"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"# PyRosettaCluster Tutorial 1A. Simple protocol\n",
"\n",
"PyRosettaCluster Tutorial 1A is a Jupyter Lab that generates a decoy using `PyRosettaCluster`. It is the simplest use case, where one protocol takes one input `.pdb` file and returns one output `.pdb` file. \n",
"\n",
"All information needed to reproduce the simulation is included in the output `.pdb` file. After completing PyRosettaCluster Tutorial 1A, see PyRosettaCluster Tutorial 1B to learn how to reproduce simulations from PyRosettaCluster Tutorial 1A."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"*Warning*: This notebook uses `pyrosetta.distributed.viewer` code, which runs in `jupyter notebook` and might not run if you're using `jupyterlab`."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"*Note:* This Jupyter notebook uses parallelization and is **not** meant to be executed within a Google Colab environment."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"*Note:* This Jupyter notebook requires the PyRosetta distributed layer which is obtained by building PyRosetta with the `--serialization` flag or installing PyRosetta from the RosettaCommons conda channel \n",
"\n",
"**Please see Chapter 16.00 for setup instructions**"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"*Note:* This Jupyter notebook is intended to be run within **Jupyter Lab**, but may still be run as a standalone Jupyter notebook."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"### 1. Import packages"
]
},
{
"cell_type": "code",
"execution_count": null,
"metadata": {},
"outputs": [],
"source": [
"import bz2\n",
"import glob\n",
"import logging\n",
"import os\n",
"import pyrosetta\n",
"import pyrosetta.distributed.io as io\n",
"import pyrosetta.distributed.viewer as viewer\n",
"\n",
"from pyrosetta.distributed.cluster import PyRosettaCluster\n",
"\n",
"logging.basicConfig(level=logging.INFO)"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
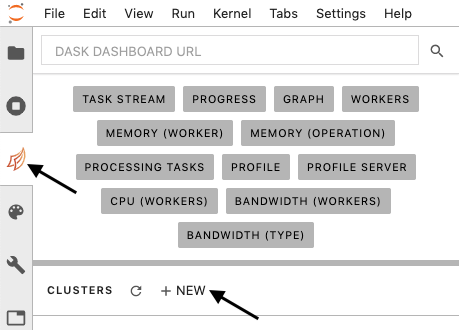
"### 2. Initialize a compute cluster using `dask`\n",
"\n",
"1. Click the \"Dask\" tab in Jupyter Lab (arrow, left)\n",
"2. Click the \"+ NEW\" button to launch a new compute cluster (arrow, lower)\n",
"\n",
"\n",
"\n",
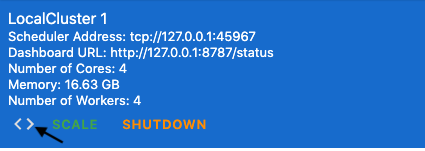
"3. Once the cluster has started, click the brackets to \"inject client code\" for the cluster into your notebook\n",
"\n",
"\n",
"\n",
"Inject client code here, then run the cell:"
]
},
{
"cell_type": "code",
"execution_count": null,
"metadata": {},
"outputs": [],
"source": [
"# This cell is an example of the injected client code. You should delete this cell and instantiate your own client with scheduler IP/port address.\n",
"if not os.getenv(\"DEBUG\"):\n",
" from dask.distributed import Client\n",
"\n",
" client = Client(\"tcp://127.0.0.1:40329\")\n",
"else:\n",
" client = None\n",
"client"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"Providing a `client` allows you to monitor parallelization diagnostics from within this Jupyter Lab Notebook. However, providing a `client` is only optional for the `PyRosettaCluster` instance and `reproduce` function. If you do not provide a `client`, then `PyRosettaCluster` will instantiate a `LocalCluster` object using the `dask` module by default, or an `SGECluster` or `SLURMCluster` object using the `dask-jobqueue` module if you provide the `scheduler` argument parameter, e.g.:\n",
"***\n",
"```\n",
"PyRosettaCluster(\n",
" ...\n",
" client=client, # Monitor diagnostics with existing client (see above)\n",
" scheduler=None, # Bypasses making a LocalCluster because client is provided\n",
" ...\n",
")\n",
"```\n",
"***\n",
"```\n",
"PyRosettaCluster(\n",
" ...\n",
" client=None, # Existing client was not input (default)\n",
" scheduler=None, # Runs the simluations on a LocalCluster (default)\n",
" ...\n",
")\n",
"```\n",
"***\n",
"```\n",
"PyRosettaCluster(\n",
" ...\n",
" client=None, # Existing client was not input (default)\n",
" scheduler=\"sge\", # Runs the simluations on the SGE job scheduler\n",
" ...\n",
")\n",
"```\n",
"***\n",
"```\n",
"PyRosettaCluster(\n",
" ...\n",
" client=None, # Existing client was not input (default)\n",
" scheduler=\"slurm\", # Runs the simluations on the SLURM job scheduler\n",
" ...\n",
")\n",
"```"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"### 3. Define or import the user-provided PyRosetta protocol(s):\n",
"\n",
"Remember, you *must* import `pyrosetta` locally within each user-provided PyRosetta protocol. Other libraries may not need to be locally imported because they are serializable by the `distributed` module. Although, it is a good practice to locally import all of your modules in each user-provided PyRosetta protocol."
]
},
{
"cell_type": "code",
"execution_count": null,
"metadata": {},
"outputs": [],
"source": [
"if not os.getenv(\"DEBUG\"):\n",
" from additional_scripts.my_protocols import my_protocol"
]
},
{
"cell_type": "code",
"execution_count": null,
"metadata": {},
"outputs": [],
"source": [
"if not os.getenv(\"DEBUG\"):\n",
" client.upload_file(\"additional_scripts/my_protocols.py\") # This sends a local file up to all worker nodes."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"#### Let's look at the definition of the user-provided PyRosetta protocol `my_protocol` located in `additional_scripts/my_protocols.py`:"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"```\n",
"def my_protocol(input_packed_pose=None, **kwargs):\n",
" \"\"\"\n",
" Relax the input `PackedPose` object.\n",
" \n",
" Args:\n",
" input_packed_pose: A `PackedPose` object to be repacked. Optional.\n",
" **kwargs: PyRosettaCluster task keyword arguments.\n",
"\n",
" Returns:\n",
" A `PackedPose` object.\n",
" \"\"\"\n",
" import pyrosetta # Local import\n",
" import pyrosetta.distributed.io as io # Local import\n",
" import pyrosetta.distributed.tasks.rosetta_scripts as rosetta_scripts # Local import\n",
" \n",
" packed_pose = io.pose_from_file(kwargs[\"s\"])\n",
" \n",
" xml = \"\"\"\n",
"
"
]
}
],
"metadata": {
"kernelspec": {
"display_name": "Python [conda env:PyRosetta.notebooks]",
"language": "python",
"name": "pyrosetta.notebooks"
},
"language_info": {
"codemirror_mode": {
"name": "ipython",
"version": 3
},
"file_extension": ".py",
"mimetype": "text/x-python",
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
"version": "3.7.6"
}
},
"nbformat": 4,
"nbformat_minor": 4
}