3.8425278854370015,
3.903346481323232,
3.9641650772094623,
4.024983673095693,
4.085802268981923,
4.1466208648681535,
4.207439460754384,
4.268258056640614,
4.329076652526845,
4.389895248413075,
4.4507138442993055,
4.511532440185536,
4.572351036071766,
4.633169631957997,
4.693988227844227,
4.7548068237304575,
4.815625419616688,
4.876444015502918,
4.967671909332264,
5.0588998031616095,
5.150127696990955,
5.241355590820301,
5.332583484649646,
5.423811378478992,
5.5150392723083375,
5.606267166137683,
5.697495059967029,
5.788722953796374,
5.87995084762572,
5.971178741455065,
6.108020582199084,
6.244862422943103,
6.381704263687122,
6.518546104431141,
6.65538794517516,
6.7922297859191785,
6.929071626663197,
7.134334387779226
],
"xaxis": "x",
"y": [
50,
47.9375,
45.9718203125,
45.503467606689455,
45.040610612130536,
44.583184741656176,
44.13112616716149,
43.68437181068289,
43.242859335582004,
42.80652713783324,
42.37531433741376,
41.94916076979458,
41.52800697753174,
41.11179420195624,
40.70046437496171,
40.29396011088851,
39.89222469850329,
39.495202093072756,
39.10283690853061,
38.71507440973658,
38.33186050482635,
37.953141737651485,
37.57886528030812,
37.20897892575354,
36.843431080509475,
36.48217075745115,
36.125147568681136,
35.77231171848688,
35.42361399638108,
35.079005770223745,
34.73843897942521,
34.40186612822897,
34.06924027907342,
33.7405150460317,
33.41564458832859,
33.094583603933565,
32.77728732322922,
32.46371150275405,
32.15381241901881,
31.847546862395507,
31.544872131078236,
31.24574602511499,
30.95012684050962,
30.657973363393076,
30.369244864263205,
30.08390109229219,
29.801902269700932,
29.52320908619953,
29.247782693493097,
28.97558469985216,
28.706577164746857,
28.440722593544198,
28.17798393226766,
27.918324562418356,
27.661708295857064,
27.408099369746434,
27.157462441552592,
26.909762584105536,
26.66496528071754,
26.42303642035896,
26.183942292890737,
25.9476495843529,
25.71412537230847,
25.483337121242084,
25.25525267801267,
25.029840267359607,
24.807068487461645,
24.58690630554808,
24.36932305356146,
24.15428842387128,
23.941772465038063,
23.731745577627223,
23.524178510072097,
23.319042354585637,
23.11630854312012,
22.915948843374363,
22.71793535484784,
22.52224050494119,
22.328837045102567,
22.137698047019235,
21.948796898853967,
21.762107301525656,
21.577603265033638,
21.395259104825218,
21.21504943820588,
21.036949180791698,
20.860933543003426,
20.686978026601807,
20.51505842126358,
20.345150801197747,
20.177231521801605,
20.011277216356067,
19.84726479275982,
19.68517143030189,
19.524974576472104,
19.366651943809064,
19.210181506785144,
19.055541498728118,
18.902710408778947,
18.751666978885332,
18.52775181180322,
18.30776489839474,
18.091636992706473,
17.87930006950633,
17.670687302763703,
17.46573304450902,
17.26437280406592,
17.066543227649603,
16.872182078324794,
16.681228216317045,
16.493621579671107,
16.3093031652503,
16.1282150100708,
15.95030017296503,
15.775502716568253,
15.60376768962276,
15.435041109593989,
15.269269945593106,
15.106402101600649,
14.946386399985913,
14.789172565316864,
14.63471120845547,
14.482953810933408,
14.333852709603219,
14.18736108156003,
14.043432929329098,
13.902023066314472,
13.763087102504185,
13.626581430427443,
13.492463211359366,
13.36069036176894,
13.23122154000585,
13.104016133222029,
12.979034244523758,
12.856236680350252,
12.735584938074755,
12.617041193824212,
12.500568290513659,
12.386129726091557,
12.273689641992348,
12.163212811792574,
12.054664630066979,
11.94801110144106,
11.843218829836612,
11.740255007906866,
11.639087406657852,
11.539684365252757,
11.442014780995988,
11.346048099493826,
11.251754304988525,
11.159103910862813,
11.068067950311784,
10.97861796717924,
10.890726006955568,
10.80436460793432,
10.677077884819875,
10.553114595310111,
10.43238687191389,
10.314809170631266,
10.200298209512985,
10.088772908844662,
9.980154332912683,
9.874365633309992,
9.771331993741068,
9.670980576286421,
9.573240469088036,
9.478042635418173,
9.385319864094955,
9.29500672120912,
9.20703950312728,
9.12135619073792,
9.037896404907269,
8.956601363113075,
8.877413837225113,
8.80027811240211,
8.725139947075567,
8.65194653399174,
8.58064646228379,
8.51118968054687,
8.443527460889623,
8.34465481545959,
8.249609280014845,
8.158239102199522,
8.070398548855577,
7.985947667272658,
7.904752055907955,
7.8266826442003925,
7.751615481118442,
7.679431532095141,
7.6100164840176365,
7.543260557951775,
7.479058329294929,
7.41730855506243,
7.35791400802463,
7.300781317422895,
7.218340565293883,
7.140593489868632,
7.067260995140461,
6.998080590395847,
6.93280540224666,
6.871203245443679,
6.8130557489740635,
6.7581575341535585,
6.706315441619872,
6.632863985666714,
6.565495622342111,
6.503668012642315,
6.446887218369929,
6.394703382568518,
6.346706795460069,
6.280433069491154,
6.221976425326633,
6.170292248126131,
6.124475760963174,
6.083743304922147,
6.047416125202848,
6.014906327756855,
5.9857047158896135,
5.9593702551598104,
5.935520948598144,
5.913825933447956,
5.893998635904289,
5.875790842218578,
5.858987563496375,
5.843402587937505,
5.828874628491694,
5.815263986222215,
5.8024496603404305,
5.790326845116018,
5.778804761872363,
5.767804781209725,
5.7572587966037965,
5.747107815728363,
5.737300740355653,
5.72779330958973,
5.71854718456777,
5.709529155691101,
5.700710455983126,
5.69206616636706,
5.679328967653936,
5.666893548309878,
5.654705235267859,
5.642720318410605,
5.63090384921902,
5.619227881458357,
5.6076700651396525,
5.5962125228177175,
5.584840951531689,
5.573543905078432,
5.562312220407454,
5.5511385591972875,
5.540017041484575,
5.528942952861405,
5.517912510468176,
5.506922675975676,
5.495971006120795,
5.485055533255019,
5.4741746698790115,
5.463327132346845,
5.452511879890534,
5.441728065888519,
5.43097499891948,
5.420252111636559,
5.409558935891619,
5.398895082854533,
5.388260227124469,
5.377654094031582,
5.367076449488452,
5.35652709187929,
5.346005845577714,
5.335512555766072,
5.325047084294959,
5.30939041741361,
5.293795804349249,
5.278262889122162,
5.262791350740035,
5.247380892944181,
5.232031237043137,
5.216742116903688,
5.201513275449505,
5.1863444622133095,
5.171235431625219,
5.156185941815565,
5.1411957537771995,
5.1262646307790405,
5.111392337955163,
5.096578642016583,
5.0818233110487725,
5.067126114369088,
5.052486822426079,
5.037905206728054,
5.023381039792104,
5.008914095107427,
4.994504147108639,
4.980150971156085,
4.965854343521027,
4.951614041374258,
4.937429842777111,
4.923301526674139,
4.909228872886978,
4.895211662109029,
4.881249675900722,
4.860389207077437,
4.839652016371997,
4.819037375245639,
4.798544559468648,
4.77817284909325,
4.757921528427404,
4.7377898860091605,
4.71777721458139,
4.697882811066787,
4.678105976543086,
4.658446016218462,
4.638902239407098,
4.619473959504905,
4.6001604939653955,
4.580961164275698,
4.561875295932717,
4.542902218419438,
4.524041265181368,
4.50529177360312,
4.486653084985133,
4.468124544520532,
4.449705501272121,
4.431395308149517,
4.413193321886414,
4.3950989030179874,
4.377111415858427,
4.350289634788691,
4.323705614785403,
4.297357248221014,
4.271242446150989,
4.245359138148196,
4.219705272138757,
4.194278814239356,
4.1690777485959885,
4.144100077224143,
4.119343819850396,
4.094807013755411,
4.070487713618334,
4.046383991362562,
4.022493936002886,
3.9988156534939776,
3.9753472665802323,
3.952086914646935,
3.9290327535727476,
3.906182955583503,
3.8835357091072957,
3.861089218630859,
3.8388417045572134,
3.805766252318255,
3.773130594967105,
3.7409288846761366,
3.7091553513746263,
3.6778043017148407,
3.6468701180518726,
3.616347257437043,
3.586230250624686,
3.556513701092141,
3.5271922840727745,
3.498260745601862,
3.469713901575156,
3.441546636819968,
3.4137539041786074,
3.386330723604006,
3.359272181267365,
3.33257342867767,
3.3062296818129147,
3.2802362202628728,
3.254588386383272,
3.229281584461213,
3.2043112798916815,
3.1796729983650205,
3.1553623250651954,
3.119381193285491,
3.0841177078300124,
3.04955755518995,
3.0156867073404845,
2.9824914160467824,
2.949958207283568,
2.9180738757659923,
2.8868254795895862,
2.856200334977119,
2.826186011130227,
2.796770325183742,
2.767941337260635,
2.739687345625607,
2.7119968819353266,
2.684858706583422,
2.658261804138292,
2.6321953788719346,
2.6066488503779373,
2.581611849276871,
2.557074213007345,
2.533025981701005,
2.5094573941398024,
2.486358883793905,
2.4637210749386185,
2.441534778848772,
2.4197909900689942,
2.3984808827583857,
2.377595807108089,
2.3571272858303303,
2.3370670107174556,
2.3174068392696316,
2.298138791389781,
2.2792550461444514,
2.260747938589283,
2.2426099566577986,
2.2248337381122383,
2.2074120675552167,
2.190337873500982,
2.173604225505085,
2.1572043313513047,
2.141131534294669,
2.117503198391883,
2.094581765839664,
2.072346087778903,
2.050775648073815,
2.0298505443823824,
2.0095514697930943,
1.98985969501111,
1.9707570510773238,
1.9522259126044894,
1.9342491815148066,
1.9168102712641417,
1.8998930915380803,
1.8834820334060334,
1.8675619549192304,
1.8521181671399392,
1.8371364205882013,
1.8226028920946353,
1.8085041720458177,
1.794827252012225,
1.7815595127450656,
1.7622533124282058,
1.7438135051831811,
1.7262012103958675,
1.7093792922693096,
1.6933122815412252,
1.6779663006505876,
1.6633089924179272,
1.6493094515482993,
1.6359381601267982,
1.6231669237756743,
1.610968816104998,
1.5993181124600018,
1.5826263318503955,
1.567058107952035,
1.5525380365322703,
1.5389946180371774,
1.526366021799498,
1.5145744448654386,
1.5036308059963888,
1.4879832957000314
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=B
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "B",
"line": {
"color": "green",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "B",
"showlegend": true,
"type": "scattergl",
"x": [
0,
0.00125,
0.0025,
0.0028125,
0.0031249999999999997,
0.0034374999999999996,
0.0037499999999999994,
0.004062499999999999,
0.0043749999999999995,
0.0046875,
0.005,
0.0053125,
0.005625000000000001,
0.005937500000000001,
0.006250000000000001,
0.0065625000000000015,
0.006875000000000002,
0.007187500000000002,
0.007500000000000002,
0.007812500000000002,
0.008125000000000002,
0.008437500000000002,
0.008750000000000003,
0.009062500000000003,
0.009375000000000003,
0.009687500000000003,
0.010000000000000004,
0.010312500000000004,
0.010625000000000004,
0.010937500000000005,
0.011250000000000005,
0.011562500000000005,
0.011875000000000005,
0.012187500000000006,
0.012500000000000006,
0.012812500000000006,
0.013125000000000006,
0.013437500000000007,
0.013750000000000007,
0.014062500000000007,
0.014375000000000008,
0.014687500000000008,
0.015000000000000008,
0.015312500000000008,
0.015625000000000007,
0.015937500000000007,
0.016250000000000007,
0.016562500000000008,
0.016875000000000008,
0.01718750000000001,
0.01750000000000001,
0.01781250000000001,
0.01812500000000001,
0.01843750000000001,
0.01875000000000001,
0.01906250000000001,
0.01937500000000001,
0.01968750000000001,
0.02000000000000001,
0.02031250000000001,
0.02062500000000001,
0.02093750000000001,
0.021250000000000012,
0.021562500000000012,
0.021875000000000012,
0.022187500000000013,
0.022500000000000013,
0.022812500000000013,
0.023125000000000014,
0.023437500000000014,
0.023750000000000014,
0.024062500000000014,
0.024375000000000015,
0.024687500000000015,
0.025000000000000015,
0.025312500000000016,
0.025625000000000016,
0.025937500000000016,
0.026250000000000016,
0.026562500000000017,
0.026875000000000017,
0.027187500000000017,
0.027500000000000017,
0.027812500000000018,
0.028125000000000018,
0.02843750000000002,
0.02875000000000002,
0.02906250000000002,
0.02937500000000002,
0.02968750000000002,
0.03000000000000002,
0.03031250000000002,
0.03062500000000002,
0.03093750000000002,
0.03125000000000002,
0.03156250000000002,
0.03187500000000002,
0.03218750000000002,
0.03250000000000002,
0.03281250000000002,
0.03328125000000002,
0.033750000000000016,
0.03421875000000001,
0.03468750000000001,
0.03515625000000001,
0.035625000000000004,
0.03609375,
0.0365625,
0.037031249999999995,
0.03749999999999999,
0.03796874999999999,
0.038437499999999986,
0.03890624999999998,
0.03937499999999998,
0.039843749999999976,
0.04031249999999997,
0.04078124999999997,
0.04124999999999997,
0.041718749999999964,
0.04218749999999996,
0.04265624999999996,
0.043124999999999955,
0.04359374999999995,
0.04406249999999995,
0.044531249999999946,
0.04499999999999994,
0.04546874999999994,
0.04593749999999994,
0.046406249999999934,
0.04687499999999993,
0.04734374999999993,
0.047812499999999925,
0.04828124999999992,
0.04874999999999992,
0.049218749999999915,
0.04968749999999991,
0.05015624999999991,
0.050624999999999906,
0.0510937499999999,
0.0515624999999999,
0.0520312499999999,
0.052499999999999894,
0.05296874999999989,
0.05343749999999989,
0.053906249999999885,
0.05437499999999988,
0.05484374999999988,
0.055312499999999876,
0.05578124999999987,
0.05624999999999987,
0.056718749999999866,
0.05718749999999986,
0.05765624999999986,
0.05812499999999986,
0.058593749999999854,
0.05929687499999985,
0.05999999999999985,
0.06070312499999985,
0.06140624999999985,
0.06210937499999985,
0.06281249999999985,
0.06351562499999985,
0.06421874999999985,
0.06492187499999985,
0.06562499999999985,
0.06632812499999985,
0.06703124999999985,
0.06773437499999985,
0.06843749999999985,
0.06914062499999984,
0.06984374999999984,
0.07054687499999984,
0.07124999999999984,
0.07195312499999984,
0.07265624999999984,
0.07335937499999984,
0.07406249999999984,
0.07476562499999984,
0.07546874999999983,
0.07617187499999983,
0.07722656249999983,
0.07828124999999983,
0.07933593749999983,
0.08039062499999983,
0.08144531249999983,
0.08249999999999982,
0.08355468749999982,
0.08460937499999982,
0.08566406249999982,
0.08671874999999982,
0.08777343749999982,
0.08882812499999981,
0.08988281249999981,
0.09093749999999981,
0.09199218749999981,
0.09357421874999981,
0.09515624999999982,
0.09673828124999982,
0.09832031249999983,
0.09990234374999983,
0.10148437499999984,
0.10306640624999984,
0.10464843749999984,
0.10623046874999985,
0.10860351562499986,
0.11097656249999986,
0.11334960937499987,
0.11572265624999988,
0.11809570312499988,
0.12046874999999989,
0.12402832031249988,
0.1275878906249999,
0.1311474609374999,
0.13470703124999991,
0.13826660156249992,
0.14182617187499993,
0.14538574218749994,
0.14894531249999995,
0.15250488281249996,
0.15606445312499997,
0.15962402343749998,
0.16318359375,
0.1667431640625,
0.17030273437500001,
0.17386230468750002,
0.17742187500000003,
0.18098144531250004,
0.18454101562500005,
0.18810058593750006,
0.19166015625000007,
0.19521972656250008,
0.1987792968750001,
0.2023388671875001,
0.20589843750000011,
0.20945800781250012,
0.21301757812500013,
0.21657714843750014,
0.22013671875000015,
0.22369628906250016,
0.22903564453125017,
0.23437500000000017,
0.23971435546875017,
0.24505371093750017,
0.25039306640625014,
0.2557324218750001,
0.2610717773437501,
0.26641113281250006,
0.27175048828125004,
0.27708984375,
0.28242919921875,
0.28776855468749996,
0.29310791015624993,
0.2984472656249999,
0.3037866210937499,
0.30912597656249985,
0.3144653320312498,
0.3198046874999998,
0.32514404296874977,
0.33048339843749974,
0.3358227539062497,
0.3411621093749997,
0.34650146484374966,
0.35184082031249964,
0.3571801757812496,
0.3625195312499996,
0.36785888671874956,
0.37319824218749953,
0.3785375976562495,
0.3838769531249995,
0.38921630859374945,
0.3945556640624994,
0.3998950195312494,
0.4079040527343744,
0.4159130859374994,
0.42392211914062444,
0.43193115234374946,
0.4399401855468745,
0.4479492187499995,
0.4559582519531245,
0.4639672851562495,
0.47197631835937454,
0.47998535156249955,
0.48799438476562457,
0.4960034179687496,
0.5040124511718745,
0.5120214843749995,
0.5200305175781245,
0.5280395507812494,
0.5360485839843744,
0.5440576171874993,
0.5520666503906243,
0.5600756835937493,
0.5680847167968742,
0.5760937499999992,
0.5841027832031241,
0.5921118164062491,
0.6001208496093741,
0.608129882812499,
0.616138916015624,
0.6241479492187489,
0.6321569824218739,
0.6401660156249989,
0.6521795654296864,
0.6641931152343739,
0.6762066650390613,
0.6882202148437488,
0.7002337646484363,
0.7122473144531238,
0.7242608642578113,
0.7362744140624988,
0.7482879638671863,
0.7603015136718738,
0.7723150634765613,
0.7843286132812488,
0.7963421630859363,
0.8083557128906238,
0.8203692626953113,
0.8323828124999988,
0.8443963623046863,
0.8564099121093738,
0.8684234619140613,
0.8804370117187488,
0.8924505615234363,
0.9044641113281238,
0.9164776611328113,
0.9284912109374988,
0.9405047607421863,
0.9525183105468737,
0.970538635253905,
0.9885589599609363,
1.0065792846679675,
1.0245996093749987,
1.04261993408203,
1.060640258789061,
1.0786605834960923,
1.0966809082031235,
1.1147012329101547,
1.1327215576171858,
1.150741882324217,
1.1687622070312482,
1.1867825317382794,
1.2048028564453106,
1.2228231811523418,
1.240843505859373,
1.2588638305664042,
1.2768841552734354,
1.2949044799804665,
1.3129248046874977,
1.330945129394529,
1.34896545410156,
1.375995941162107,
1.4030264282226539,
1.4300569152832008,
1.4570874023437477,
1.4841178894042946,
1.5111483764648415,
1.5381788635253884,
1.5652093505859352,
1.5922398376464821,
1.619270324707029,
1.646300811767576,
1.6733312988281228,
1.7003617858886697,
1.7273922729492166,
1.7544227600097635,
1.7814532470703104,
1.8084837341308573,
1.8355142211914042,
1.862544708251951,
1.889575195312498,
1.9166056823730448,
1.9436361694335917,
1.9706666564941386,
1.9976971435546855,
2.0382428741455056,
2.0787886047363258,
2.119334335327146,
2.159880065917966,
2.200425796508786,
2.2409715270996062,
2.2815172576904263,
2.3220629882812465,
2.3626087188720666,
2.4031544494628867,
2.443700180053707,
2.484245910644527,
2.524791641235347,
2.565337371826167,
2.6058831024169873,
2.6464288330078074,
2.6869745635986275,
2.7275202941894476,
2.7680660247802678,
2.808611755371088,
2.849157485961908,
2.889703216552728,
2.930248947143548,
2.9707946777343683,
3.0113404083251885,
3.0518861389160086,
3.0924318695068287,
3.132977600097649,
3.173523330688469,
3.214069061279289,
3.254614791870109,
3.2951605224609293,
3.3357062530517494,
3.3762519836425695,
3.4167977142333896,
3.4573434448242097,
3.49788917541503,
3.53843490600585,
3.57898063659667,
3.61952636718749,
3.6600720977783103,
3.7208906936645407,
3.781709289550771,
3.8425278854370015,
3.903346481323232,
3.9641650772094623,
4.024983673095693,
4.085802268981923,
4.1466208648681535,
4.207439460754384,
4.268258056640614,
4.329076652526845,
4.389895248413075,
4.4507138442993055,
4.511532440185536,
4.572351036071766,
4.633169631957997,
4.693988227844227,
4.7548068237304575,
4.815625419616688,
4.876444015502918,
4.967671909332264,
5.0588998031616095,
5.150127696990955,
5.241355590820301,
5.332583484649646,
5.423811378478992,
5.5150392723083375,
5.606267166137683,
5.697495059967029,
5.788722953796374,
5.87995084762572,
5.971178741455065,
6.108020582199084,
6.244862422943103,
6.381704263687122,
6.518546104431141,
6.65538794517516,
6.7922297859191785,
6.929071626663197,
7.134334387779226
],
"xaxis": "x",
"y": [
0,
1.875,
3.6609375,
4.086203100585938,
4.506413417053986,
4.921627870578563,
5.33190518398381,
5.737303389950974,
6.137879839129328,
6.53369120815177,
6.9247935075562195,
7.311242089613917,
7.69309165606572,
8.070396265767478,
8.443209342245556,
8.811583681163563,
9.175571457701325,
9.535224233847135,
9.890592965604306,
10.241728010113023,
10.588679132688503,
10.931495513776424,
11.270225755826631,
11.60491789008604,
11.935619383311721,
12.262377144405074,
12.585237530968046,
12.904246355782293,
13.219448893212197,
13.530889885532625,
13.838613549182329,
14.142663580943843,
14.443083164050766,
14.73991497422325,
15.033201185632574,
15.322983476795605,
15.60930303639999,
15.892200569060888,
16.17171630101005,
16.447889985718025,
16.7207609094503,
16.99036789675814,
17.25674931590491,
17.519943084228586,
17.77998667344129,
18.036917114866505,
18.290771004614765,
18.541584508698502,
18.78939336808678,
19.034232903700644,
19.276138021349727,
19.51514321661087,
19.75128257964939,
19.9845897999837,
20.215098171193898,
20.442840595575067,
20.667849588735855,
20.89015728414301,
21.109795437612526,
21.326795431747982,
21.54118828032674,
21.75300463263458,
21.962274777749396,
22.169028648774553,
22.37329582702249,
22.575105546149135,
22.774486696239773,
22.97146782784685,
23.166077155980354,
23.358342564051274,
23.548291607768736,
23.735951518991328,
23.921349209533158,
24.104511274925187,
24.285463998132357,
24.464233353227026,
24.640845009019245,
24.81532433264435,
24.98769639310842,
25.157985964792022,
25.32621753091284,
25.492415286947544,
25.65660314401349,
25.818804732210662,
25.97904340392432,
26.137342237088866,
26.293724038413337,
26.448211346568975,
26.600826435339354,
26.75159131673348,
26.900527744062316,
27.047657214979107,
27.19300097448404,
27.33658001789353,
27.478415093774654,
27.618526706845056,
27.756935120838822,
27.893660361338622,
28.028722218574607,
28.162140250190387,
28.359830550869578,
28.55390870786946,
28.744438428219066,
28.931482295862864,
29.115101791459367,
29.29535731183075,
29.472308189069555,
29.6460127093086,
29.81652813115998,
29.983910703829018,
30.148215684908877,
30.309497357861495,
30.46780904919034,
30.62320314531042,
30.77573110912092,
30.92544349628566,
31.07238997122656,
31.216619322835164,
31.35817947990717,
31.497117526304898,
31.633479715852424,
31.767311486968165,
31.89865747703949,
32.02756153654393,
32.15406674292145,
32.278215414202165,
32.40004912239382,
32.51960870663326,
32.63693428610605,
32.75206527273837,
32.86504038366513,
32.97589765347829,
33.084674446259285,
33.1914074673993,
33.2961327752112,
33.39888579233672,
33.49970131695261,
33.59861353377916,
33.69565602489471,
33.79086178035953,
33.88426320865233,
33.97589214692288,
34.065779871063846,
34.15395710560505,
34.240454033433366,
34.325300305341194,
34.40852504940668,
34.490156880208495,
34.570223907878265,
34.64875374699331,
34.72577352531269,
34.80130989235925,
34.87538902785029,
34.94803664997972,
35.01927802355414,
35.1240679402012,
35.22580450048185,
35.32456854252967,
35.42043876682677,
35.51349179273015,
35.60380221350325,
35.691442649892345,
35.776483802286194,
35.85899450149641,
35.93904175819507,
36.01669081104495,
36.0920051735572,
36.16504667970976,
36.23587552835966,
36.30455032648077,
36.37112813125833,
36.435664491070256,
36.49821348538485,
36.55882776360346,
36.61755858287601,
36.674455844916565,
36.729568131845404,
36.782942741083275,
36.83462571932295,
36.88466189560249,
36.95731142245461,
37.02644944427066,
37.09221557090486,
37.1547438744482,
37.21416310888267,
37.27059692102297,
37.32416405309122,
37.37497853725665,
37.42314988245885,
37.46878325382077,
37.51197964494532,
37.55283604337787,
37.59144558950572,
37.62789772915483,
37.6622783601339,
37.71086577938162,
37.755156149167064,
37.79540622611957,
37.83185748972529,
37.86473705127457,
37.894258508729514,
37.92062275072937,
37.94401871275987,
37.964624088332634,
37.991596952610806,
38.013020665092135,
38.029394131681855,
38.04117172877335,
38.04876727732274,
38.05255766225416,
38.05305036214574,
38.04645711341405,
38.033738794290514,
38.015727630039194,
37.99314441558984,
37.96661343260069,
37.93667536959462,
37.903798512494376,
37.86838843709625,
37.83079640402559,
37.791326629871136,
37.750242584942555,
37.70777244795579,
37.66411383050716,
37.61943786908854,
37.573892769310056,
37.52760687566249,
37.48069133033481,
37.43324237609933,
37.385343350912834,
37.337066415503024,
37.28847404968539,
37.23962034837007,
37.190552144074026,
37.14130997916401,
37.0919289479466,
37.042439426028785,
36.992867702039966,
36.94323652478614,
36.86873010334279,
36.79417251029523,
36.71961345098013,
36.645092546193645,
36.57064135746877,
36.49628500567028,
36.42204346457037,
36.34793259466982,
36.27396496942411,
36.2001505355602,
36.126497140799074,
36.053010955609366,
35.97969681027085,
35.90655846425398,
35.83359882150647,
35.760820102509015,
35.68822398178106,
35.61581169777424,
35.543584140698314,
35.471541922710585,
35.39968543401048,
35.32801488766945,
35.25653035545822,
35.185231796479144,
35.11411908004842,
35.043192003982796,
34.97245030921359,
34.90189369146551,
34.83152181058962,
34.7613342980216,
34.69133076274166,
34.621510796037,
34.55187397530738,
34.44769281300491,
34.343921759695455,
34.240559302228725,
34.13760390251718,
34.03505400694111,
33.9329080529134,
33.83116447345951,
33.729821700410575,
33.62887816662732,
33.5283323075468,
33.42818256225593,
33.32842737423439,
33.229065191866475,
33.1300944687916,
33.03151366414195,
32.93332124270145,
32.83551567500969,
32.73809543742738,
32.641059012175084,
32.54440488735316,
32.448131556948695,
32.35223752083333,
32.25672128475478,
32.161581360323964,
32.066816264999034,
31.972424522067396,
31.87840466062623,
31.784755215562054,
31.69147472752962,
31.59856174293039,
31.459741349370894,
31.321741335936334,
31.18455685447352,
31.048183085476808,
30.912615237920313,
30.777848549090447,
30.643878284419035,
30.51069973731726,
30.37830822901043,
30.24669910837369,
30.11586775176867,
29.98580956288105,
29.856519972559127,
29.727994438653273,
29.600228445856377,
29.47321750554522,
29.346957155622782,
29.221442960361475,
29.09667051024732,
28.97263542182503,
28.849333337544,
28.72675992560524,
28.60491087980918,
28.483781919404386,
28.363368788937176,
28.243667258102118,
28.065176053339048,
27.88826707978895,
27.71292631179933,
27.539139848047768,
27.366893910439806,
27.196174843016582,
27.026969110872177,
26.85926329908053,
26.693044111631895,
26.52829837037869,
26.365013013990726,
26.203175096919683,
26.042771788372754,
25.8837903712954,
25.726218241363124,
25.57004290598218,
25.41525198329913,
25.26183320121919,
25.109774396433288,
24.959063513453724,
24.809688603658394,
24.66163782434348,
24.441530244505074,
24.224349372597413,
24.010056293016962,
23.798612607609908,
23.589980428791772,
23.384122372758522,
23.18100155278794,
22.98058157263007,
22.782826519985544,
22.58770096007064,
22.39516992926788,
22.205198928861066,
22.01775391885363,
21.83280131186916,
21.65030796713305,
21.47024118453414,
21.292568698765365,
21.117258673542274,
20.944279695898445,
20.77360077055674,
20.60519131437541,
20.439021150868058,
20.27506050479644,
20.113279996835193,
19.873835959043653,
19.6391676502267,
19.40917981811809,
19.183779110265046,
18.962874036136338,
18.74637492998612,
18.53419391445846,
18.326244864917754,
18.122443374490608,
17.922706719804935,
17.726953827412395,
17.535105240880554,
17.34708308854137,
17.16281105188297,
16.9822143345718,
16.805219632092715,
16.63175510199451,
16.46175033472897,
16.295136325071493,
16.131845444111782,
15.971811411803131,
15.814969270059247,
15.661255356387658,
15.51060727804899,
15.362963886731658,
15.218265253731651,
15.076452645627379,
14.937468500439671,
14.801256404267269,
14.667761068388339,
14.53692830681867,
14.4087050143175,
14.283039144832008,
14.159879690371731,
14.039176660304333,
13.920881061064323,
13.804944876266482,
13.691321047215936,
13.579963453806943,
13.470826895802668,
13.363867074488333,
13.206627324791262,
13.054091812682511,
12.90611979861416,
12.76257475362927,
12.623324233391052,
12.48823975598081,
12.35719668335184,
12.23007410632998,
12.106754733054638,
11.987124780757446,
11.871073870778547,
11.758494926723886,
11.649284075669197,
11.543340552319988,
11.440566606038496,
11.340867410652598,
11.244150976962484,
11.150328067865615,
11.059312116020049,
10.971019143972221,
10.842541958023167,
10.719830367687743,
10.60262563369859,
10.490680628084286,
10.383759313078851,
10.281636243474766,
10.184096091165577,
10.090933191372997,
10.001951108392555,
9.916962222881523,
9.83578733267357,
9.758255283648136,
9.647176239654135,
9.543574480383842,
9.446946465866878,
9.356823629195162,
9.272764477334546,
9.194375760178556,
9.121213891980563,
9.019168479129675
],
"yaxis": "y"
},
{
"hovertemplate": "Chemical=S
SYSTEM TIME=%{x}
Concentration=%{y}",
"legendgroup": "S",
"line": {
"color": "red",
"dash": "solid"
},
"marker": {
"symbol": "circle"
},
"mode": "lines",
"name": "S",
"showlegend": true,
"type": "scattergl",
"x": [
0,
0.00125,
0.0025,
0.0028125,
0.0031249999999999997,
0.0034374999999999996,
0.0037499999999999994,
0.004062499999999999,
0.0043749999999999995,
0.0046875,
0.005,
0.0053125,
0.005625000000000001,
0.005937500000000001,
0.006250000000000001,
0.0065625000000000015,
0.006875000000000002,
0.007187500000000002,
0.007500000000000002,
0.007812500000000002,
0.008125000000000002,
0.008437500000000002,
0.008750000000000003,
0.009062500000000003,
0.009375000000000003,
0.009687500000000003,
0.010000000000000004,
0.010312500000000004,
0.010625000000000004,
0.010937500000000005,
0.011250000000000005,
0.011562500000000005,
0.011875000000000005,
0.012187500000000006,
0.012500000000000006,
0.012812500000000006,
0.013125000000000006,
0.013437500000000007,
0.013750000000000007,
0.014062500000000007,
0.014375000000000008,
0.014687500000000008,
0.015000000000000008,
0.015312500000000008,
0.015625000000000007,
0.015937500000000007,
0.016250000000000007,
0.016562500000000008,
0.016875000000000008,
0.01718750000000001,
0.01750000000000001,
0.01781250000000001,
0.01812500000000001,
0.01843750000000001,
0.01875000000000001,
0.01906250000000001,
0.01937500000000001,
0.01968750000000001,
0.02000000000000001,
0.02031250000000001,
0.02062500000000001,
0.02093750000000001,
0.021250000000000012,
0.021562500000000012,
0.021875000000000012,
0.022187500000000013,
0.022500000000000013,
0.022812500000000013,
0.023125000000000014,
0.023437500000000014,
0.023750000000000014,
0.024062500000000014,
0.024375000000000015,
0.024687500000000015,
0.025000000000000015,
0.025312500000000016,
0.025625000000000016,
0.025937500000000016,
0.026250000000000016,
0.026562500000000017,
0.026875000000000017,
0.027187500000000017,
0.027500000000000017,
0.027812500000000018,
0.028125000000000018,
0.02843750000000002,
0.02875000000000002,
0.02906250000000002,
0.02937500000000002,
0.02968750000000002,
0.03000000000000002,
0.03031250000000002,
0.03062500000000002,
0.03093750000000002,
0.03125000000000002,
0.03156250000000002,
0.03187500000000002,
0.03218750000000002,
0.03250000000000002,
0.03281250000000002,
0.03328125000000002,
0.033750000000000016,
0.03421875000000001,
0.03468750000000001,
0.03515625000000001,
0.035625000000000004,
0.03609375,
0.0365625,
0.037031249999999995,
0.03749999999999999,
0.03796874999999999,
0.038437499999999986,
0.03890624999999998,
0.03937499999999998,
0.039843749999999976,
0.04031249999999997,
0.04078124999999997,
0.04124999999999997,
0.041718749999999964,
0.04218749999999996,
0.04265624999999996,
0.043124999999999955,
0.04359374999999995,
0.04406249999999995,
0.044531249999999946,
0.04499999999999994,
0.04546874999999994,
0.04593749999999994,
0.046406249999999934,
0.04687499999999993,
0.04734374999999993,
0.047812499999999925,
0.04828124999999992,
0.04874999999999992,
0.049218749999999915,
0.04968749999999991,
0.05015624999999991,
0.050624999999999906,
0.0510937499999999,
0.0515624999999999,
0.0520312499999999,
0.052499999999999894,
0.05296874999999989,
0.05343749999999989,
0.053906249999999885,
0.05437499999999988,
0.05484374999999988,
0.055312499999999876,
0.05578124999999987,
0.05624999999999987,
0.056718749999999866,
0.05718749999999986,
0.05765624999999986,
0.05812499999999986,
0.058593749999999854,
0.05929687499999985,
0.05999999999999985,
0.06070312499999985,
0.06140624999999985,
0.06210937499999985,
0.06281249999999985,
0.06351562499999985,
0.06421874999999985,
0.06492187499999985,
0.06562499999999985,
0.06632812499999985,
0.06703124999999985,
0.06773437499999985,
0.06843749999999985,
0.06914062499999984,
0.06984374999999984,
0.07054687499999984,
0.07124999999999984,
0.07195312499999984,
0.07265624999999984,
0.07335937499999984,
0.07406249999999984,
0.07476562499999984,
0.07546874999999983,
0.07617187499999983,
0.07722656249999983,
0.07828124999999983,
0.07933593749999983,
0.08039062499999983,
0.08144531249999983,
0.08249999999999982,
0.08355468749999982,
0.08460937499999982,
0.08566406249999982,
0.08671874999999982,
0.08777343749999982,
0.08882812499999981,
0.08988281249999981,
0.09093749999999981,
0.09199218749999981,
0.09357421874999981,
0.09515624999999982,
0.09673828124999982,
0.09832031249999983,
0.09990234374999983,
0.10148437499999984,
0.10306640624999984,
0.10464843749999984,
0.10623046874999985,
0.10860351562499986,
0.11097656249999986,
0.11334960937499987,
0.11572265624999988,
0.11809570312499988,
0.12046874999999989,
0.12402832031249988,
0.1275878906249999,
0.1311474609374999,
0.13470703124999991,
0.13826660156249992,
0.14182617187499993,
0.14538574218749994,
0.14894531249999995,
0.15250488281249996,
0.15606445312499997,
0.15962402343749998,
0.16318359375,
0.1667431640625,
0.17030273437500001,
0.17386230468750002,
0.17742187500000003,
0.18098144531250004,
0.18454101562500005,
0.18810058593750006,
0.19166015625000007,
0.19521972656250008,
0.1987792968750001,
0.2023388671875001,
0.20589843750000011,
0.20945800781250012,
0.21301757812500013,
0.21657714843750014,
0.22013671875000015,
0.22369628906250016,
0.22903564453125017,
0.23437500000000017,
0.23971435546875017,
0.24505371093750017,
0.25039306640625014,
0.2557324218750001,
0.2610717773437501,
0.26641113281250006,
0.27175048828125004,
0.27708984375,
0.28242919921875,
0.28776855468749996,
0.29310791015624993,
0.2984472656249999,
0.3037866210937499,
0.30912597656249985,
0.3144653320312498,
0.3198046874999998,
0.32514404296874977,
0.33048339843749974,
0.3358227539062497,
0.3411621093749997,
0.34650146484374966,
0.35184082031249964,
0.3571801757812496,
0.3625195312499996,
0.36785888671874956,
0.37319824218749953,
0.3785375976562495,
0.3838769531249995,
0.38921630859374945,
0.3945556640624994,
0.3998950195312494,
0.4079040527343744,
0.4159130859374994,
0.42392211914062444,
0.43193115234374946,
0.4399401855468745,
0.4479492187499995,
0.4559582519531245,
0.4639672851562495,
0.47197631835937454,
0.47998535156249955,
0.48799438476562457,
0.4960034179687496,
0.5040124511718745,
0.5120214843749995,
0.5200305175781245,
0.5280395507812494,
0.5360485839843744,
0.5440576171874993,
0.5520666503906243,
0.5600756835937493,
0.5680847167968742,
0.5760937499999992,
0.5841027832031241,
0.5921118164062491,
0.6001208496093741,
0.608129882812499,
0.616138916015624,
0.6241479492187489,
0.6321569824218739,
0.6401660156249989,
0.6521795654296864,
0.6641931152343739,
0.6762066650390613,
0.6882202148437488,
0.7002337646484363,
0.7122473144531238,
0.7242608642578113,
0.7362744140624988,
0.7482879638671863,
0.7603015136718738,
0.7723150634765613,
0.7843286132812488,
0.7963421630859363,
0.8083557128906238,
0.8203692626953113,
0.8323828124999988,
0.8443963623046863,
0.8564099121093738,
0.8684234619140613,
0.8804370117187488,
0.8924505615234363,
0.9044641113281238,
0.9164776611328113,
0.9284912109374988,
0.9405047607421863,
0.9525183105468737,
0.970538635253905,
0.9885589599609363,
1.0065792846679675,
1.0245996093749987,
1.04261993408203,
1.060640258789061,
1.0786605834960923,
1.0966809082031235,
1.1147012329101547,
1.1327215576171858,
1.150741882324217,
1.1687622070312482,
1.1867825317382794,
1.2048028564453106,
1.2228231811523418,
1.240843505859373,
1.2588638305664042,
1.2768841552734354,
1.2949044799804665,
1.3129248046874977,
1.330945129394529,
1.34896545410156,
1.375995941162107,
1.4030264282226539,
1.4300569152832008,
1.4570874023437477,
1.4841178894042946,
1.5111483764648415,
1.5381788635253884,
1.5652093505859352,
1.5922398376464821,
1.619270324707029,
1.646300811767576,
1.6733312988281228,
1.7003617858886697,
1.7273922729492166,
1.7544227600097635,
1.7814532470703104,
1.8084837341308573,
1.8355142211914042,
1.862544708251951,
1.889575195312498,
1.9166056823730448,
1.9436361694335917,
1.9706666564941386,
1.9976971435546855,
2.0382428741455056,
2.0787886047363258,
2.119334335327146,
2.159880065917966,
2.200425796508786,
2.2409715270996062,
2.2815172576904263,
2.3220629882812465,
2.3626087188720666,
2.4031544494628867,
2.443700180053707,
2.484245910644527,
2.524791641235347,
2.565337371826167,
2.6058831024169873,
2.6464288330078074,
2.6869745635986275,
2.7275202941894476,
2.7680660247802678,
2.808611755371088,
2.849157485961908,
2.889703216552728,
2.930248947143548,
2.9707946777343683,
3.0113404083251885,
3.0518861389160086,
3.0924318695068287,
3.132977600097649,
3.173523330688469,
3.214069061279289,
3.254614791870109,
3.2951605224609293,
3.3357062530517494,
3.3762519836425695,
3.4167977142333896,
3.4573434448242097,
3.49788917541503,
3.53843490600585,
3.57898063659667,
3.61952636718749,
3.6600720977783103,
3.7208906936645407,
3.781709289550771,
3.8425278854370015,
3.903346481323232,
3.9641650772094623,
4.024983673095693,
4.085802268981923,
4.1466208648681535,
4.207439460754384,
4.268258056640614,
4.329076652526845,
4.389895248413075,
4.4507138442993055,
4.511532440185536,
4.572351036071766,
4.633169631957997,
4.693988227844227,
4.7548068237304575,
4.815625419616688,
4.876444015502918,
4.967671909332264,
5.0588998031616095,
5.150127696990955,
5.241355590820301,
5.332583484649646,
5.423811378478992,
5.5150392723083375,
5.606267166137683,
5.697495059967029,
5.788722953796374,
5.87995084762572,
5.971178741455065,
6.108020582199084,
6.244862422943103,
6.381704263687122,
6.518546104431141,
6.65538794517516,
6.7922297859191785,
6.929071626663197,
7.134334387779226
],
"xaxis": "x",
"y": [
0,
0.1875,
0.3672421875,
0.4103292927246094,
0.45297597081548313,
0.49518738776526755,
0.5369686488547025,
0.5783247993661397,
0.6192608252886747,
0.6597816540149926,
0.6998921550300232,
0.7395971405915039,
0.7789013664025428,
0.8178095322762787,
0.856326282792729,
0.8944562079479184,
0.9322038437953779,
0.9695736730801061,
1.006570125865078,
1.0431975801503923,
1.0794603624851407,
1.1153627485720876,
1.150908963865243,
1.186103184160411,
1.2209495361788,
1.2554520981437722,
1.2896149003508157,
1.3234419257308183,
1.3569371104067207,
1.3901043442436278,
1.422947471392455,
1.455470290827185,
1.4876765568758112,
1.5195699797450402,
1.5511542260388278,
1.5824329192708222,
1.6134096403707827,
1.6440879281850485,
1.6744712799711245,
1.7045631518864557,
1.734366959471455,
1.7638860781268575,
1.7931238435854613,
1.822083552378327,
1.850768462295496,
1.8791817928412962,
1.9073267256842938,
1.9352064051019608,
1.9628239384201134,
1.9901823964471874,
2.0172848139034096,
2.0441341898449252,
2.0707334880829404,
2.0970856375979388,
2.1231935329490312,
2.1490600346784925,
2.174687969711546,
2.2000801317514482,
2.22523928166993,
2.25016814789305,
2.274869426782515,
2.2993457830125132,
2.323599849942125,
2.3476342299833535,
2.371451494964831,
2.39505418649125,
2.418444816298572,
2.4416258666050576,
2.464599790458178,
2.4873690120774397,
2.5099359271931916,
2.53230290338144,
2.5544722803947346,
2.576446370489165,
2.598227458747511,
2.6198178033986004,
2.6412196361329077,
2.6624351624144484,
2.6834665617890052,
2.704315988188733,
2.7249855702331827,
2.7454774115267884,
2.7657935909528586,
2.78593616296411,
2.8059071578697914,
2.825708582119426,
2.8453424185832272,
2.8648106268292124,
2.884115143397063,
2.9032578820687664,
2.9222407341360745,
2.941065568664822,
2.959734232756135,
2.978248551804574,
2.996610329753238,
3.0148213493458758,
3.0328833723760296,
3.0507981399332538,
3.0685673726464384,
3.0861927709242734,
3.112417637327194,
3.1383263937357926,
3.163924579074454,
3.189217634630803,
3.2142109057769233,
3.2389096436602265,
3.2633190068645206,
3.2874440630417916,
3.311289790515219,
3.334861079853933,
3.3581627354200108,
3.3811994768882006,
3.403975940738855,
3.4264966817245446,
3.448766174310821,
3.4707888140915744,
3.492568919179446,
3.514110731571726,
3.5354184184921738,
3.556496073709183,
3.577347718830708,
3.5979773045763643,
3.618388712027103,
3.6385857538528517,
3.658572175518519,
3.6783516564687355,
3.6979278112917076,
3.717304190862558,
3.736484283466508,
3.755471515902259,
3.7742692545659255,
3.7928808065158552,
3.811309420518683,
3.8295582880769397,
3.8476305444385477,
3.8655292695885195,
3.8832574892231753,
3.900818175707183,
3.9182142490137317,
3.9354485776481254,
3.9525239795551,
3.9694432230101415,
3.9862090274950948,
4.002824064558332,
4.019290958659764,
4.035612288000945,
4.051790585340558,
4.067828338795507,
4.083727992627901,
4.09949194801816,
4.115122563824487,
4.1306221573289585,
4.145993004970459,
4.161237343064697,
4.176357368511522,
4.19885417497891,
4.221080904208024,
4.2430445855564285,
4.26475206254195,
4.286209997756853,
4.3074248776520765,
4.328403017194961,
4.349150564403802,
4.369673504762505,
4.389977665518499,
4.410068719866996,
4.429952191024613,
4.449633456195267,
4.469117750431203,
4.488410170391926,
4.50751567800373,
4.526439104022458,
4.545185151502058,
4.56375839917141,
4.582163304721865,
4.60040420800785,
4.618485334162838,
4.636410796632918,
4.65418460013016,
4.671810643507866,
4.69803376208578,
4.723941275714475,
4.749545326895599,
4.774857576696206,
4.799889223844652,
4.824651023069055,
4.849153302708363,
4.873405981624883,
4.897418585445984,
4.9212002621615705,
4.944759797102883,
4.968105627327176,
4.991245855431829,
5.014188262820515,
5.036940322443186,
5.070793655324475,
5.104250360964285,
5.137332778739954,
5.170061919878847,
5.202457546478753,
5.234538245826789,
5.266321500296547,
5.297823753086552,
5.329060470047477,
5.375539061722464,
5.421483712565737,
5.466937855675816,
5.511941052856708,
5.556529340108732,
5.600735542285761,
5.6665165683630985,
5.731566461259307,
5.795968957583348,
5.859796608997625,
5.9231122794880084,
5.985970442196456,
6.048418302648519,
6.110496771616005,
6.172241307743934,
6.233682647376258,
6.294847436680899,
6.355758779153149,
6.416436709825621,
6.476898605996451,
6.537159542973938,
6.597232602198237,
6.657129138115283,
6.716859009324753,
6.776430778784639,
6.835851887214792,
6.895128803287238,
6.954267153710803,
7.01327183590156,
7.0721471155703135,
7.130896711246255,
7.189523867485624,
7.248031418280107,
7.306421841976903,
7.3646973088468,
7.4519409290032685,
7.538933941394881,
7.625681313752006,
7.712187135395744,
7.798454793312201,
7.884487112871353,
7.970286470289965,
8.055854882512447,
8.141194079044189,
8.226305559361357,
8.311190638793455,
8.395850485193332,
8.480286148244558,
8.564498582884594,
8.648488668025333,
8.732257221515285,
8.815805012098124,
8.899132768970714,
8.982241189422648,
9.065130944942544,
9.147802686098961,
9.230257046442008,
9.312494645622275,
9.39451609188427,
9.476321984059934,
9.557912913162648,
9.639289463661916,
9.720452214502885,
9.801401739921902,
9.882138610099082,
9.962663391680604,
10.0429766481969,
10.123078940397633,
10.24291676958145,
10.362282435955265,
10.481177808649084,
10.599604746742758,
10.71756510011468,
10.835060710043434,
10.95209340963677,
11.068665024139886,
11.184777371159337,
11.300432260827948,
11.415631495928473,
11.53037687198838,
11.64467017735445,
11.758513193253208,
11.871907693841438,
11.984855446249743,
12.097358210621188,
12.209417740146502,
12.321035781096823,
12.432214072854697,
12.542954347943843,
12.653258332057995,
12.763127744089093,
12.872564296154973,
12.981569693626673,
13.090145635155459,
13.198293812699594,
13.306015911550931,
13.413313610361314,
13.520188581168851,
13.679869443551631,
13.838606647691629,
13.9964057702808,
14.153272355054504,
14.309211912986395,
14.464229922482108,
14.618331829571762,
14.771523048101308,
14.923808959922743,
15.075194915083184,
15.225686232012832,
15.375288197711814,
15.52400606793593,
15.671845067381296,
15.81881038986789,
15.964907198522027,
16.110140625957744,
16.25451577445712,
16.39803771614952,
16.5407114931898,
16.68254211793543,
16.8235345731226,
16.963693812041264,
17.10302475870916,
17.241532308044796,
17.379221326039417,
17.584534311872225,
17.78802730542561,
17.989716439979624,
18.18961770580121,
18.387746951411966,
18.58411988484463,
18.778752074888434,
18.97165895232345,
19.16285581114393,
19.352357809770886,
19.540179972253835,
19.726337189461955,
19.910844220264657,
20.09371569270169,
20.274966105142873,
20.45460982743756,
20.63266110205391,
20.809134045208037,
20.984042647983184,
21.157400777438955,
21.329222177710722,
21.499520471099284,
21.752703503176647,
22.00252003243546,
22.24901482230688,
22.492232041015445,
22.732215269493366,
22.969007509189584,
23.202651189774997,
23.433188176745226,
23.660659778922295,
23.885106755856565,
24.106569325130238,
24.32508716956376,
24.540699444326382,
24.75344478395221,
24.963361309262925,
25.17048663419848,
25.374857872556948,
25.576511644644793,
25.775484083838663,
25.971810843059966,
26.165527101163352,
26.356667569240237,
26.54526649683852,
26.73135767809959,
27.006782847670838,
27.27671464194327,
27.54126262669194,
27.800534182394447,
28.054634547816857,
28.30366686273029,
28.547732209775525,
28.78692965549264,
29.021356290532253,
29.251107269064818,
29.476275847403844,
29.696953421858794,
29.913229565833007,
30.12519206618169,
30.332926958844762,
30.536518563768976,
30.736049519133537,
30.931600814893077,
31.123251825651618,
31.311080342880853,
31.495162606495846,
31.67557333580093,
31.852385759818418,
32.02567164701237,
32.19550133441955,
32.36194375619934,
32.52506647161422,
32.68493569245222,
32.841616309902385,
32.99517192089419,
33.14566485391168,
33.293156194292706,
33.43770580902353,
33.57937237103897,
33.718213383037856,
33.854285200823426,
33.987643056178285,
34.11834107928307,
34.24643232068796,
34.37196877284602,
34.49500139121699,
34.67586947681684,
34.851326421477815,
35.021534113606926,
35.1866495982969,
35.346825222226556,
35.502208774226084,
35.65294362163704,
35.799168842592685,
35.94101935434086,
36.07862603772774,
36.2121158579573,
36.341611981738026,
36.46723389092476,
36.58909749276077,
36.70731522682156,
36.82199616875919,
36.93324613094287,
37.041167760088555,
37.145860631967714,
37.2474213432827,
37.39520472954862,
37.53635612712907,
37.671173155905535,
37.7999400796464,
37.92292840537991,
38.04039745587463,
38.15259491641648,
38.25975735707869,
38.362110731480634,
38.45987085334279,
38.553243851221424,
38.642426603891856,
38.770197428495464,
38.88936741166412,
39.000515497600844,
39.10418175276765,
39.20086950086595,
39.291049794955995,
39.37515530202304,
39.492848225170285
],
"yaxis": "y"
}
],
"layout": {
"autosize": true,
"legend": {
"title": {
"text": "Chemical"
},
"tracegroupgap": 0
},
"template": {
"data": {
"bar": [
{
"error_x": {
"color": "#2a3f5f"
},
"error_y": {
"color": "#2a3f5f"
},
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "bar"
}
],
"barpolar": [
{
"marker": {
"line": {
"color": "#E5ECF6",
"width": 0.5
},
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "barpolar"
}
],
"carpet": [
{
"aaxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"baxis": {
"endlinecolor": "#2a3f5f",
"gridcolor": "white",
"linecolor": "white",
"minorgridcolor": "white",
"startlinecolor": "#2a3f5f"
},
"type": "carpet"
}
],
"choropleth": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "choropleth"
}
],
"contour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "contour"
}
],
"contourcarpet": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "contourcarpet"
}
],
"heatmap": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmap"
}
],
"heatmapgl": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "heatmapgl"
}
],
"histogram": [
{
"marker": {
"pattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
}
},
"type": "histogram"
}
],
"histogram2d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2d"
}
],
"histogram2dcontour": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "histogram2dcontour"
}
],
"mesh3d": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"type": "mesh3d"
}
],
"parcoords": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "parcoords"
}
],
"pie": [
{
"automargin": true,
"type": "pie"
}
],
"scatter": [
{
"fillpattern": {
"fillmode": "overlay",
"size": 10,
"solidity": 0.2
},
"type": "scatter"
}
],
"scatter3d": [
{
"line": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatter3d"
}
],
"scattercarpet": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattercarpet"
}
],
"scattergeo": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergeo"
}
],
"scattergl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattergl"
}
],
"scattermapbox": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scattermapbox"
}
],
"scatterpolar": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolar"
}
],
"scatterpolargl": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterpolargl"
}
],
"scatterternary": [
{
"marker": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"type": "scatterternary"
}
],
"surface": [
{
"colorbar": {
"outlinewidth": 0,
"ticks": ""
},
"colorscale": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"type": "surface"
}
],
"table": [
{
"cells": {
"fill": {
"color": "#EBF0F8"
},
"line": {
"color": "white"
}
},
"header": {
"fill": {
"color": "#C8D4E3"
},
"line": {
"color": "white"
}
},
"type": "table"
}
]
},
"layout": {
"annotationdefaults": {
"arrowcolor": "#2a3f5f",
"arrowhead": 0,
"arrowwidth": 1
},
"autotypenumbers": "strict",
"coloraxis": {
"colorbar": {
"outlinewidth": 0,
"ticks": ""
}
},
"colorscale": {
"diverging": [
[
0,
"#8e0152"
],
[
0.1,
"#c51b7d"
],
[
0.2,
"#de77ae"
],
[
0.3,
"#f1b6da"
],
[
0.4,
"#fde0ef"
],
[
0.5,
"#f7f7f7"
],
[
0.6,
"#e6f5d0"
],
[
0.7,
"#b8e186"
],
[
0.8,
"#7fbc41"
],
[
0.9,
"#4d9221"
],
[
1,
"#276419"
]
],
"sequential": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
],
"sequentialminus": [
[
0,
"#0d0887"
],
[
0.1111111111111111,
"#46039f"
],
[
0.2222222222222222,
"#7201a8"
],
[
0.3333333333333333,
"#9c179e"
],
[
0.4444444444444444,
"#bd3786"
],
[
0.5555555555555556,
"#d8576b"
],
[
0.6666666666666666,
"#ed7953"
],
[
0.7777777777777778,
"#fb9f3a"
],
[
0.8888888888888888,
"#fdca26"
],
[
1,
"#f0f921"
]
]
},
"colorway": [
"#636efa",
"#EF553B",
"#00cc96",
"#ab63fa",
"#FFA15A",
"#19d3f3",
"#FF6692",
"#B6E880",
"#FF97FF",
"#FECB52"
],
"font": {
"color": "#2a3f5f"
},
"geo": {
"bgcolor": "white",
"lakecolor": "white",
"landcolor": "#E5ECF6",
"showlakes": true,
"showland": true,
"subunitcolor": "white"
},
"hoverlabel": {
"align": "left"
},
"hovermode": "closest",
"mapbox": {
"style": "light"
},
"paper_bgcolor": "white",
"plot_bgcolor": "#E5ECF6",
"polar": {
"angularaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"radialaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"scene": {
"xaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"yaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
},
"zaxis": {
"backgroundcolor": "#E5ECF6",
"gridcolor": "white",
"gridwidth": 2,
"linecolor": "white",
"showbackground": true,
"ticks": "",
"zerolinecolor": "white"
}
},
"shapedefaults": {
"line": {
"color": "#2a3f5f"
}
},
"ternary": {
"aaxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"baxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
},
"bgcolor": "#E5ECF6",
"caxis": {
"gridcolor": "white",
"linecolor": "white",
"ticks": ""
}
},
"title": {
"x": 0.05
},
"xaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
},
"yaxis": {
"automargin": true,
"gridcolor": "white",
"linecolor": "white",
"ticks": "",
"title": {
"standoff": 15
},
"zerolinecolor": "white",
"zerolinewidth": 2
}
}
},
"title": {
"text": "Same plot as above, both only showing initial detail"
},
"xaxis": {
"anchor": "y",
"domain": [
0,
1
],
"range": [
0,
0.3
],
"title": {
"text": "SYSTEM TIME"
},
"type": "linear"
},
"yaxis": {
"anchor": "x",
"autorange": true,
"domain": [
0,
1
],
"range": [
-2.7777777777777777,
52.77777777777778
],
"title": {
"text": "Concentration"
},
"type": "linear"
}
}
},
"image/png": "iVBORw0KGgoAAAANSUhEUgAAAu8AAAFoCAYAAADn3hWsAAAgAElEQVR4Xu29C7QdxXmgW3rzkM4REg+DECiE+EaWwY69wusKYq69QOM1gOzBwKwZbI0BD8yYO0bMDWBlEcKKwGQuwmvkDFxAjGwyd3gtLMNdE5mVXDOGACbXiY2syIkJEQ9hg0HogQV63/1vqY77tLp3V1X/1bX77K/X0pJ0TtVf1V/V3vvr2n9Xj9vbOQwHBCAAAQhAAAIQgAAEIND3BMYh730/RnQQAhCAAAQgAAEIQAACXQLIOxMBAhCAAAQgAAEIQAACLSGAvLdkoOgmBCAAAQhAAAIQgAAEkHfmAAQgAAEIQAACEIAABFpCAHlvyUDRTQhAAAIQgAAEIAABCCDvzAEIQAACEIAABCAAAQi0hADy3pKBopsQgAAEIAABCEAAAhBA3pkDEIAABCAAAQhAAAIQaAkB5L0lA0U3IQABCEAAAhCAAAQggLwzByAAAQhAAAIQgAAEINASAsh7SwaKbkIAAhCAAAQgAAEIQAB5Zw5AAAIQgAAEIAABCECgJQSQ95YMFN2EAAQgAAEIQAACEIAA8s4cgAAEIAABCEAAAhCAQEsIIO8tGSi6CQEIQAACEIAABCAAAeSdOQABCEAAAhCAAAQgAIGWEEDeWzJQdBMCEIAABCAAAQhAAALIO3MAAhCAAAQgAAEIQAACLSGAvLdkoOgmBCAAAQhAAAIQgAAEkHfmAAQgAAEIQAACEIAABFpCAHlvyUDRTQhAAAIQgAAEIAABCCDvzAEIQAACEIAABCAAAQi0hADy3pKBopsQgAAEIAABCEAAAhBA3pkDEIAABCAAAQhAAAIQaAkB5L0lA0U3IQABCEAAAhCAAAQggLwzByAAAQhAAAIQgAAEINASAsh7SwaKbkIAAhCAAAQgAAEIQAB5Zw5AAAIQgAAEIAABCECgJQSQ95YMFN2EAAQgAAEIQAACEIAA8s4cgAAEIAABCEAAAhCAQEsIIO8tGSi6CQEIQAACEIAABCAAAeSdOQABCEAAAhCAAAQgAIGWEEDeWzJQdBMCEIAABCAAAQhAAALIO3MAAhCAAAQgAAEIQAACLSGAvLdkoOgmBCAAAQhAAAIQgAAEkHfmAAQgAAEIQAACEIAABFpCAHnv84H65EWLzQeOnGH+2zf+oM97qt+9sXju8z6xyPyz/+1U83/eeJU+sFzEbz38XXPbn/53c8cf/Xtzzu/9bvT2XBv4T3c+YFY+uNo8dPdNZt4H57hWq12uSfa1O0sAozF//+PNd5o//39/YNY+ubIviGqcU1+cCJ2AAASSEuh7eX/if/61ueYP//QASIsuXmD+j6suSQqvicbrCGyduk2cW1Ubbe9/0fnFEMiymP0qCoMu72v/Yb256Es3mY9++MQkF+V12/9XX/5j86OfvDjq4qvstWrnYMj7tcb8rSPvdd9/ijhrnFPV+ya/hwAExj6BvpZ3+8Z73b//l+bznzt3ZDTsh39TK5gpp0GdD5A6dVOes2277f1H3otnEfKOvLu8v2iILvLuQpoyEIBA2wj0rbzbFfdeKzbyxtxE+kHKQa0jsHXqpjxn5N2PPivvbrxifOvh1vLYLxXjvabt8l406hrnNPZnE2cIAQhUEeh7ec+vupedkGt6jf0qUy4K3vjlO918SHvYtuzXwvbnZTnDIgPZw7Wv2ZXH//0P/rP5xZsbR8Lk84DLPhTzfcx/CyH1snFtA71yPy2bPOOibziKyrp+E5LnJu0VpRDYc/9C51uXbOpUWTv5c85f+Em7ZakKUleOv3xo2cjp21U7+wONNAcrkCfPPaGbj26PsotUl3PKj5fcIyHnkRWFb3by3yXVoWpOF72+isY6/5qwrz95Dbyw7qXC15WNnV95t3O5aG7aMXDJj8/Pq/x4lbEvm09Vr7GibxCy7y/ZtL786zh/IeE7VkVjIucrY+yS312n/fxqdq/3muy8sN+e2nPNz7X8+6ev6BYxkdeCvA/mmdixs32wrxn7/17n5PM+2YtzP92HUiUK/B4CEOgvAn0r79k3SJcPI/mQEBnKylfR6n02bvbDIvtmXvTzrDzYuNkPfZdvCvLyIv/Pxi2SmCJ5L5LQop/5roZZNlne9mdV52o/aF0uYKSveRkr67988GYlrKg/rj8rS9ewY5cVUvvhnWVRJPi+L2crmFU8Xc9J2q9aeZcy2XPzSSUoGtein2UvnqteP/lxKHvtlIlwnrnrHO3FPj9vXV5jRWJqzy0rg0XnUSZ1LmNVxqtozpbNzzrtF82fsveaMnn/buf9OnsTfq955nLDdRmTovfUopx91/de4en6Pln02vS9IPF9f6E8BCAwGAT6Vt4Ff9kKjcubuR0+eVOWw35QlAlB0YeM1Cv6edkHlf3grrrYKJPIor7l26oS0KyE+Mp72ZSXD+u//cnPRi6MevX/hz/++1H3J7i+jIrYVXG2FwBlMpr/oCySPOmfz/nJjYYuFyiu0mTL5c/B9Zxc5L3XKnn2XpKiPpeNQb5/Za+fojldNH/kdSoXatmLb9fXUy8hkhh2BbzsIqfOaywf077fZG/oLLo4dF2R9Xn/8bkoq9N+XXkve23IOPzOh39rJBXSR3Rd52mvmMIk+w2Y7/tn/n0EeXd996ccBCDgS6Cv5T17MkWpFkUSVfR1Z9UqWHY1JZ++kJePXquB9oOh6iv+Xjfs5T8w8v8vkhzLKf+B7Pvhk73gyaZY2J/bi5LsRVXVhUqvC4JsypItl2XnuppXVq5orPLiUVSminHIzhllY2R/npcKn3OqWnkPlXeXuW5j15X3IqnKy1zZXMqu+vd67bnKe9X4Z781yQqb5SV9kHS4BWef0r1wkNf76u89P+rCJFSee41J2+S9Kn3OVd59mPRilJ9vVe+f+bSq/Psk8u6rI5SHAARcCbRG3vMnZN/4rTxaIc7nueY/iMve6F1/XpZbn+1f1TcDdeS91wdK/qv+qg+fPFP7YZnP/yz6wCv6VsQlJzzLLyv+RUxc5b1MynqlU9gLv7IV/6L7BSwv19z+ohehq2j7nJNrTNufMtHO97dXuXyaQ115l7az4+16IZw/p+w55Oexq7z7vMay/fwff/mc+dHaF7vf8mWFXd6Djj5y5qib60PlvdeYtEXebT+L7tPJPtPCVd59mJQJt5032fewsnng8z7pOs6uH9iUgwAEICAEWivv+RvZyt5oteXdCmGd1Ile8l61eu6zKugr72WxXaTAlqlalS6L5SPvdVap85JYtLqbT7XSfKsoE8j8+Q/ayrswzorwLf/5z7rYQx9OVrSdrKu8+7zGsu8Hksf90XkndlfbrVDKKnxRqpWr1OXF1EdUe83b0PYlZt20GddxiCHvRaktZZxcP1Ns/SIurpw132OIBQEIjH0CfSvv8sYtqzBld+TnP2DLPhC05T0vfyFTpE7euk9dXwl1zRst26KzbAyyjMrEyEfe6+SHZyVRLjTkSZ/5b0pi7kPuOk99ct7L0kvK5Md15b3XXNfOebdzxPKRlKqqb7BsHRmvT3/ytAOe1pqfz67S6PMayzIqe3BR/udSx1Xqisaq7DxcLrLznO1Wuz5zpaidsveafP97pbjkx8tV3ot4lgm1T8yyc3J9n/QZ55DPEepAAAKDS6Cv5d1upZfPq7YfHtnV7167CmjmvMtUKdptJvuhWJUHbuUgLyf5tJcyeZJyRSkB+bQVXwnNf5uRXWWTf9vzsuWy/Xfdbabo3LM7/eRz3qXd7E2MRe1YIajalSb7MrdpV722jizaYk7OXbZ5tDd6ZncvqrrXwX6Yl41TlqfPOfmIfnb+unx7ZHlnv1HptdtMPqbrDatZEZcLKpcUrHydbNtFu4+4yrsdJ5fXWPY1Ujau+Tg+UtdrR5ui11/2ddrrY63OxUPRfPO54BH5zb+u7b1KWYY+ol31vpJ9Ty7abUb6IzHksDc4l52T6/ukzzgProJw5hCAQAiBvpX37ofQ/seI50+s6MNQyhTtzy77W2d3sXDNbbdt9lopKrrhyiUf2n4o2D2IbVtFdXt9dZu9qdR1r+qqCwv7wWT7lN23u+gDMDs2PiulImj2EDGUQ35WJO/5/POydvI3K1c94EtWd3sJbJ7FyJx4cuVI34vktkqa8uMu5cvE3/WcsuWK9nnPfoPls/Je9jp0vQnWV959+2ZZF41V0faPMV5jZReuZbsb+UhdGY/sBa/Ek/OSI7srVNU8zLKou/Je9P4r7xdl/c/Pa3uTb0jOuz3PPJPsszyq9nm3MfKvw/xnSn4Bo+p90vUiqddY8TsIQAACeQJ9Le9jdbh8V8THKoexcF4+qQpj4Xxjn0OvfPPYbbc9vm+aXNvPl/5DAAIQGFQCyHuCkUfeE0CP1KSsrFXdpBup6TEXtuxBO2PuRGuekHCSbxSLHnLk+u1XzS5QHQIQgAAEEhJA3hPAR94TQI/QZHZHkXkfnBOhhcEKWXTfymARcDvbsu1qXe65cGuBUhCAAAQg0M8EkPd+Hh36BgEIQAACEIAABCAAgQwB5J3pAAEIQAACEIAABCAAgZYQQN5bMlB0EwIQgAAEIAABCEAAAsg7cwACEIAABCAAAQhAAAItIYC8t2Sg6CYEIAABCEAAAhCAAASQd+YABCAAAQhAAAIQgAAEWkIAeW/JQNFNCEAAAhCAAAQgAAEIIO/MAQhAAAIQgAAEIAABCLSEAPLekoGimxCAAAQgAAEIQAACEEDemQMQgAAEIAABCEAAAhBoCQHkvSUDRTchAAEIQAACEIAABCCAvDMHIAABCEAAAhCAAAQg0BICyHtLBopuQgACEIAABCAAAQhAAHlnDkAAAhCAAAQgAAEIQKAlBJD3lgwU3YQABCAAAQhAAAIQgADyzhyAAAQgAAEIQAACEIBASwgg7y0ZKLoJAQhAAAIQgAAEIAAB5J05AAEIQAACEIAABCAAgZYQQN5bMlB0EwIQgAAEIAABCEAAAsg7cwACEIAABCAAAQhAAAItIYC8t2Sg6CYEIAABCEAAAhCAAASQd+YABCAAAQhAAAIQgAAEWkIAeW/JQNFNCEAAAhCAAAQgAAEIIO/MAQhAAAIQgAAEIAABCLSEAPLekoGimxCAAAQgAAEIQAACEEDemQMQgAAEIAABCEAAAhBoCQHkvSUDRTchAAEIQAACEIAABCCAvDMHIAABCEAAAhCAAAQg0BICyHtLBopuQgACEIAABCAAAQhAAHlnDkAAAhCAAAQgAAEIQKAlBJD3lgwU3YQABCAAAQhAAAIQgADyzhyAAAQgAAEIQAACEIBASwgg7y0ZKLoJAQhAAAIQgAAEIAAB5J05AAEIQAACEIAABCAAgZYQQN5bMlB0EwIQgAAEIAABCEAAAsg7cwACEIAABCAAAQhAAAItIYC8t2Sg6CYEIAABCEAAAhCAAASQd+YABCAAAQhAAAIQgAAEWkIAeW/JQNFNCEAAAhCAAAQgAAEIIO/MAQhAAAIQgAAEIAABCLSEAPLekoGimxCAAAQgAAEIQAACEEDemQMQgAAEIAABCEAAAhBoCQHkvSUDRTchAAEIQAACEIAABCCAvDMHIAABCEAAAhCAAAQg0BICyHtLBopuQgACEIAABCAAAQhAAHlnDkAAAhCAAAQgAAEIQKAlBJD3lgwU3YQABCAAAQhAAAIQgADyrjAHXn/7PYUohPAlcNDkCeaQKRPMxq07fKtSXoHAhPHjzOHDU8wb77yvEI0QIQSOmXmw4f0nhJxOnSM683/Tr3aanbv26AQkiheB6VMnmx07d5tt23d71aOwHgF5D+JongDyrsCcD08FiAEhkPcAaIpVkHdFmIGhkPdAcErVkHclkIFhkPdAcIrVkHdFmB6hkHcPWGVFkXcFiAEhkPcAaIpVkHdFmIGhkPdAcErVkHclkIFhkPdAcIrVkHdFmB6hkHcPWMi7AizFEMi7IsyAUMh7ADTlKsi7MlDPcMi7JzDl4si7MtCAcMh7ADSFKsi7AkRW3hUgBoRA3gOgKVZB3hVhBoZC3gPBKVVD3pVABoZB3gPBKVZD3hVheoRC3j1gsfKuAEsxBPKuCDMgFPIeAE25CvKuDNQzHPLuCUy5OPKuDDQgHPIeAE2hCvKuAJGVdwWIASGQ9wBoilWQd0WYgaGQ90BwStWQdyWQgWGQ90BwitWQd0WYHqGQdw9YrLwrwFIMgbwrwgwIhbwHQFOugrwrA/UMh7x7AlMujrwrAw0I14S8X7BoiZk5Y8jct+y6gB6mrbJm3UvmkqtuNg/ceaM5ae4Jap1B3muiPO0fXjSPzpxVMwrVQwgg7yHU9Oog73osQyMh76HkdOoh7zocQ6Mg76Hk9OppyPsXF99mfvA360Z1asb0aeapVcu7P0sh76tWP22WfO1es/T6y83CBfODgSHvwejiVhz3ty+YR46aZU6fckjchoh+AAHkPe2kQN7T8pfWkfe0Y4C8p+WPvKflb9+D6vRi3icWmayo21gi9Ecdfpi59atfSiLvdc4pWxd51yKpHEfk/aKpw+aOGUcqRyZcFQHkvYpQ3N8j73H5ukRH3l0oxSuDvMdj6xIZeXehFLdMnZV3EfSfvfTayAp7WU/tyrv83q7Qlwl/dgU/m6py5sKrzfxTTjJPP7/GbNy0tdvUlZeeb2bPOrK7wm4PW6dIuvPfEEj9qy/7rCn65mDtkyu7IZH3uPMvOLrI++yJk8xzx8wJjkHFMALIexg3rVrIuxbJ8DjIezg7jZrIuwbF8BjIezg7rZp15F1W3c8/54zu6nqvQ+T9xfUburItsiyHyPhvnXDsSB68CPTbG7eY76xc2v398hWPmrvuf8xYiZbyIu1Wzu3v8+k5Uldi5KU7f6Ehv7/jnoe77cvvrrnicyM57dLfsjha3Ml5r0nyd/7+Z+ZH294zK444xiw4+NCa0ajuQwB596GlXxZ512fqGxF59yWmWx551+XpGw159yWmXz5U3q0cu+SUF+W833DL3ebv/uHlQtG2ZynCftF5Z3eF36682wuFohVxiSkr85Jrn/29xJObTl36ai8cHnr8ewfE4YZV/fkXHHHl2xvNv3nlNbPgkKlmxeFHB8ehoj8B5N2fmWYN5F2TZlgs5D2Mm1Yt5F2LZFgc5D2Mm2atfpB3e3Np0XnZ1foyec8KuazGF0n3P778eje1xq7iF7VjV/azv5PypM1ozjbFWOt37DC/sfan3YiSOiMpNBzNEEDem+Fc1grynpa/tI68px0D5D0tf+Q9LX/7HhTaC5+0mfxWkdmVdyvvVXItOe/5lXcNeZfzOPVjc0dSeLIpO8h76OxooN4Z6140z76/rXvTqty8ytEMAeS9Gc7Ie1rOvVpH3tOODfKelj/ynpZ/XXmvumFVBL1st5mitJleaS11Vt7lPMvSZoouHJD39PPSqQdff/kX5pqNb5rTDzrEPHIke747QVMohLwrQKwRgpX3GvCUqiLvSiADwyDvgeCUqiHvSiBrhAlNm7FNFm0VaYXY3sxalfMuseyOL9nVdxH8Uz/2oe4+7XXkXXLVpQ8bN20Z2RnH3rAqN6rmxV7OSQ7SZmpMrCaqvv72e2bua/9otuzZQ+pME8D3t4G8Nwi7oCnkPS1/u+ol7z8caQgg72m421aR97T87XtQ3V4UbbWYXUV3kfeswGf7k91tJjRtxt5oane9sfFtH+Ui4bEnnhlpVvLs7U43pM3UnR0R68uHp6y8P/TuZvZ8j8g5Hxp5bxA28p4WdknrrLynHRbkPS1/5D0tfy15T38W7esBW0UqjJnI+7Pbt5kL39jAnu8KPF1DIO+upOKUY+U9DlefqMi7Dy39ssi7PlOfiMi7D604ZeumzcTp1diPirwrjLH92vq019ebV3ftZM93BaYuIZB3F0rxyiDv8di6RkbeXUnFKYe8x+HqGhV5dyUVrxzyHo9tr8jIuwJ3K++SNiPpM7LjjOw8wxGXAPIel29VdOS9ilD83yPv8Rn3agF5T8sfeU/LX1pH3tOMAfKuwN3Ku6y6y+q7HOz5rgC2IgTyHp9xrxaQ97T87QcnN6ymGwfkPR17aRl5T8sfeU/HH3lXYJ/98LzsrZ+b1dveZc93Ba5VIZD3KkJxf4+8x+XrEp2VdxdK8cog7/HYukRG3l0oxS3DyntcvmXRkXcF7ll5X/3er8xlv3ydG1cVuFaFQN6rCMX9PfIel69LdOTdhVK8Msh7PLYukZF3F0pxyyDvcfki7xH55r+2tjeuPnLULHP6lEMitjzYoZH3tOOPvKflL60j72nHAHlPyx95T8vfvgel78Xg9YCVd4Uxz8v7PVveMTdteosbVxXY9gqBvEcGXBEeeU/LH3lPzx95TzsGyHta/sh7Ov7IuwL7vLxz46oCVIcQyLsDpIhFkPeIcB1Ds/LuCCpSMeQ9EljHsMi7I6iIxUibiQi3R2jkXYF70W4P9omr1w7PNIuHZyi0Qog8AeQ97ZxA3tPyZ+U9PX/kPe0YIO9p+Y/1lfd5n1hkTpwzy3xn5dL0oHM9QN4VhqRI3nniqgLYihDIe3zGvVpA3tPyR97T80fe044B8p6W/1iW9+UrHjV/8dQPzcZNW8x/ufUac9LcE9LDzvQAeVcYjrJ9lu2Nq/LAJnlwE4cuAeRdl6dvNOTdl5h+edJm9Jn6RETefWjpl0Xe9Zn6RhyraTMXLFpiPnXmx83frv2ZOerww8ytX/2SL5qo5ZF3Bbxl8m6fuHr6QYeYR46cpdASIbIEkPe08wF5T8uflff0/JH3tGOAvKflr7Hyvn7HDrN++47GT2TOlMlmzuTJhe2uWfeSueSqm80Dd95o/vHl183tdz1onlq1vPE+9moQeVcYjjJ537JnjznnF68YuYGVJ64qgM6FQN71mfpERN59aMUpy8p7HK6uUZF3V1JxyiHvcbj6RK278v71N39prtnwc58mVcp+5cgjzB2zji6MZVNmbK675L6LyPdT6gzyrjANej2e3N64Kmkzkj7DoUcAeddjGRIJeQ+hplsHedfl6RsNefclplseedflGRKtrryvfHuj+ebGTSFN16rzhRnTzaKZxZuJ2JSZqy/7bLeNLy6+re9SZ5D3WsO/r3IveWfbSAXAJSGQ93hsXSIj7y6U4pZB3uPyrYqOvFcRivt75D0uX5fodeXdpY0my9iUmXybM6ZP66vUGeRdYVb0kncJb1ffb5p+uLli6DCFFgkhBJD3tPMAeU/LX1pH3tOOAfKelj/ynpa/fQ9K3wu9HuRTZmxkSZ1Zev3lZuGC+XqN1YiEvNeAZ6tWyTvbRipALgiBvMfh6hoVeXclFa8c8h6PrUtk5N2FUrwyyHs8tq6Rx9rK+5kLrzYXnXe2sSkzloOkzshx37LrXNFELYe8Z/DecMvd5rEnnjngxgTJf3px/YZuyaIN+6vkXeqxbaT+PEbe9Zn6RETefWjFKYu8x+HqGhV5dyUVpxzyHoerT9SxJu8+556yLPK+n/6q1U+b//rAn3clPXtXsVxtvb1xy8gTtkTkZ84YGnX15SLvdtvI2RMndXee4ahPAHmvz7BOBOS9Dj2dusi7DsfQKMh7KDmdesi7Dsc6UZD3OvTC6yLv+9nZrYDs3p52SyD5CuXaKy8eyXMSyc/v+eki79KMXX1fccQxZsHBh4aPGjW7BJD3tBMBeU/LX1pH3tOOAfKelj/ynpa/fQ9K34vB6wHy3hlzWU3/N5f8M/Obxx8zsjG/yHt2o34r80U/e+Od951mzoNbN5v/8PYb5oyDDzGPHnWsUx0KlROYMnmCOXjyeLPp3Z1gSkBA5P2waZPNW5u3J2idJoXAUYcdZFzffyCmT2Dm0GSzZdsus3PXHv3gRKwkMHTopC7797bvrixLgTgE5D2Io3kCAy/vkuf+xlvvdNNg8mLuKu+79+x1Gjl5ktin/vGfjPz9ww+eaD5y8MFO9ShUTGBc58fjxo0ze/a68YejPgEReNf5r986EeGfdg6M5/0n6QAI/72d938+AdINg7wHcTRPIIm8SyrKxk1bC8927ZMrG6OQT4EJlXfXtBk5MR7apDe8pM3osQyJRNpMCDXdOqTN6PL0jUbajC8x3fKkzejyDIlGznsItfp1Gpf3ohs+659GWASR9yVfu7ew8pWXnt/dKqgo513qZC8yfOSdhzaFjVVRLeRdj2VIJOQ9hJpuHeRdl6dvNOTdl5hueeRdl2dINOQ9hFr9Oo3Le79tdJ9FWJQmo7XbTLYdVt/rT1yJgLzrcAyNgryHktOrh7zrsQyJhLyHUNOrg7zrsQyNhLyHkqtXD3nP8CuSd/m1xj7v2WFi9b3epLW1kXcdjqFRkPdQcnr1kHc9liGRkPcQanp1kHc9lqGRkPdQcvXqNS7vIsKfOvPjBzy9qt5ppK3tkzZje8rqe/0xQ97rM6wToa68b9m+2Wzevqm0C1t2bDZbevy+Tt816x477XivcLOH/Mr3Co68e6FXL4y8qyP1Coi8e+GKUhh5j4K1Mmjj8l60T3plL/u8QIi8s/pef1CR9/oMJUJWol/b+vJI0Fe37Pv35o5kb+2ItD1e3fpK95+djR7ML371qtmR2SZPZFukO3vYODq9HYwoVYI/NHnYDE2ZbqZMGm+27xy9TeHwlH2/KztmTzuu8FfTOjGlbtFR1p+yC5eq/o+VUUTe044k8p6Wv7Q+1uTdZmDkyS69/vKR5/2kp975/O1ss9ToLkuS897raHK3Ga0BCJF3aZvV93ojgLz/mp8I+Ksd8bYr1SLMr+2XbCvbVqKtYKeW6qGOKA73kEwrqPVmSfza2Qsel9ZSc3fpY6wyPhcBZRch3Z93LjTyx7EFFyVyEVN0QVJ00SFzUeakz4G8+9DSL4u86zP1jThW5f2BO2809vk+sqX408+vMU+tWu6LJ1r5xuU92pkkDBwq76y+1xu0sS7vWSG3Mi6r2vJvWQ0XCd8n7KNXun2pZiU6KzVWtPKyZCVJ9lj+8DEnmne27hhpUmQpL1aDsgrry71X+cEyn1MAACAASURBVCrBtylFM4emmLe3jH5Ilp0bRfHz36Jky9iLvKJ6Zf0pu3Cp6r8mq9ixiuavvbCcPHGc2bV7b+dZE6Z7kVD0jUfRxUbZtxxFbWldaMTmlCI+8p6C+ug2B0He7c6E/bS4jLwrzP1QeZemWX0PH4C2y/vat17YJ+D7V8lFnuTf8jP5nc9hBdx+0IsEWGmwomDFwAp2yEpjtk91c959zo+yxQTalvPucxFQdhFSdgFSdPEhryUpnz+KLjrk/ou6F8Kp52mvC41s38ouNIpSqoq+0ZBY+baKLtyLymkyQt41aYbFqivv6zetN/Kn6WPO9DlG/uSPsl0HpZw8zLNfjiTyXrS/er/lE/kMUB15Z/Xdh/Tosv0u73blXITl7zoybuVcxMFlZVKEfHbnZkj5UJQPSvlgzYq4/bBMtbKNvIfPXa2abZN3rfNuMk7Ra9V+8zF86GTz7vu7zO7de0a+Dcv3rehiQ+oXXVQUtTUWLzTKUuaKvmUoen+zCxPyGSBPeN7Zue+m7NuMotSpFBcaTc7ZJtuqK+9ff+7r5prvXtNkl7ttfeW0r5g7zr2jVN7zv7DP/mm8oyUNNi7vy1c8au66/zGTzSeyVzr9Bsd1kOrIu7TB6rsr6f6Ud5H0tW/92Pzkly+YDe++0v3bZfVcPpTkw8qKuaxwyb+7st6Vdr/82zCK4bWQ93B2WjWRdy2SYXH6Lee914VG9gzLvtWw98lky7p+o1F0s7rEcVmoCKPfTK0Y32aMpbSpuvK+8kcrzTd//M1mBjPTyhc+8gWz6KOLSuU966ikzXQwyRNLLzrv7AO2ihSpf+jx7/XVDQGus6muvLP67ko6vbxLOotdSbf/7pXiMu/wk/fJeefPrKnHmQ8fcXJXzuXnbT+Q9/QjiLynHYN+k/e0NNxbL9smtuhbhiL5t99mZFfey77NKEqdGqv3atS90PC5P8N+o/EvTj7XfeBbULLseT+y2UpW6FOfSuMr72VPWO3HKxvXwakr79IOq++utH9dLnbajHxoiJhLysszG57qrq6X5cSePuusbo75acecOSLrY0HQe40K8u4/Z7VrIO/aRP3iIe9+vLRLN5HzHuPbjLGUNrX3DxvdsFB7CjmtvNuMkYG+YZWV9+K5x+q7/2tSW95F1J957fvmuddF1PetsOcPSWWZd/hHzBmzzjQf2r+qPtYlvWxkkHf/OatdA3nXJuoXD3n346Vdugl51+5zzHh1LzR87s+w32g8c/n/jHlKjccu2+e9n8RdoDS+8k7Oe/lcZPXd73VaV96zsv7Mhu8fsKqeF3WbAuPXy7FbGnlPP7bIe9oxQN7T8kfe0/KX1uvmvKc/g3b2oHF5F0zsNlO++v65NzcYWYV/5KhZ5vQph7RzVjXUa195l1WJZzuS/t1/+n8KV9YlX1DSX04/Zr6Zd8RHxkReesyhQN5j0nWLjby7cYpVCnmPRdYtLvLuxilmKeQ9Jt3y2EnkPc2pxmtVI+fd9m7Z5o3m9s1vm9kTJ5nnjpkTr9NjILKLvFtZl3SY/I2lWVkXaU+15WJbhwJ5Tz9yyHvaMUDe0/JH3tPyZ+U9HX/kXYG9prxLd057fX139f2OGUeai6b293aBCviCQ5TJ++qXHu+urou4Z3MAJQ3mjI6kn/sb/7y7wo6sB6PvVkTe6/HTqI28a1AMj4G8h7PTqIm8a1CsF4OV93r8Qms3Ju+yy4zs4y57vPc6+u2mABew2vL+0Lubu7vPsPrem35W3q2wr37psVG56yLoC044b0TYXcaTMm4EkHc3TjFLIe8x6VbHRt6rGcUsgbzHpOsWG3l346RdqjF51+54P8XTlnc5N7v6fu3wTLN4eEY/nW7f9OWHb/yVefjv/8w8uf7JUSvscmPp5377X5szjj2LvPWIo4W8R4TrGBp5dwQVqRjyHgmsY1jk3RFUxGLIe0S4PUI3Lu9l+7wP8kOaisbn2e3bzIVvbDBD48ebJz7QefJmJweeY9/T+la88KdGVtqzKTFW2GWVnXSYZmYK8t4M516tIO9pxwB5T8sfeU/LX1pH3tOMQd/I+6A/pKlo+Nk68tdUHlp3v3nop/+tm8duj+M6KTHn/y8Lzb+eeyXCnuD9A3lPAD3XJPKedgyQ97T8kfe0/JH3dPz7Rt5vuOVu8/Tza8xTq5anoxHYcoy0GelK9sFNTxx9nJk3aUpgD9tZrWyV/aK5l5qLfvtfmbN/42xzyJQJZuPWHe08wZb3GnlPP4DIe9oxQN7T8kfe0/JH3tPxb0Tei/Z1LzrlpddfbhYumJ+ORmDLseRdujOIW0fKlo73/vhPjay220NSYRb/7ldH7RLjslVk4JBSzYEA8u4AKXIR5D0y4IrwyHta/sh7Wv7Iezr+jch79vTKct7TIajfckx5l94NytaRRakxdpVdtnbMH8h7/blbJwLyXoeeTl3kXYdjaBTkPZScTj3kXYdjnShjNeddXDV7nDhnlvnOyqV1UKnWbVzeVXvfJ8Fiy3t268iHj5w15m5eFWlf9te3jNyAah+eJCvtvW4+Rd7TvgCQ97T87apX7Pef9GfZvz1A3tOODfKelr99D0rfC90enLnwajP/lJPMrV/90kjgCxYtQd51MaeP1sSH51i8ebVI2kXYF5xwvpEHKlUdyHsVobi/R97j8nWJzsq7C6V4ZZD3eGxdIiPvLpTilhlrK+9r1r1kLrnqZvPAnTeak+aeEBdejeiNr7xbMGV95iFNxWSyN6+uOOIYs+DgQ2sMe9qqZdIuKTI+B/LuQ0u/LPKuz9Q3IvLuS0y3PPKuy9M3GvLuS0y/fG15X7/eGPnT9DFnjjHyp+CQlfcZ04f6aqU9383G5d1+HXHqxz5kbr/rwZHdZeQriU+d+XFz9WWfbXoIa7fXxMq7dLLtN6/KjajX/MWXjPwth70J1Vfa7YAh77Wnbq0AyHstfCqVkXcVjMFBkPdgdCoVkXcVjLWC1Jb3r3/dmGuuqdWHoMpf+Yoxd9xRWjWf837lpef3lZ82Lu/2htXfPP4Y8+9uuGNE3mVHmqzMBw1GokpNybucXhufvCpbPkpOu909pq60I++JJnquWeQ9/Tgg72nHAHlPyx95T8tfWq8t7ytXGvPNbzZ/Il/4gjGLFjm1Kw8Rvev+x0w/7YiYTN5lS0gReZsmw0OanObQqL3fnztmTl/fvLpl++bOlo/fMLc/v+8Obcljv+IjXzaXd/645LRXEWHlvYpQ3N8j73H5ukRH3l0oxSuDvMdj6xIZeXehFLdMbXmP2z216JI1ctF5Z/fN6nvj8i7pMR/64PHdu3iz/+YhTe5zrA3pM/m8dkmNqdo9xp3AvpLIuy8x3fLIuy7PkGjIewg1vTrIux7LkEjIewg13TpjTd5lIfm/PvDno/Ld+3FxuXF5z0+bbF5Rv9/dWzblm0ybkT5s2bPHnPOLV7qr8NcOzzSLh2fovhprRCtKkXl44eqeWz6GNoe8h5LTqYe863CsEwV5r0Ovfl3kvT7DOhGQ9zr0dOqONXkXKrKw/OL6DaMA9dtmKsnlXWf6pI3StLzL2T77/jZz4Zv7Jle/pM8s66TH3NNJk5F0Ga289l4ji7ynnffIe1r+0jrynnYMkPe0/JH3tPzte1D6XgxeDxqXd56wqjfJbtr0lrlnyzvdvHcR+FSHrLZ/btWCkYcsxUiRKTo35D3ViO9rF3lPyx95T88feU87Bsh7Wv7Iezr+yLsC+xQr79LtfkifkdV2e0NqE6vt2eFC3hUmb40QyHsNeEpVWXlXAhkYBnkPBKdUDXlXAlkjzFhMm6mBo7Gqjct7m/dzLxuVVPIu/ck+vOmRI2eZ0w86pJHJI6vt1/zlvzXPbvh+t72mVtuR90aG16kR5N0JU9RCyHtUvJXBkfdKRFELIO9R8ToFR96dMKkXalze5Qmr2f3d1c8oQcCU8i6nm9195uGOwEsaTcwju5NM06vtyHvMkfWLjbz78YpRGnmPQdU9JvLuzipGSeQ9BlW/mMi7Hy+t0o3Le/6pVfkT6bc7el1Ap5Z36aN9eNNFU4fNHTOOdOl2UJmbnv59c8+PvtGtm2K1HXkPGrYolZD3KFi9giLvXrjUCyPv6ki9AiLvXriiFEbeo2CtDNq4vFf2qIUF+kHeJX1Gto+UPPibph9urhg6TJVk/qbUa09ZYhZ3/qQ8yHlPSZ8bVtPS39c68p52FJD3tPyR97T87XtQ+l4MXg8al/ey3Wbk8bMPPf4989Sq5a0bhX6Qd4H20LubzTUb3zRD48ebJz5wnFr6zOqXHjey4i4CL2kysfZt9x145N2XmG55Vt51eYZEQ95DqOnVQd71WIZEQt5DqOnWYeVdl6drtL6R9358gpUrxH6Rd+lvdvtIEXgR+TpHdjeZBSecZ26a/ydRHrgU0kfkPYSaXh3kXY9laCTkPZScTj3kXYdjaBTkPZScXj3kXY+lT6S+kfcbbrnbPP38GlbefUavoGx2+0hJnZEUmpBDHrS07K87D13an9/eD2ky+fNA3kNGVq8O8q7HMjQS8h5KTqce8q7DMTQK8h5KTq8e8q7H0idSI/JuV9WrOrb0+svNwgXzq4r13e/7aeVd4GTz368dnmkWD8/wYpbNbx+a0rkB9pN3G1l177cDeU87Ish7Wv7SOvKedgyQ97T8kfe0/O17UPpeDF4PGpH3LFaesNrMJLP579LaE0cfZ+ZNmuLUcFbc+ym/vajzyLvTkEYrhLxHQ+scGHl3RhWlIPIeBatzUOTdGVW0gqy8R0PbM3Dj8p7mNOO22m8r7/Zsffd/X/vWC+bCb59rJGVm3uEnmxWffrBv8tuR97hzOCQ68h5CTbcO8q7L0zca8u5LTLc88q7LMyQa8h5CrX4d5L0+Q9Ov8i6nduEbG8yz27d1V95lBb7skAcvyRNT5ZD92+/45P+lQCZuCFbe4/Ktio68VxGK/3vkPT7jXi0g72n5I+9p+UvryHuaMUgi72cuvNps3LS18Ix5SJPuRJD898+9uaGbB192A2t2R5l+vDG1jAjyrjtXfKMh777E9Msj7/pMfSIi7z609Msi7/pMfSMi777EdMo3Lu8XLFpiZs4YMvctu07nDPogSj+vvAueXjewtlXc5byQ97STH3lPy9+uevX7+096SvF6gLzHY+sSGXl3oRS3DPIel29Z9MblnRtW0wz06m3vmsve+nm38TtmHGkumjps2izuyHuaeZRtFXlPPwasvKcdA+Q9LX/kPS1/u4CQvheD1wPkXWHM27LyZW9glVO+4ucPmHv++tZ9Mt/Jb5c897YdrLynHTHkPS1/+8HZlvef9LT0e4C86zP1iYi8+9CKU5aV9zhcq6I2Lu+SNvOpMz9urr7ss1V9a83v2/Th2RX45//YmL+/q9Xizsp7+pcH8p5+DFh5TzsGyHta/sh7Wv6svKfj37i8ywObbr/rwVY+SbVsmFol788v7cj70u6pzP7dW83DH/93ZvbESelmYI2WWXmvAU+hKvKuALFmCOS9JsCa1ZH3mgBrVkfeawJUqM7KuwLEgBCNy7vkvPc62G0mYBQdq2Rz3EXcXz36011xf+IDx5mh8eMdo/RPMeQ97Vgg72n521WvNi0epCem2wPkXZenbzTk3ZeYfnnkXZ+pS8TG5d2lU20r04YPz6y4S4776b91ycgWkrIH/CNHHds6gUfe075SkPe0/JH39PyR97RjgLyn5W/fg9L3YvB6gLwrjHm/y3v2AUw3zf8Tc8VHv9w96+we8FUPcVLApB4CeVdH6hUQeffCFaUwaTNRsDoHRd6dUUUpiLxHweoVlJV3L1xqhZPIu9y0+uL6Dd2TWHr95WbhgvlG0mlO/djcVu7/3s/y/uyG75sLv72gy7roAUxZgZftI2UbybYcyHvakULe0/K3q179/P6TnlDcHiDvcflWRUfeqwjF/z3yHp9xUQuNy3v2IU3ypNVrr7y4K+/LVzxqHnr8e628kbVfPzzXvvVCR9zPNVu2by4Udzsh2irwyHuaNw3bKvKelj/ynp4/8p52DJD3tPzte1D6XgxeDxqXd1lhf+DOG81Jc08wWXmXXWiWfO1eww2rOpPw1S0vm8+tWmDkb0mTkXSZXkcbBR5515kroVGQ91ByevVIm9FjGRIJeQ+hplcHeddjGRqJlfdQcvXqNS7vIuz/5dZrDpD3VCvvX1x8m/nB36wbRTF/AZFN8zlxzizznZX7tlq0R7+tvMtK+zkPntYV99NnnWUe+cxqp1mSF/g/mn54X9/Eirw7DWu0Qsh7NLTOgZF3Z1RRCiLvUbA6B0XenVFFK4i8R0PbM3Dj8n7DLXebp59f002PsSvvv3n8MeaSq242559zhrn1q19qlIT0Qfpij2z/5Gci929v3DIi7Nm0n36Vd8lxl1z32UPHmycufs4MTRl2Zpq/ibWfd6FB3p2HNUpB5D0KVq+gyLsXLvXCyLs6Uq+AyLsXriiFkfcoWCuDNi7v0iObIpPt3ZWXnt8XT11ds+6l7oVEUWqP7Xv+IVP9tPJ+09O/b+750Te64v7wwtXdv32Ptgg88u47srrlkXddniHRkPcQanp1kHc9liGRkPcQarp1kHddnq7Rksi7a+dSlMum7+RFXvpT9LN+kXeRdpF3WWl/5DPfNfMOPzkYYVbg5UFODx85q++exIq8Bw+vSkXkXQVjrSDIey18tSsj77UR1gqAvNfCp1IZeVfB6B2kcXm3Oeb5vPJ+2CrSirndvtJV3t/est0bvHaFv3rt++aCh8/phl1+zj3mX867tHYTr+zaaRZueNXI38d1BH7VrNndv/vlmDxxgpkyebzZum1nv3RpoPoxftw4Mzx1knln646BOu9+OtmZQ1NMP7z/9BOTJvsyfOhk8+77u8zu3XuabJa29hOYevAks3PXHrN9526YJCIg70EczRNoXN4lx/yi884+IEUm1Q2rFrkV9Wz6jqu8b9+Z9o375c3rzTl/9kkjf199yn8w/+lTy9Rm0qbdu825/7Te/Pj998z0CRPMQ7OPM2dNPVQtfp1A48cbM6EjkDt3760ThrqBBDrozaQJ482OzocnRxoCUyaN74gL/NPQN2byxHFmV+f9Zw9vQUmGYOKEcWbv3r2Ga6ck+LuNynsQR/MEGpd3WWG3K9vZ0025VaRt2+a5Z/uV3c5Sfl7Uz9RpM+c8cJqRPd19dpbxmWpb9uwxf7jpLfPQu5u71W7q7EJzxdBhPiGilCVtJgpW56CkzTijilaQtJloaJ0CkzbjhClaIdJmoqF1DkzajDMq1YKNy3u/rbyLjOdvQM0S7vfdZrI3qPruLOM7k5Zt3mhu3/x2t9q1wzPN4uEZviFUyyPvqji9gyHv3sjUKyDv6ki9AiLvXrjUCyPv6ki9AyLv3shUKjQu75Iec9f9j43s5iJnUZSyonJ2FUFsu0XFst8O9Os+76tfetxc9j8u7nb/iUueq3WDqivve7a8Y27qrMLLcfqUQ8x9RxydbC945N111OKUQ97jcPWJirz70NIvi7zrM/WJiLz70IpTFnmPw7UqauPyLh0q2iqyKJWmqvP98vsUaTPyACZ5EJM8kEmenipPUW3qePb9beaajW8a2ZEm5U40yHtTI17cDvKelr+0jrynHQPkPS1/5D0tf/selL4Xg9eDJPI+1jCnkHeb577ghPPMik8/2DhSEffLfvlzs3bn9u7KuzyN9aKp7g+D0ugw8q5BMTwG8h7OTqsm8q5FMiwO8h7GTasW8q5FMjwOK+/h7OrURN7r0Ntft2l5bzLPvRceuZF12ZaNRlJp5JCbWBcPzWgsjQZ5V5i8NUIg7zXgKVVF3pVABoZB3gPBKVVD3pVA1giDvNeAV6NqEnmXm1Y3btpa2O38/u81zq2xqk3K+7Mbvm8u/PaC7rk98pnV3R1mUh8i7yLxIvPzJk0xKzp58JJOE/tA3mMT7h0feU/LX1pH3tOOAfKelj/ynpa/fQ9K34vB60Hj8i43f86cMWTuW3bdmKHdlLxLfrvkuUu+e9N57lWDJekzkkYj6TSSRiMr8LG3k0Teq0Yl7u+R97h8XaIj7y6U4pVB3uOxdYmMvLtQiluGlfe4fMuiNy7vZfu8pzl9nVabknfZWUZ2mIm1n3tdGvk0mgWHTDV3zDgqWhoN8l53xOrVR97r8dOojbxrUAyPgbyHs9OoibxrUKwXA3mvxy+0NvIeSi5Trwl5t9tCDk0ZNrKf++yh4xV6HieEPMxJHuokMi+r8CLwIvLaB/KuTdQvHvLuxytGaeQ9BlX3mMi7O6sYJZH3GFT9YiLvfry0Sjcu75I286kzP26uvuyzWueQPE5seZd0mVO/9dtJtoUMhSvpM9e8/aZ5dvu2bogYq/DIe+jo6NRD3nU41omCvNehV78u8l6fYZ0IyHsdejp1kXcdjr5RGpf3qiea+p5AP5SPLe/9ni7TawyyN7Nq58Ij72lnP/Kelr+0jrynHQPkPS1/5D0tf/selL4Xg9eDxuVdct57Hew2M5pOm9JlysY1vwovT2a9Y+aRtXekQd7TvmEh72n5I+/p+SPvaccAeU/LH3lPx79xeU93qvFajrXy3sZ0mV6Us7nwUu7a4Znm8mnTg29oRd7jzWmXyMi7C6W4ZVh5j8u3KjryXkUo7u+R97h8XaKTNuNCSb8M8q7ANJa8X/OX/9Y8tO7+vt1dJgRdfkeaOk9nRd5DRkCvDvKuxzI0EvIeSk6nHvKuwzE0CvIeSk6vHvKux9InUhJ5l7z3JV+7d1Q/l15/uVm4YL5P3/umbAx5tw9jasPuMiED8ez728ztm98ZuaFVUmmuHT7MnH7QIc7hkHdnVFEKIu9RsHoFRd69cKkXRt7VkXoFRN69cEUpjLxHwVoZtHF5X77iUXPX/Y+ZB+680Zw094RuB9ese8lcctXN5spLz2/lLjTa8t7PD2OqnFGeBSSVZlnnCa2SFy+Hj8Qj756wlYsj78pAA8Ih7wHQFKsg74owA0Ih7wHQlKsg78pAHcM1Lu9nLrzaXHTe2QdIukj9Q49/zzy1arlj1/unmLa8L3t+qbm982fe4SebJy55rn9ONFJPJJXm3q2bzD1b3+nuDS/HRVOHO09pPaznTa3Ie6QBcQyLvDuCilgMeY8I1yE08u4AKWIR5D0iXMfQyLsjKOVijct72RNWbSrNoO828+qWl81p35rbHeZHPrO6m+8+KIevxCPvaWcG8p6Wv7SOvKcdA+Q9LX/kPS1/+x6UvheD14PG5Z2V996T7MJvLzCS737FR79sbpr/J4M3IztnnL+pVSAUpdMg72mnB/Kelj/ynp4/8p52DJD3tPyR93T8G5d3ct7LBzu7p/sPPv9TIzerDvIhefAr3t1kHnx3y0g6TVbikfe0swN5T8sfeU/PH3lPOwbIe1r+yHs6/o3Lu5wqu80cOOCDdJOq73QvSqcRif/n06aZyw4/zOzetts3JOUVCCDvChBrhiBtpibAmtWR95oAa1ZH3msCVKhOzrsCxIAQSeQ9oJ99XUXjhlV7k6rkuEuuO0fBBU7Bja3DEyaYy6dOr/WwJ1iHEUDew7hp1kLeNWn6x0Le/Zlp1kDeNWmGxULew7jVrYW81yXYqV9X3uUm1XMePM3I6vug3aQagl9W4ldv22oeee9d81fvbRsJ4bPNZEi71BlNAHlPPyOQ97RjgLyn5Y+8p+UvrSPvacagMXm3ue5Fe7n3+l0aLH6t1pV3+yTVBSecZ1Z8+kG/xge4tOS8/3+73jcr3nqnK/N2m0l5ausV0zoPfJpykNdDnwYYZdCpI+9B2FQrIe+qOL2DIe/eyFQrIO+qOIOCIe9B2GpXakzeL1i0xMycMWTuW3ZdYae/uPg28/bGLeY7K5fWPqmmA9SR97H+JNWYY5G9YdXmxT/z/nsjT22VtmU1/tyDDzUXTx0yIvUcegSQdz2WoZGQ91ByOvWQdx2OoVGQ91ByevWQdz2WPpEak/ey/d1tZwd1n3e7NeS1pywxizt/ONwJlO028+z728x33//VqF1qEHl3rq4lkXdXUvHKIe/x2LpERt5dKMUrg7zHY+saGXl3JaVbDnlX4Bm68p5ddWdrSP+BqNoqUlbj1+5439y++R2zduf7I2k1iLw/66IayLsOxzpRkPc69OrXRd7rM6wTAXmvQ0+nLvKuw9E3SmPyLg9nuvbKi83CBfML+ygr77ff9aB5atVy33NIXj5U3uVJqnKzqjyMSR7KxOFHoEres9H23eT6rnnoV1sLRf6Mgw4mR94Pv0HePYFFKI68R4DqERJ594AVoSjyHgGqZ0jk3ROYUvHG5P2GW+42f/cPL5fmtFflxCudb5QwIfJ+z4++YW56+vfN7KHjzXOfXxelX2M9qI+8F4m8pNY800mxsTe6ShnJi19w8NSOyB9sFhwylTz5HpMIeU//CkPe044B8p6WP/Kelr+0jrynGYPG5F1OT1bf5civrsvPN27qrIg+uTINhZqt+sq7bAl56rd+m60ha3IPlfe8yEtqjYj86m2/MvJU1+wxb/IUI6vy5x50qJk3+SBkPgMHea85gRWqI+8KEGuEQN5rwFOoirwrQKwZAnmvCTCweqPyLn2UFfjHnnhmVHdP/djc0l1oAs+r0Wq+8s4DmXSGR0Pe8z15dedOs/q9X5nvdv7k8+SlrOxe8+Epk81pnb/P6KzOD/IONsi7zjyuEwV5r0Ovfl3kvT7DOhGQ9zr0dOoi7zocfaM0Lu++HWxDeR95Z9Vdb0RjyHu2d/aGV1mV/8n2HaO2oLTlZk+c1E2xsX9mT5qkd4J9Hgl5Tz9AyHvaMUDe0/JH3tPyl9aR9zRjgLwrcPeRd1bdFYDvDxFb3vM9zct80cq8rMTPm3RQd3X+QxOndKV+rAo98q43l0MjIe+h5HTqIe86HEOjIO+h5PTqIe96LH0iIe8+tErKuso7q+4KsDMhmpb3Mpn/yY4d5u92bu+szL93Xgvx8AAAGYpJREFUQM681MkL/bwpU8zsCZNan3KDvOvO55BoyHsINb06yLsey5BIyHsINd06yLsuT9doyLsrqR7lXOWdVXcF2H0k70VnI6vzz3Qk/rnt2zoiv6uzz/z2QqGXupJyIzfEzp44sbtK3zapR95153NINOQ9hJpeHeRdj2VIJOR9NLXxmzebcZs3jfxw/JbNZnzm//KLcZ0y+Z9NePWVA/DnfyZ1pG72kJ9NWvPjkKGjTk0CyHtNgFLdRd5ZdVcAnQuReuXd9Yys0MvNsLJCv1b+dKS+7BCpP7azMt+V+kkdue/k0c+bONkMT5jQV6v1yLvrDIhXDnmPx9YlMvLuQilemZTyPuGVl0eLbKAoi2DnpTgfWxqa+Oro9kTSRdaTH3v3Ju/CIHYAeVcYdRd5Z9VdAXRL5b3szEXmJdXmtd07963Sd6RetqrM7jufrytivy8NZ9+KvUj+UEfqU8g98q4/p30jIu++xHTLI++6PMuilYny1IMnmZ279pjtO3c7rSiHinKRTDdz5n6t7BkeNnuHp49U2jM0bPZk/m9/sfu440cF3j37uAMayv9M4uztxM8e8rMjfu9Uv05SWoUA8q6AsUreWXVXgFwQoi0r775nL1K/dtcOI39v2LNf7Dsr9Zv37O4p9tKOiL0Iflfyx8m/4wk+8u47svrlkXd9pj4Rx6K851MvhEd+1Vd+lhdal9SLVq0oV0yEvAAXibLIbl6eD5DijmDnpTgfW7qya/Zo4RZJF1lPfZDznmYEkHcF7lXyzqq7AuQBkvdetEToXxOh37lrZMX+1d2df3dW7PMPmOoVx67gZyV/2rgJZljkf9LEzkVAJ0XHyL/Lt75E3uPMa5+oyLsPLf2yGvKulaecX1Uuy1GWctmjDavKZaI8ccI4s7eTtrF7j+kK8FgXZf0ZXD8i8l6fYUgE5D2EWq5OL3ln1V0BcEmIsbryXoeYyP0Ws6ebhmNX7jd3Ptms4Lus3ufbl9X8YZH5zt8i9bMnTOwWOa4j9kcfMtmMf39PN3VnePw4c+z4fbLfb/n5dZj2c13k3X90XFaWXW7qEwk+eNtWs6OTtrGnk/Y7VmQ5n3ohhPOrvvKzkNQL7RXllDnv/jNvbNZA3tOMK/KuwL2XvLPqrgAYeVeHmJX8Lbt3d1fxt+zdY7KiL436rOYXddKKv/zu2E4qjxxW/ocndC4GOqk93d91cvftISv/3Z/tvxDgYqB8+Nsi7y6ry/kV4K5A51eJc7tiuKRh9PvKskuecsiqclmOsqR3ZI8ioVZ/w4kUEHmPBNYjLPLuAUuxKPKuALNM3ll1V4DbIwQr73H52uhyA+3mjuDbFX2R/c2dn73bWeHfOWmc+fm2HfvEv/MzSd+RI2SF3/VsshcE2YuC7IXByIVAJ+ffHjYtKNuOvVCwP8teMNif9fO3CL3k3UWY8yvMIcIsK85tSsVwWVl2leWpRx9utm3fbXZ3vt0aBFl2fY02VQ55b4p07wWE9L0YvB4g7wpjXibv9/zoG+amp3/fnD7rLPPIZ1YrtESILAHkPe18cMl5t+IvPZVcfTkkX3+f4O8xW/fu3vezTpqPPSTFp1t+/4XAvrLVN+s2SSN/AZFt+8PvvW+mb9068iP59/StW7o3Edtjzuuvj+ru8Z3/y83F9nDJX5abCGUMdkvORufo5xVml9Xl/ApwV6Dzq8S5XTFc0jBirixr5Lw3OW/HWlvIe/oRZeU9zRgg7wrcy+T9tG/NNa9uebkr7iLwHLoEkHddnr7RXOTdN6ZL+ewFQVfy918UyL/lwmBiJrXi1U46kJXaLfsvAPKSe/zPR4t0Xqzz/98n4xk5f3f0/13OoYkym6ZNM5umThtpqvv/zp/skf9ZUZmXjz5mVJ31x4z+/6ZpQwfEzZfJBuh14RPCxaZjhdTtVUdu3rZpXWXlDpo0fiTnvU772Qu3OnF86mZT1XzqaZW198nUiZfdKjI0TtG3baGx6tbrtUFA3dix6iPvscj2jou8K3AvkvdnN3zfXPjtBWb20PHmuc+vU2iFEHkCyHvaOdFL3vMpG/mt5rICnU/VyK86H7DHcy5No99WnENWmfPbx20uEO9Xph7a/bbCHus7Un3IlAndtA05rDDbby5cZkf22w2X8rZM3XshfNqiLAQg0DyB7DeFvVp/5aS5zXeOFg3yrjAJiuRdxF0E/qb5f2Ku+OiXFVohBPIePgeygttLpPN7NWf/n5fkbNpG3zztr4MonyZx4P7Io7eUy4vzAf/PPdBEcpuz6Rwp91tuyw2r+ZkrN0xrHdlvXrRiSpzNnXQkub+j1zHtkEkjOe912pYbxps+sqlqTbct7dn7ZOq0nd0qMjRO6AVsaHu96rXxonjv75wcAwUxKwgg7wpTJC/vdtV9aMqw+cHnf2rkbw59AmNl5d1Ksaw4y81/9rA/z69Mlwl19sbB1DKdX33OC3RWsPO5zSLG2YeWHLDHc06eY+Y0689a3YhtlXddCumikfOejr20TM57PP6uF9infmAoXieIXEoAeVeYHHl5t6vu156yxCzu/OGIQ6BpeS+S7CLBzqZ9lK14N53qkRXcXiLdaxX6gBsKj59jDps22by1eXv3kdz98LS/ODOtf6Mi72nHBnlPyx95T8tfWifnPc0YIO8K3LPyLjeoyo2qrLorgK0I4SLvNvc6u6ptxdmuYFvZzj5kxaaWNLGCbaU4/3jtrCxnpXrUvzMpHdl0jiZkOtUNq/FnVntaQN7TjhXynpY/8p6WP/Kejj/yrsA+K+/2oUyS5y757hz1CFj5/rVM70stEfGe2HnIz+QNr5jtO/eM7ChiU0dirGwXSbbdDzqb+pFN+yhb8R4LqR7Ie725rVEbedegGB4DeQ9np1ETedegWC8GK+/1+IXWRt5DyWXqWXnnoUy9YYqIT+jsTW1XuK2E25VvK9wi6lor3jb3OruqbcXZrmBb2c4+ZMWmljSxgq0wBZOEQN6TYB/VKPKedgyQ97T8kfe0/Fl5T8cfeVdgb+V9EB/KJMK9T7Z/vSIuMi4///XPRNg3B5HOy7dd6RbxlpX3STOnm3enTB3ZYcSmjoyFle0gYA1WQt4bhF3SFPKedgyQ97T8kfe0/JH3dPyRdwX2Vt7PeeA0s/atF8bEQ5myUt7992uv1BZyEfHds48feYy4iLZd/Za/R9JS9u8mUiXgLjnvCsNLiBICyHv6qYG8px0D5D0tf+Q9LX/kPR1/5F2Bvch7dnvIdVf8XCFqvBBZMZ/0kxeMrJRPXNP5u5tL7rdKLoItKSbZFXFJQ5Gf//pnIuz622Ui7/HmiEtk5N2FUtwyyHtcvlXRkfcqQnF/j7zH5esSnZx3F0r6ZZB3BaYi7/20PaSkqEz6yY+7aSsi53JzpxV21xs5s1Iu/+4K+f7V8X0r5nGE3Gc4kHcfWvplkXd9pr4RkXdfYrrlkXddnr7RkHdfYvrlkXd9pi4RkXcXShVlfvr6L8zce47ulpJV9yYeymQFXWRc5Lwr6d1/V6+cZ8V854dP3ifm+1fKd374I1FWyRUwHxACeY9B1T0m8u7OKlZJ5D0WWbe4yLsbp1ilkPdYZN3jIu/urDRLIu8KNP/jn/+Buf35peb0WWd18901j25aSyedRdJaJO9c/pZV9V43gEqKiki4pK1YOd91UkfSOyvm8v+xciDvaUcSeU/LX1pH3tOOAfKelj/ynpa/fQ9K34vB6wHyrjDmw1+bbmSbSBF3EfiQw66ku0p6XtBFyvfJentWzkM4Zesg73UJ1quPvNfjp1EbedegGB4DeQ9np1ETedegWC8GK+/1+IXWRt4dyF2waIl5cf2GbskT58wy31m5dKTWk+ufNGd/82wz7/CTzROXPOcQzXRXzWX1fPJfPTWS7iIr7EWHlXRZOd917HFG/h4kQe8FFHl3mm7RCiHv0dA6B0benVFFKYi8R8HqHBR5d0YVrSDyHg1tz8DIewX3Ly6+zby9ccuIsIvIz5wxZO5bdl23poi7CPy1pywxizt/8kde1G1uelGz2//Xs7pyjqS7vRiQdzdOsUoh77HIusdF3t1ZxSiJvMeg6h4TeXdnFask8h6LbO+4yHsF9zMXXm2uvfJis3DB/G7JVaufNrff9aB5atXy7v/H/dG47t/2RlWR80lrOqvqzzzd/btoRT27mr79jDO7N4uOpVz0pqYy8t4U6eJ2kPe0/KV15D3tGCDvafkj72n52/eg9L0YvB4g7z3GfM26l8wlV91sHrjzRnPS3BO6JfM/+52rxpmz3hk2t209ayQFJh/Srqgj6rovMORdl6dvNOTdl5h+eeRdn6lPROTdh5Z+WeRdn6lvRFbefYnplEfea8q7Gbdv5X3kmDPHmI9+1Jjf+719f3/iEzojRRQIQAACEIAABCAAgYEngLzXlPedxx1rJn38d4254IJ9si5/OCAAAQhAAAIQgAAEIBCBAPJeAbUo533J1+41a59cOVJTnrDK0TwB0maaZ55tkbSZtPylddJm0o4BaTNp+ZM2k5a/fQ9K34vB6wHyXjHmVbvNSHXkPc0LB3lPw922iryn5Y+8p+ePvKcdA+Q9LX/kPR1/5N2Bfa993pF3B4CRiiDvkcA6hkXeHUFFLMbKe0S4DqGRdwdIEYsg7xHhOobmhlVHUMrFkHcFoKy8K0AMCIG8B0BTrIK8K8IMDIW8B4JTqoa8K4EMDIO8B4JTrIa8K8L0CIW8e8AqK4q8K0AMCIG8B0BTrIK8K8IMDIW8B4JTqoa8K4EMDIO8B4JTrIa8K8L0CIW8e8BC3hVgKYZA3hVhBoRC3gOgKVdB3pWBeoZD3j2BKRdH3pWBBoRD3gOgKVRB3hUgsvKuADEgBPIeAE2xCvKuCDMwFPIeCE6pGvKuBDIwDPIeCE6xGvKuCNMjFPLuAYuVdwVYiiGQd0WYAaGQ9wBoylWQd2WgnuGQd09gysWRd2WgAeGQ9wBoClWQdwWIhIAABCAAAQhAAAIQgEATBJD3JijTBgQgAAEIQAACEIAABBQIIO8KEAkBAQhAAAIQgAAEIACBJggg701Qpg0IQAACEIAABCAAAQgoEEDeAyFWPXU1MOzAV/Pl2qv8qtVPmyVfu/cApmufXDnwnF0B+I6HxF2z7iVzyVU3mwfuvNGcNPcE16Yot5+AJnNeA/Wnlc94fHHxbeYHf7NuVKO83/iNgSZv5r8f+6LSPuNxwy13m8eeeIb5Xx97ZQTkvRLRgQXkDfrtjVvMd1Yu7f5SJvfMGUPmvmXXBUSjiiXgy7WqvLxx337Xg+apVcuBHECgim9RyDMXXm02btra/RXy7g9dmzmvAf8xyNbwHQ+Z/9n3G5GZp59fw3uQ4zBo82b+O4IvKeY7HuJCf3zdZSOLNstXPGoeevx7zP96w1BYG3kPgCpv0NdeebFZuGB+tzZvEAEQC6r4cq0qz7jUG5cqvmXRWXkP567NnNdA+FhIzdDxsK3yWvDjr82b+e/HP19aezzq9YbaWQLIu+d8KHoz5g3aE2JBcV+uLuWLvjLlK2y3sXLhi7y7sXQtFYM5rwFX+geWqzMeNhorj+78Y/Bm/rvzz5fUGA9Zuf/ZS6+x8h4+DKU1kXdPqBoT2rPJgSjuy9W3vEDMfwU4EGADTzKEL6uNgbD3V2uCOa8B9zGqMx7Siq2/9PrLR76ldW998Eo2wZv57z6v6oxHNn2SBTN35j4lkXcfWpk35Gw+LyvvnhALivu+UfiWz36Y8mZSPV4hfJH3aq69SjTB3LbBa6B6rDTG48pLzzdXX/bZ6sYoUXiju+tnqy1XxZv57z7R6sx/24p883TX/Y8Z3m/cubuWRN5dSWXKFeWBya4mTNAAmDW4+o6D/QqVcXIbJ1++yLsb116lYjPnNeA3RiHjYRlzw7Yfaykdmzfz329MQsYj38K8Tyxi8wI/7E6lkXcnTKML+d6BHdDEQFap4ip3ssthd/mpKp/f+YFdgfymVRXf/Hgg7358i0prM+c1UG9MfMeDGyT7izfzv9nxYLelerx9aiPvPrQyZX32Pg1sYiCr9eJaJItV5V9cv2GE46kfm8t2np6zqopv9mLKrpzZrSLl/zOmT+NmpYTMs+Mn3eA14DkYneKurwGbZlDUAnnv7tw1eTP/3bmXlXQdD6mf5y0/45vu+mNQFAF5j8OVqBCAAAQgAAEIQAACEFAngLyrIyUgBCAAAQhAAAIQgAAE4hBA3uNwJSoEIAABCEAAAhCAAATUCSDv6kgJCAEIQAACEIAABCAAgTgEkPc4XIkKAQhAAAIQgAAEIAABdQLIuzpSAkIAAhCAAAQgAAEIQCAOAeQ9DleiQgACEIAABCAAAQhAQJ0A8q6OlIAQgAAEIAABCEAAAhCIQwB5j8OVqBCAAAQgAAEIQAACEFAngLyrIyUgBCAAAQhAAAIQgAAE4hBA3uNwJSoEIAABCEAAAhCAAATUCSDv6kgJCAEIQAACEIAABCAAgTgEkPc4XIkKAQhAAAIQgAAEIAABdQLIuzpSAkIAAhCAAAQgAAEIQCAOAeQ9DleiQgACEIAABCAAAQhAQJ0A8q6OlIAQgAAEIAABCEAAAhCIQwB5j8OVqBCAAAQgAAEIQAACEFAngLyrIyUgBCAAAQhAAAIQgAAE4hBA3uNwJSoEIAABCEAAAhCAAATUCSDv6kgJCAEItI3A8hWPmrvuf+yAbl956fnm6ss+a85ceHX3d0+tWn5AGfndjOlD5jsrl3Z/VxVr3icW9cQzY/q0bjtfXHyb+cHfrCssu/T6y83CBfPNBYuWmBfXbzD2/7bwqtVPmyVfu9ecOGfWSL/ygVz6Mf+Uk8xjTzwzUvX8c84wt371S17tupxH2+YL/YUABCCQkgDynpI+bUMAAskJWLl84M4bzUlzTxjpj0j4Xzz1wxH5Fdk99WNzzX3Lrhspc8Mtd5unn18zIvWusfKSnZdv+b3EenvjllL5ljJW3vP9sj/vJe9Z8Fb2i/pR9Dufdl3OI/kkoAMQgAAEWkQAeW/RYNFVCEBAn4BIuV1R7hU9L7Fr1r1kLrnq5lGr3q6xNOV95oyh7gq9vfiw/RKhr5J/l36Uybtru8i7/pwlIgQgMNgEkPfBHn/OHgIDT0DSXn7rhGNHraiXQRER/dlLr3VX2mX1WQQ2uxLvE0va6LXi7SK90ocPffB488Zb75ijDj+sm9Ii3wbIIT+LKe+u7bqcx8BPQgBAAAIQ8CCAvHvAoigEIDD2CFiBtmdmc87LzjSbK772yZWjivnGqpJ3l5x3kehTP/ahbo679Ef6J6vwd9zzcHR5d2mXnPex95rhjCAAgbQEkPe0/GkdAhDoIwI25cR2qSidxgq3vZm1rPs+serkvIu825tIpS/22wCfFe+QnHfXdn360UdTga5AAAIQ6FsCyHvfDg0dgwAEUhKQ9BPZaSW/ul6U617Vz7JYVSvvVWkvNm1G5N3ucmMvBHykuY68V7Xr048qjvweAhCAAASMQd6ZBRCAwMASEBH/v7/9F92V6/xhpTS/C02ZvIfE0pR36b/k3NvtLH2kuY68V7Xr04+BnYicOAQgAAEPAsi7ByyKQgACY4tANrUlu8Ke3bEle0OqnH0veZfdZ+RwjaUt79nR8ZHmuvLeq12ffoyt2cXZQAACEIhDAHmPw5WoEIBAiwgUPbCoLKe9Km3GJ1aVvLvesFr0zYGPNJf1w6b72KHMPqTJ5rznhznfLjestuiFQFchAIFWEEDeWzFMdBICEIAABCAAAQhAAALkvDMHIAABCEAAAhCAAAQg0BoCrLy3ZqjoKAQgAAEIQAACEIDAoBNA3gd9BnD+EIAABCAAAQhAAAKtIYC8t2ao6CgEIAABCEAAAhCAwKATQN4HfQZw/hCAAAQgAAEIQAACrSGAvLdmqOgoBCAAAQhAAAIQgMCgE0DeB30GcP4QgAAEIAABCEAAAq0hgLy3ZqjoKAQgAAEIQAACEIDAoBNA3gd9BnD+EIAABCAAAQhAAAKtIYC8t2ao6CgEIAABCEAAAhCAwKATQN4HfQZw/hCAAAQgAAEIQAACrSGAvLdmqOgoBCAAAQhAAAIQgMCgE0DeB30GcP4QgAAEIAABCEAAAq0hgLy3ZqjoKAQgAAEIQAACEIDAoBNA3gd9BnD+EIAABCAAAQhAAAKtIYC8t2ao6CgEIAABCEAAAhCAwKATQN4HfQZw/hCAAAQgAAEIQAACrSGAvLdmqOgoBCAAAQhAAAIQgMCgE0DeB30GcP4QgAAEIAABCEAAAq0hgLy3ZqjoKAQgAAEIQAACEIDAoBNA3gd9BnD+EIAABCAAAQhAAAKtIYC8t2ao6CgEIAABCEAAAhCAwKATQN4HfQZw/hCAAAQgAAEIQAACrSGAvLdmqOgoBCAAAQhAAAIQgMCgE0DeB30GcP4QgAAEIAABCEAAAq0hgLy3ZqjoKAQgAAEIQAACEIDAoBNA3gd9BnD+EIAABCAAAQhAAAKtIYC8t2ao6CgEIAABCEAAAhCAwKATQN4HfQZw/hCAAAQgAAEIQAACrSGAvLdmqOgoBCAAAQhAAAIQgMCgE0DeB30GcP4QgAAEIAABCEAAAq0hgLy3ZqjoKAQgAAEIQAACEIDAoBNA3gd9BnD+EIAABCAAAQhAAAKtIYC8t2ao6CgEIAABCEAAAhCAwKATQN4HfQZw/hCAAAQgAAEIQAACrSGAvLdmqOgoBCAAAQhAAAIQgMCgE0DeB30GcP4QgAAEIAABCEAAAq0h8P8DbYAcepdSKbEAAAAASUVORK5CYII=",
"text/html": [
""
]
},
"metadata": {},
"output_type": "display_data"
}
],
"source": [
"dynamics.plot_history(colors=[\"darkturquoise\", \"green\", \"red\"],\n",
" title=\"Same plot as above, both only showing initial detail\", xrange=[0, 0.3])"
]
},
{
"cell_type": "code",
"execution_count": 25,
"id": "5c44014b-9dee-4d8c-a2de-13478ee7bb63",
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(0.02405919499545674, 23.73396682504195)"
]
},
"execution_count": 25,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"# Look at where the curves intersect\n",
"dynamics.curve_intersect(\"A\", \"B\", t_start=0, t_end=0.1)"
]
},
{
"cell_type": "code",
"execution_count": 26,
"id": "ac0382c7-5940-4db7-b1df-80edd29c3336",
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(0.14412951669101942, 6.026379520544665)"
]
},
"execution_count": 26,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"dynamics.curve_intersect(\"A\", \"S\", t_start=0.1, t_end=0.2)"
]
},
{
"cell_type": "code",
"execution_count": null,
"id": "d70f2d6b",
"metadata": {},
"outputs": [],
"source": []
}
],
"metadata": {
"kernelspec": {
"display_name": "Python 3 (ipykernel)",
"language": "python",
"name": "python3"
},
"language_info": {
"codemirror_mode": {
"name": "ipython",
"version": 3
},
"file_extension": ".py",
"mimetype": "text/x-python",
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
"version": "3.8.10"
}
},
"nbformat": 4,
"nbformat_minor": 5
}