{
"cells": [
{
"cell_type": "markdown",
"metadata": {},
"source": [
"# 5.11 Batch Normalization\n",
"- Let's review some of the practical challenges when training deep networks.\n",
" - Data preprocessing is a key aspect of effective modeling. \n",
" - We standardized input data to zero mean and unit variance. \n",
" - Standardizing input data makes the distribution of features similar, which generally makes it easier to train effective models since parameters are a-priori at a similar scale.\n",
" - The activations in intermediate layers will assume rather different orders of magnitude. \n",
" - This can lead to issues with the convergence of the network due to scale of activations\n",
" - If one layer has activation values that are 100x that of another layer, we need to adjust learning rates adaptively per layer (or even per parameter group per layer).\n",
" - Deeper networks are fairly complex and they are more prone to overfitting. \n",
" - This means that regularization becomes more critical. \n",
" - Dropout is nontrivial to use in convolutional layers and does not perform as well\n",
" - Dence we need a more appropriate type of regularization.\n",
" - When the last layers will converge first, at which point the layers below start converging. \n",
" - Unfortunately, once this happens, the weights for the last layers are no longer optimal and they need to converge again. \n",
" - As training progresses, this gets worse.\n",
"- Batch normalization (BN) can be used to cope with the challenges of deep model training. \n",
" - During training, BN continuously adjusts the intermediate output of the neural network by utilizing the mean and standard deviation of the mini-batch.\n",
" - Ioffe and Szegedy, \"Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift,\" 2015 - https://arxiv.org/abs/1502.03167 \n",
"- In a nutshell, the idea in Batch Normalization is to transform the activation at a given layer from $\\mathbf{x}$ to $$\\mathrm{BN}(\\mathbf{x}) = \\mathbf{\\gamma} \\odot \\frac{\\mathbf{x} - \\hat{\\mathbf{\\mu}}}{\\hat\\sigma} + \\mathbf{\\beta}$$\n",
" - $\\hat{\\mathbf{\\mu}}$ is the estimate of the mean\n",
" - $\\hat{\\mathbf{\\sigma}}$ is the estimate of the variance\n",
" - The activations are approximately rescaled to zero mean and unit variance. \n",
" - Since this may not be quite what we want, we allow for a coordinate-wise scaling coefficient $\\mathbf{\\gamma}$ and an offset $\\mathbf{\\beta}$. \n",
" - To address the fact that in some cases the activations actually need to differ from standardized data, we need to introduce scaling coefficients $\\mathbf{\\gamma}$ and an offset $\\mathbf{\\beta}$.\n",
" - Consequently the activations for intermediate layers cannot diverge any longer\n",
" - we are actively rescaling it back to a given order of magnitude via $\\mathbf{\\mu}$ and $\\sigma$. \n",
" - Consequently we can be more aggressive in picking large learning rates on the data. "
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.11.1 Batch Normalization Layers \n",
"- Batch Normalization for Fully Connected Layers\n",
" - We will put the batch normalization layer between the affine transformation and the activation function in the fully connected layer. \n",
" - We denote by $\\mathbf{u}$ the input and by $\\mathbf{x} = \\mathbf{W}\\mathbf{u} + \\mathbf{b}$ the output of the linear transform.\n",
" - This yields the following variant of the batch norm: $$\\mathbf{y} = \\phi(\\mathrm{BN}(\\mathbf{x})) = \\phi(\\mathrm{BN}(\\mathbf{W}\\mathbf{u} + \\mathbf{b}))$$\n",
" - Recall that mean and variance are computed on the same minibatch $\\mathcal{B}$ on which this transformation is applied to. \n",
" - Also recall that the scaling coefficient $\\mathbf{\\gamma}$ and the offset $\\mathbf{\\beta}$ are parameters that need to be learned. \n",
" - They ensure that the effect of batch normalization can be neutralized as needed.\n",
"
\n",
"- Batch Normalization for Convolutional Layers\n",
" - Batch normalization occurs after the convolution computation and before the application of the activation function. \n",
" - If the convolution computation outputs multiple channels, we need to carry out batch normalization for each of the outputs of these channels, and each channel has an independent scale parameter and shift parameter. \n",
" - Assume that there are $m$ examples in the mini-batch. \n",
" - On a single channel, we assume that the height and width of the convolution computation output are $p$ and $q$, respectively. \n",
" - We need to carry out batch normalization for $m \\times p \\times q$ elements in this channel simultaneously. \n",
" - While carrying out the standardization computation for these elements, we use the same mean and variance. \n",
" - In other words, we use the means and variances of the $m \\times p \\times q$ elements in this channel rather than one per pixel.\n",
"
\n",
"- Batch Normalization During Prediction\n",
" - At prediction time we might be required to make one prediction at a time. \n",
" - $\\mathbf{\\mu}$ and $\\mathbf{\\sigma}$ arising from a minibatch are highly undesirable once we've trained the model. \n",
" - One way to mitigate this is to compute more stable estimates on a larger set for once (e.g. via a moving average) and then fix them at prediction time.\n",
" - Consequently, Batch Normalization behaves differently during training and test time."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.11.2 Implementation Starting from Scratch"
]
},
{
"cell_type": "code",
"execution_count": 1,
"metadata": {},
"outputs": [],
"source": [
"import gluonbook as gb\n",
"from mxnet import autograd, gluon, init, nd\n",
"from mxnet.gluon import nn\n",
"\n",
"def batch_norm(X, gamma, beta, moving_mean, moving_var, eps, momentum):\n",
" # Use autograd to determine whether the current mode is training mode or prediction mode.\n",
" if not autograd.is_training():\n",
" # If it is the prediction mode, directly use the mean and variance obtained\n",
" # from the incoming moving average.\n",
" X_hat = (X - moving_mean) / nd.sqrt(moving_var + eps)\n",
" else:\n",
" assert len(X.shape) in (2, 4)\n",
" if len(X.shape) == 2:\n",
" # When using a fully connected layer, calculate the mean and variance on the feature dimension.\n",
" mean = X.mean(axis=0)\n",
" var = ((X - mean) ** 2).mean(axis=0)\n",
" else:\n",
" # When using a two-dimensional convolutional layer, calculate the mean\n",
" # and variance on the channel dimension (axis=1). Here we need to maintain\n",
" # the shape of X, so that the broadcast operation can be carried out later.\n",
" mean = X.mean(axis=(0, 2, 3), keepdims=True)\n",
" var = ((X - mean) ** 2).mean(axis=(0, 2, 3), keepdims=True)\n",
" \n",
" # In training mode, the current mean and variance are used for the standardization.\n",
" X_hat = (X - mean) / nd.sqrt(var + eps)\n",
"\n",
" # Update the mean and variance of the moving average.\n",
" moving_mean = momentum * moving_mean + (1.0 - momentum) * mean\n",
" moving_var = momentum * moving_var + (1.0 - momentum) * var\n",
"\n",
" Y = gamma * X_hat + beta # Scale and shift.\n",
" return Y, moving_mean, moving_var"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"- `BatchNorm` retains the scale parameter `gamma` and the shift parameter `beta` involved in gradient finding and iteration\n",
"- It also maintains the mean and variance obtained from the moving average, so that they can be used during model prediction. "
]
},
{
"cell_type": "code",
"execution_count": 2,
"metadata": {},
"outputs": [],
"source": [
"class BatchNorm(nn.Block):\n",
" def __init__(self, num_features, num_dims, **kwargs):\n",
" super(BatchNorm, self).__init__(**kwargs)\n",
" \n",
" if num_dims == 2:\n",
" shape = (1, num_features)\n",
" else:\n",
" shape = (1, num_features, 1, 1)\n",
" \n",
" # The scale parameter and the shift parameter involved in gradient finding and iteration are initialized to 0 and 1 respectively.\n",
" self.gamma = self.params.get('gamma', shape=shape, init=init.One())\n",
" self.beta = self.params.get('beta', shape=shape, init=init.Zero())\n",
"\n",
" # All the variables not involved in gradient finding and iteration are initialized to 0 on the CPU.\n",
" self.moving_mean = nd.zeros(shape)\n",
" self.moving_var = nd.zeros(shape)\n",
"\n",
" def forward(self, X):\n",
" # If X is not on the CPU, copy moving_mean and moving_var to the device where X is located.\n",
" if self.moving_mean.context != X.context:\n",
" self.moving_mean = self.moving_mean.copyto(X.context)\n",
" self.moving_var = self.moving_var.copyto(X.context)\n",
" \n",
" # Save the updated moving_mean and moving_var.\n",
" Y, self.moving_mean, self.moving_var = batch_norm(\n",
" X, \n",
" self.gamma.data(), \n",
" self.beta.data(), \n",
" self.moving_mean,\n",
" self.moving_var, \n",
" eps=1e-5, \n",
" momentum=0.9\n",
" )\n",
" return Y"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"- The `num_features` parameter required by the `BatchNorm` instance is the number of outputs for a fully connected layer and the number of output channels for a convolutional layer. \n",
"- The `num_dims` parameter also required by this instance is 2 for a fully connected layer and 4 for a convolutional layer."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.11.3 Use a Batch Normalization LeNet"
]
},
{
"cell_type": "code",
"execution_count": 3,
"metadata": {},
"outputs": [],
"source": [
"net = nn.Sequential()\n",
"net.add(\n",
" nn.Conv2D(6, kernel_size=5),\n",
" BatchNorm(6, num_dims=4),\n",
" nn.Activation('sigmoid'),\n",
" nn.MaxPool2D(pool_size=2, strides=2),\n",
" nn.Conv2D(16, kernel_size=5),\n",
" BatchNorm(16, num_dims=4),\n",
" nn.Activation('sigmoid'),\n",
" nn.MaxPool2D(pool_size=2, strides=2),\n",
" nn.Dense(120),\n",
" BatchNorm(120, num_dims=2),\n",
" nn.Activation('sigmoid'),\n",
" nn.Dense(84),\n",
" BatchNorm(84, num_dims=2),\n",
" nn.Activation('sigmoid'),\n",
" nn.Dense(10)\n",
")"
]
},
{
"cell_type": "code",
"execution_count": 5,
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"training on cpu(0)\n",
"epoch 1, loss 0.6653, train acc 0.761, test acc 0.806, time 142.5 sec\n",
"epoch 2, loss 0.3917, train acc 0.859, test acc 0.847, time 141.8 sec\n",
"epoch 3, loss 0.3493, train acc 0.874, test acc 0.854, time 142.9 sec\n",
"epoch 4, loss 0.3212, train acc 0.884, test acc 0.872, time 138.8 sec\n",
"epoch 5, loss 0.3033, train acc 0.889, test acc 0.861, time 136.9 sec\n"
]
}
],
"source": [
"lr = 1.0\n",
"num_epochs = 5\n",
"batch_size = 256\n",
"\n",
"ctx = gb.try_gpu()\n",
"\n",
"net.initialize(ctx=ctx, init=init.Xavier(), force_reinit=True)\n",
"\n",
"trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': lr})\n",
"\n",
"train_iter, test_iter = gb.load_data_fashion_mnist(batch_size)\n",
"\n",
"gb.train_ch5(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)"
]
},
{
"cell_type": "code",
"execution_count": 6,
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(\n",
" [1.3520054 1.3801662 1.8764832 1.4937813 0.93755937 1.8829043 ]\n",
" , \n",
" [ 0.6523215 0.2318771 0.2659682 0.7197848 -0.51127845 -2.0419025 ]\n",
" )"
]
},
"execution_count": 6,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"net[1].gamma.data().reshape((-1,)), net[1].beta.data().reshape((-1,))"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.11.4 Gluon Implementation for Batch Normalization"
]
},
{
"cell_type": "code",
"execution_count": 7,
"metadata": {},
"outputs": [],
"source": [
"net = nn.Sequential()\n",
"net.add(\n",
" nn.Conv2D(6, kernel_size=5),\n",
" nn.BatchNorm(),\n",
" nn.Activation('sigmoid'),\n",
" nn.MaxPool2D(pool_size=2, strides=2),\n",
" nn.Conv2D(16, kernel_size=5),\n",
" nn.BatchNorm(),\n",
" nn.Activation('sigmoid'),\n",
" nn.MaxPool2D(pool_size=2, strides=2),\n",
" nn.Dense(120),\n",
" nn.BatchNorm(),\n",
" nn.Activation('sigmoid'),\n",
" nn.Dense(84),\n",
" nn.BatchNorm(),\n",
" nn.Activation('sigmoid'),\n",
" nn.Dense(10)\n",
")"
]
},
{
"cell_type": "code",
"execution_count": null,
"metadata": {},
"outputs": [],
"source": [
"lr = 1.0\n",
"num_epochs = 5\n",
"batch_size = 256\n",
"\n",
"ctx = gb.try_gpu()\n",
"\n",
"net.initialize(ctx=ctx, init=init.Xavier(), force_reinit=True)\n",
"\n",
"trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': lr})\n",
"\n",
"train_iter, test_iter = gb.load_data_fashion_mnist(batch_size)\n",
"\n",
"gb.train_ch5(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"- During model training, batch normalization continuously adjusts the intermediate output of the neural network by utilizing the mean and standard deviation of the mini-batch, so that the values of the intermediate output in each layer throughout the neural network are more stable.\n",
"\n",
"- Like a dropout layer, batch normalization layers have different computation results in training mode and prediction mode.\n",
"\n",
"- Batch Normalization has many beneficial side effects, primarily that of regularization. \n",
"\n",
"- On the other hand, the original motivation of reducing covariate shift seems not to be a valid explanation."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"# 5.12 Residual Networks (ResNet)"
]
},
{
"attachments": {},
"cell_type": "markdown",
"metadata": {},
"source": [
"- Can we add a new layer to the neural network so that the fully trained model can reduce training errors more effectively?\n",
" - The added layer might make it easier to reduce training errors.\n",
"- In practice, however, with the addition of too many layers, training errors increase rather than decrease.\n",
" - Adding layers doesn't only make the network more expressive.\n",
"- Function Classes\n",
" - Consider $\\mathcal{F}$, the class of functions that a specific network architecture (together with learning rates and other hyperparameter settings) can reach. \n",
" - That is, for all $f \\in \\mathcal{F}$ there exists some set of parameters $W$ that can be obtained through training on a suitable dataset. \n",
" - Let's assume that $\\hat{f}$ is the function that we really would like to find. \n",
" \n",
" )\n",
" - Only if larger function classes contain the smaller ones are we guaranteed that increasing them strictly increases the expressive power of the network.\n",
"- He Kaiming and his colleagues proposed the ResNet.\n",
" - Papers\n",
" - He, K., Zhang, X., Ren, S., & Sun, J. (2016). Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition (pp. 770-778). https://arxiv.org/abs/1512.03385\n",
"\n",
" - He, K., Zhang, X., Ren, S., & Sun, J. (2016, October). Identity mappings in deep residual networks. In European Conference on Computer Vision (pp. 630-645). Springer, Cham. - https://arxiv.org/abs/1603.05027\n",
" - It won the ImageNet Visual Recognition Challenge in 2015\n",
" - It had a profound influence on the design of subsequent deep neural networks.\n",
" - At the heart of ResNet is the idea that ***every additional layer should contain the identity function as one of its elements***. \n",
" - This means that if we can train the newly-added layer into an identity mapping $f(\\mathbf{x}) = \\mathbf{x}$, the new model will be as effective as the original model. \n",
" - These considerations are rather profound but they led to a surprisingly simple solution, a ***residual block***"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.12.1 Residual Blocks\n",
"\n",
"- Each block can be expressed in a general form: $$ y_l = h(x_l) + F(x_l, W_l) \\\\ x_{l+1} = f(y_l) $$\n",
" - $x_l$ and $x_{l+1}$ are input and output of the $l$-th unit\n",
" - $F$ is a residual function. \n",
" - $h(x_l) = x_l$ is an identity mapping \n",
" - $f$ is a ReLU function\n",
"- The central idea of ResNets\n",
" - ***Learn the additive residual function $F$ with respect to an identity mapping $h(x_l) = x_l$***.\n",
" - This is realized by attaching an identity skip connection (“shortcut”)."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"- ResNet follows VGG's full $3\\times 3$ convolutional layer design. \n",
"- The residual block has two $3\\times 3$ convolutional layers with the same number of output channels. \n",
"- Each convolutional layer is followed by a batch normalization layer and a ReLU activation function. \n",
"- Then, we skip these two convolution operations and add the input directly before the final ReLU activation function. \n",
" - The output of the two convolutional layers should be of the same shape as the input, so that they can be added together. \n",
"- If we want to change the number of channels or the the stride, we need to introduce an additional $1\\times 1$ convolutional layer to transform the input into the desired shape for the addition operation. "
]
},
{
"cell_type": "code",
"execution_count": 1,
"metadata": {},
"outputs": [],
"source": [
"import gluonbook as gb\n",
"from mxnet import gluon, init, nd\n",
"from mxnet.gluon import nn\n",
"\n",
"class Residual(nn.Block): # This class is part of the gluonbook package\n",
" def __init__(self, num_channels, use_1x1conv=False, strides=1, **kwargs):\n",
" super(Residual, self).__init__(**kwargs)\n",
" self.conv1 = nn.Conv2D(num_channels, kernel_size=3, padding=1, strides=strides)\n",
" self.conv2 = nn.Conv2D(num_channels, kernel_size=3, padding=1)\n",
" self.bn1 = nn.BatchNorm()\n",
" self.bn2 = nn.BatchNorm()\n",
" if use_1x1conv:\n",
" self.conv3 = nn.Conv2D(num_channels, kernel_size=1, strides=strides)\n",
" else:\n",
" self.conv3 = None \n",
"\n",
" def forward(self, X):\n",
" Y = nd.relu(self.bn1(self.conv1(X)))\n",
" Y = self.bn2(self.conv2(Y))\n",
" if self.conv3:\n",
" X = self.conv3(X)\n",
" return nd.relu(Y + X)"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"- The above code generates two types of networks\n",
" - 1) Add the input to the output before applying the ReLu nonlinearity\n",
" - 2) Whenever `use_1x1conv=True`, adjust channels and resolution by means of a $1 \\times 1$ convolution before adding.\n",
""
]
},
{
"cell_type": "code",
"execution_count": 2,
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(4, 3, 6, 6)"
]
},
"execution_count": 2,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"blk = Residual(num_channels=3)\n",
"blk.initialize()\n",
"X = nd.random.uniform(shape=(4, 3, 6, 6))\n",
"blk(X).shape"
]
},
{
"cell_type": "code",
"execution_count": 3,
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(4, 6, 3, 3)"
]
},
"execution_count": 3,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"blk = Residual(num_channels=6, use_1x1conv=True, strides=2)\n",
"blk.initialize()\n",
"X = nd.random.uniform(shape=(4, 3, 6, 6))\n",
"blk(X).shape"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.12.2 ResNet Model"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
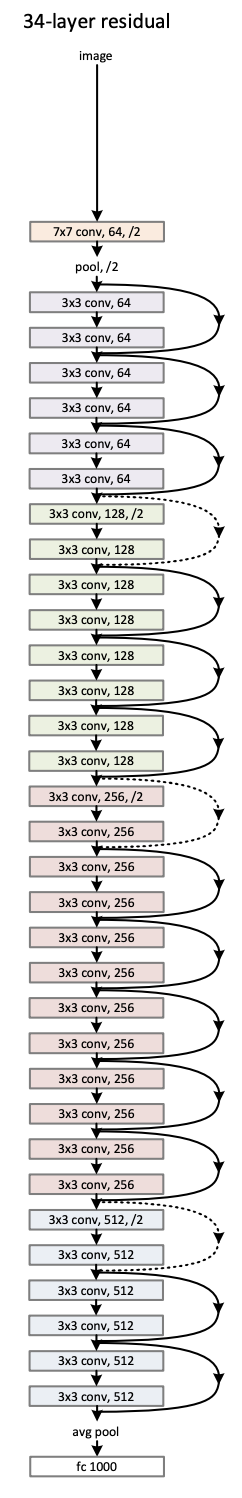
"- The first two layers of ResNet are the same as those of the GoogLeNet\n",
" - $7\\times 7$ convolutional layer with 64 output channels and a stride of 2\n",
" - Then, the $3\\times 3$ maximum pooling layer with a stride of 2 and padding of 1. \n",
"- The difference is the batch normalization layer added after each convolutional layer in ResNet."
]
},
{
"cell_type": "code",
"execution_count": 18,
"metadata": {},
"outputs": [],
"source": [
"net = nn.Sequential()\n",
"net.add(\n",
" nn.Conv2D(64, kernel_size=7, strides=2, padding=3),\n",
" nn.BatchNorm(), \n",
" nn.Activation('relu'),\n",
" nn.MaxPool2D(pool_size=3, strides=2, padding=1)\n",
")"
]
},
{
"cell_type": "code",
"execution_count": 19,
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"conv52 output shape:\t (1, 64, 14, 14)\n",
"batchnorm45 output shape:\t (1, 64, 14, 14)\n",
"relu1 output shape:\t (1, 64, 14, 14)\n",
"pool4 output shape:\t (1, 64, 7, 7)\n"
]
}
],
"source": [
"X = nd.random.uniform(shape=(1, 1, 28, 28))\n",
"net.initialize()\n",
"for layer in net:\n",
" X = layer(X)\n",
" print(layer.name, 'output shape:\\t', X.shape)"
]
},
{
"cell_type": "code",
"execution_count": 20,
"metadata": {},
"outputs": [],
"source": [
"net.add(\n",
" #Since a maximum pooling layer with a stride of 2 has already been used, \n",
" #it is not necessary to reduce the height and width at the first residual block.\n",
" Residual(num_channels=64), \n",
" Residual(num_channels=64),\n",
" Residual(num_channels=64), \n",
" Residual(num_channels=128, use_1x1conv=True, strides=2), # height and width are halved\n",
" Residual(num_channels=128),\n",
" Residual(num_channels=128), \n",
" Residual(num_channels=256, use_1x1conv=True, strides=2), # height and width are halved\n",
" Residual(num_channels=256),\n",
" Residual(num_channels=256), \n",
" Residual(num_channels=512, use_1x1conv=True, strides=2), # height and width are halved\n",
" Residual(num_channels=512),\n",
" Residual(num_channels=512) \n",
")"
]
},
{
"cell_type": "code",
"execution_count": 21,
"metadata": {},
"outputs": [],
"source": [
"net.add(\n",
" nn.GlobalAvgPool2D(),\n",
" nn.Dense(10)\n",
")"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
""
]
},
{
"cell_type": "code",
"execution_count": 23,
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"conv52 output shape:\t (1, 64, 14, 14)\n",
"batchnorm45 output shape:\t (1, 64, 14, 14)\n",
"relu1 output shape:\t (1, 64, 14, 14)\n",
"pool4 output shape:\t (1, 64, 7, 7)\n",
"residual22 output shape:\t (1, 64, 7, 7)\n",
"residual23 output shape:\t (1, 64, 7, 7)\n",
"residual24 output shape:\t (1, 64, 7, 7)\n",
"residual25 output shape:\t (1, 128, 4, 4)\n",
"residual26 output shape:\t (1, 128, 4, 4)\n",
"residual27 output shape:\t (1, 128, 4, 4)\n",
"residual28 output shape:\t (1, 256, 2, 2)\n",
"residual29 output shape:\t (1, 256, 2, 2)\n",
"residual30 output shape:\t (1, 256, 2, 2)\n",
"residual31 output shape:\t (1, 512, 1, 1)\n",
"residual32 output shape:\t (1, 512, 1, 1)\n",
"residual33 output shape:\t (1, 512, 1, 1)\n",
"pool5 output shape:\t (1, 512, 1, 1)\n",
"dense3 output shape:\t (1, 10)\n"
]
}
],
"source": [
"X = nd.random.uniform(shape=(1, 1, 28, 28))\n",
"net.initialize(force_reinit=True)\n",
"for layer in net:\n",
" X = layer(X)\n",
" print(layer.name, 'output shape:\\t', X.shape)"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.12.3 Data Acquisition and Training"
]
},
{
"cell_type": "code",
"execution_count": 24,
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"training on cpu(0)\n",
"epoch 1, loss 0.8086, train acc 0.760, test acc 0.861, time 1187.2 sec\n",
"epoch 2, loss 0.3464, train acc 0.872, test acc 0.863, time 1174.8 sec\n"
]
},
{
"name": "stderr",
"output_type": "stream",
"text": [
"Process Process-17:\n",
"Process Process-19:\n",
"Process Process-18:\n",
"Process Process-20:\n",
"Traceback (most recent call last):\n",
"Traceback (most recent call last):\n",
"Traceback (most recent call last):\n",
"Traceback (most recent call last):\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 93, in get\n",
" with self._rlock:\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 93, in get\n",
" with self._rlock:\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 94, in get\n",
" res = self._recv_bytes()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 93, in get\n",
" with self._rlock:\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/synchronize.py\", line 95, in __enter__\n",
" return self._semlock.__enter__()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/synchronize.py\", line 95, in __enter__\n",
" return self._semlock.__enter__()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/connection.py\", line 216, in recv_bytes\n",
" buf = self._recv_bytes(maxlength)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/synchronize.py\", line 95, in __enter__\n",
" return self._semlock.__enter__()\n",
"KeyboardInterrupt\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/connection.py\", line 407, in _recv_bytes\n",
" buf = self._recv(4)\n",
"KeyboardInterrupt\n",
"KeyboardInterrupt\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/connection.py\", line 379, in _recv\n",
" chunk = read(handle, remaining)\n",
"KeyboardInterrupt\n"
]
},
{
"ename": "KeyboardInterrupt",
"evalue": "",
"output_type": "error",
"traceback": [
"\u001b[0;31m---------------------------------------------------------------------------\u001b[0m",
"\u001b[0;31mKeyboardInterrupt\u001b[0m Traceback (most recent call last)",
"\u001b[0;32m\u001b[0m in \u001b[0;36m\u001b[0;34m\u001b[0m\n\u001b[1;32m 11\u001b[0m \u001b[0mtrain_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtest_iter\u001b[0m \u001b[0;34m=\u001b[0m \u001b[0mgb\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mload_data_fashion_mnist\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mbatch_size\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 12\u001b[0m \u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m---> 13\u001b[0;31m \u001b[0mgb\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mtrain_ch5\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mnet\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtrain_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtest_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mbatch_size\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtrainer\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mctx\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mnum_epochs\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0m",
"\u001b[0;32m~/anaconda3/envs/gluon/lib/python3.6/site-packages/gluonbook/utils.py\u001b[0m in \u001b[0;36mtrain_ch5\u001b[0;34m(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)\u001b[0m\n\u001b[1;32m 682\u001b[0m \u001b[0ml\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mbackward\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 683\u001b[0m \u001b[0mtrainer\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mstep\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mbatch_size\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m--> 684\u001b[0;31m \u001b[0mtrain_l_sum\u001b[0m \u001b[0;34m+=\u001b[0m \u001b[0ml\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mmean\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0masscalar\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0m\u001b[1;32m 685\u001b[0m \u001b[0mtrain_acc_sum\u001b[0m \u001b[0;34m+=\u001b[0m \u001b[0maccuracy\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0my_hat\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0my\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 686\u001b[0m \u001b[0mtest_acc\u001b[0m \u001b[0;34m=\u001b[0m \u001b[0mevaluate_accuracy\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mtest_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mnet\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mctx\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n",
"\u001b[0;32m~/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/ndarray/ndarray.py\u001b[0m in \u001b[0;36masscalar\u001b[0;34m(self)\u001b[0m\n\u001b[1;32m 1892\u001b[0m \u001b[0;32mif\u001b[0m \u001b[0mself\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mshape\u001b[0m \u001b[0;34m!=\u001b[0m \u001b[0;34m(\u001b[0m\u001b[0;36m1\u001b[0m\u001b[0;34m,\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m:\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1893\u001b[0m \u001b[0;32mraise\u001b[0m \u001b[0mValueError\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m\"The current array is not a scalar\"\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m-> 1894\u001b[0;31m \u001b[0;32mreturn\u001b[0m \u001b[0mself\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0masnumpy\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m[\u001b[0m\u001b[0;36m0\u001b[0m\u001b[0;34m]\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0m\u001b[1;32m 1895\u001b[0m \u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1896\u001b[0m \u001b[0;32mdef\u001b[0m \u001b[0mastype\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mself\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mdtype\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mcopy\u001b[0m\u001b[0;34m=\u001b[0m\u001b[0;32mTrue\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m:\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n",
"\u001b[0;32m~/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/ndarray/ndarray.py\u001b[0m in \u001b[0;36masnumpy\u001b[0;34m(self)\u001b[0m\n\u001b[1;32m 1874\u001b[0m \u001b[0mself\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mhandle\u001b[0m\u001b[0;34m,\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1875\u001b[0m \u001b[0mdata\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mctypes\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mdata_as\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mctypes\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mc_void_p\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m,\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m-> 1876\u001b[0;31m ctypes.c_size_t(data.size)))\n\u001b[0m\u001b[1;32m 1877\u001b[0m \u001b[0;32mreturn\u001b[0m \u001b[0mdata\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1878\u001b[0m \u001b[0;34m\u001b[0m\u001b[0m\n",
"\u001b[0;31mKeyboardInterrupt\u001b[0m: "
]
}
],
"source": [
"lr = 0.05\n",
"num_epochs = 5\n",
"batch_size = 256\n",
"\n",
"ctx = gb.try_gpu()\n",
"\n",
"net.initialize(force_reinit=True, ctx=ctx, init=init.Xavier())\n",
"\n",
"trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': lr})\n",
"\n",
"train_iter, test_iter = gb.load_data_fashion_mnist(batch_size)\n",
"\n",
"gb.train_ch5(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"- Residual blocks allow for a parametrization relative to the identity function $f(\\mathbf{x}) = \\mathbf{x}$.\n",
"- Adding residual blocks increases the function complexity in a well-defined manner.\n",
"- We can train an effective deep neural network by having residual blocks pass through cross-layer data channels.\n",
"- ResNet had a major influence on the design of subsequent deep neural networks, both for convolutional and sequential nature."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"# 5.13 Densely Connected Networks (DenseNet)\n",
"- DenseNet is a logical extension of DenseNet. \n",
"- To understand how to arrive at it, let's take a small detour to theory. \n",
"- Recall the Taylor expansion for functions. $$f(x) = f(0) + f'(x) x + \\frac{1}{2} f''(x) x^2 + \\frac{1}{6} f'''(x) x^3 + o(x^3)$$\n",
"- Function Decomposition\n",
" - It decomposes the function into increasingly higher order terms. \n",
" - ResNet decomposes functions into $$f(\\mathbf{x}) = \\mathbf{x} + g(\\mathbf{x})$$\n",
" - That is, ResNet decomposes $f$ into a simple linear term and a more complex nonlinear one. \n",
" - What if we want to go beyond two terms?\n",
" - A solution was proposed by Huang et al, 2016 in the form of DenseNet.\n",
" - Huang, G., Liu, Z., Weinberger, K. Q., & van der Maaten, L. (2017). Densely connected con- volutional networks. In Proceedings of the IEEE conference on computer vision and pattern recognition (Vol. 1, No. 2). https://arxiv.org/abs/1608.06993\n",
"\n",
"\n",
"- The key difference between ResNet and DenseNet\n",
" - In DenseNet, outputs are concatenated rather than added. \n",
" - As a result we perform a mapping from $\\mathbf{x}$ to its values after applying an increasingly complex sequence of functions. $$\\mathbf{x} \\to \\left[\\mathbf{x}, f_1(\\mathbf{x}), f_2(\\mathbf{x}, f_1(\\mathbf{x})), f_3(\\mathbf{x}, f_1(\\mathbf{x}), f_2(\\mathbf{x}, f_1(\\mathbf{x})), \\ldots\\right]$$\n",
" - In the end, all these functions are combined in an MLP to reduce the number of features again. \n",
" - The name DenseNet arises from the fact that...\n",
" - The dependency graph between variables becomes quite dense. \n",
" - The last layer of such a chain is densely connected to all previous layers.\n",
" - The main components that compose a DenseNet are dense blocks and transition layers. \n",
" - The dense block defines how the inputs and outputs are concatenated, \n",
" - The transition layers control the number of channels so that it is not too large."
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.13.1 Dense Blocks"
]
},
{
"cell_type": "code",
"execution_count": 25,
"metadata": {},
"outputs": [],
"source": [
"import gluonbook as gb\n",
"from mxnet import gluon, init, nd\n",
"from mxnet.gluon import nn\n",
"\n",
"def conv_block(num_channels):\n",
" blk = nn.Sequential()\n",
" blk.add(\n",
" nn.BatchNorm(), \n",
" nn.Activation('relu'),\n",
" nn.Conv2D(num_channels, kernel_size=3, padding=1)\n",
" )\n",
" return blk"
]
},
{
"cell_type": "code",
"execution_count": 33,
"metadata": {},
"outputs": [],
"source": [
"class DenseBlock(nn.Block):\n",
" def __init__(self, num_convs, num_channels, **kwargs):\n",
" super(DenseBlock, self).__init__(**kwargs)\n",
" self.net = nn.Sequential()\n",
" for _ in range(num_convs):\n",
" self.net.add(conv_block(num_channels))\n",
"\n",
" def forward(self, X):\n",
" for blk in self.net:\n",
" Y = blk(X)\n",
" print(\"X.shape: {0} --> Y.shape: {1}\".format(X.shape, Y.shape)) \n",
" X = nd.concat(X, Y, dim=1) # Concatenate the input and output of each block on the channel dimension.\n",
" return X"
]
},
{
"cell_type": "code",
"execution_count": 34,
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"X.shape: (4, 3, 8, 8) --> Y.shape: (4, 10, 8, 8)\n",
"X.shape: (4, 13, 8, 8) --> Y.shape: (4, 10, 8, 8)\n"
]
},
{
"data": {
"text/plain": [
"(4, 23, 8, 8)"
]
},
"execution_count": 34,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"blk = DenseBlock(num_convs=2, num_channels=10)\n",
"blk.initialize(force_reinit=True)\n",
"X = nd.random.uniform(shape=(4, 3, 8, 8))\n",
"Y = blk(X)\n",
"Y.shape"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.13.2 Transition Layers\n",
"- Since each dense block will increase the number of channels, adding too many of them will lead to an excessively complex model. \n",
"- A transition layer is used to control the complexity of the model. \n",
"- It reduces the number of channels by using the $1\\times 1$ convolutional layer and halves the height and width of the average pooling layer with a stride of 2, further reducing the complexity of the model."
]
},
{
"cell_type": "code",
"execution_count": 35,
"metadata": {},
"outputs": [],
"source": [
"def transition_block(num_channels):\n",
" blk = nn.Sequential()\n",
" blk.add(\n",
" nn.BatchNorm(), \n",
" nn.Activation('relu'),\n",
" nn.Conv2D(num_channels, kernel_size=1),\n",
" nn.AvgPool2D(pool_size=2, strides=2)\n",
" )\n",
" return blk"
]
},
{
"cell_type": "code",
"execution_count": 37,
"metadata": {},
"outputs": [
{
"data": {
"text/plain": [
"(4, 10, 4, 4)"
]
},
"execution_count": 37,
"metadata": {},
"output_type": "execute_result"
}
],
"source": [
"blk = transition_block(num_channels=10)\n",
"blk.initialize(force_reinit=True)\n",
"blk(Y).shape"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.13.3 DenseNet Model"
]
},
{
"cell_type": "code",
"execution_count": 38,
"metadata": {},
"outputs": [],
"source": [
"net = nn.Sequential()\n",
"net.add(\n",
" nn.Conv2D(64, kernel_size=7, strides=2, padding=3),\n",
" nn.BatchNorm(), \n",
" nn.Activation('relu'),\n",
" nn.MaxPool2D(pool_size=3, strides=2, padding=1)\n",
")"
]
},
{
"cell_type": "code",
"execution_count": 39,
"metadata": {},
"outputs": [],
"source": [
"num_channels = 64 # Num_channels: the current number of channels.\n",
"growth_rate = 32 \n",
"num_convs_in_dense_blocks = [4, 4, 4, 4]\n",
"\n",
"for i, num_convs in enumerate(num_convs_in_dense_blocks):\n",
" net.add(DenseBlock(num_convs=num_convs, num_channels=growth_rate))\n",
" \n",
" # This is the number of output channels in the previous dense block.\n",
" num_channels += num_convs * growth_rate\n",
"\n",
" # A transition layer that haves the number of channels is added between the dense blocks.\n",
" if i != len(num_convs_in_dense_blocks) - 1:\n",
" net.add(transition_block(num_channels // 2))"
]
},
{
"cell_type": "code",
"execution_count": 40,
"metadata": {},
"outputs": [],
"source": [
"net.add(\n",
" nn.BatchNorm(), \n",
" nn.Activation('relu'), \n",
" nn.GlobalAvgPool2D(),\n",
" nn.Dense(10)\n",
")"
]
},
{
"cell_type": "markdown",
"metadata": {},
"source": [
"## 5.13.4 Data Acquisition and Training"
]
},
{
"cell_type": "code",
"execution_count": 42,
"metadata": {},
"outputs": [
{
"name": "stdout",
"output_type": "stream",
"text": [
"training on cpu(0)\n",
"X.shape: (256, 64, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 96, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 128, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 160, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 96, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 128, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 160, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 192, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 160, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 192, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 224, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 256, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 224, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 256, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 288, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 320, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 64, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 96, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 128, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 160, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 96, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 128, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 160, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 192, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 160, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 192, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 224, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 256, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 224, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 256, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 288, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 320, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 64, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 96, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 128, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 160, 24, 24) --> Y.shape: (256, 32, 24, 24)\n",
"X.shape: (256, 96, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 128, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 160, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 192, 12, 12) --> Y.shape: (256, 32, 12, 12)\n",
"X.shape: (256, 160, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 192, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 224, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 256, 6, 6) --> Y.shape: (256, 32, 6, 6)\n",
"X.shape: (256, 224, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 256, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 288, 3, 3) --> Y.shape: (256, 32, 3, 3)\n",
"X.shape: (256, 320, 3, 3) --> Y.shape: (256, 32, 3, 3)\n"
]
},
{
"name": "stderr",
"output_type": "stream",
"text": [
"Process Process-26:\n",
"Process Process-28:\n",
"Process Process-25:\n",
"Process Process-27:\n",
"Traceback (most recent call last):\n",
"Traceback (most recent call last):\n",
"Traceback (most recent call last):\n",
"Traceback (most recent call last):\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 258, in _bootstrap\n",
" self.run()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/process.py\", line 93, in run\n",
" self._target(*self._args, **self._kwargs)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 93, in get\n",
" with self._rlock:\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/gluon/data/dataloader.py\", line 116, in worker_loop\n",
" idx, samples = key_queue.get()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 94, in get\n",
" res = self._recv_bytes()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 93, in get\n",
" with self._rlock:\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/queues.py\", line 93, in get\n",
" with self._rlock:\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/connection.py\", line 216, in recv_bytes\n",
" buf = self._recv_bytes(maxlength)\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/synchronize.py\", line 95, in __enter__\n",
" return self._semlock.__enter__()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/synchronize.py\", line 95, in __enter__\n",
" return self._semlock.__enter__()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/synchronize.py\", line 95, in __enter__\n",
" return self._semlock.__enter__()\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/connection.py\", line 407, in _recv_bytes\n",
" buf = self._recv(4)\n",
"KeyboardInterrupt\n",
"KeyboardInterrupt\n",
"KeyboardInterrupt\n",
" File \"/Users/yhhan/anaconda3/envs/gluon/lib/python3.6/multiprocessing/connection.py\", line 379, in _recv\n",
" chunk = read(handle, remaining)\n",
"KeyboardInterrupt\n"
]
},
{
"ename": "KeyboardInterrupt",
"evalue": "",
"output_type": "error",
"traceback": [
"\u001b[0;31m---------------------------------------------------------------------------\u001b[0m",
"\u001b[0;31mKeyboardInterrupt\u001b[0m Traceback (most recent call last)",
"\u001b[0;32m\u001b[0m in \u001b[0;36m\u001b[0;34m\u001b[0m\n\u001b[1;32m 11\u001b[0m \u001b[0mtrain_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtest_iter\u001b[0m \u001b[0;34m=\u001b[0m \u001b[0mgb\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mload_data_fashion_mnist\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mbatch_size\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mresize\u001b[0m\u001b[0;34m=\u001b[0m\u001b[0;36m96\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 12\u001b[0m \u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m---> 13\u001b[0;31m \u001b[0mgb\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mtrain_ch5\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mnet\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtrain_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtest_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mbatch_size\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mtrainer\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mctx\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mnum_epochs\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0m",
"\u001b[0;32m~/anaconda3/envs/gluon/lib/python3.6/site-packages/gluonbook/utils.py\u001b[0m in \u001b[0;36mtrain_ch5\u001b[0;34m(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)\u001b[0m\n\u001b[1;32m 682\u001b[0m \u001b[0ml\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mbackward\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 683\u001b[0m \u001b[0mtrainer\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mstep\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mbatch_size\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m--> 684\u001b[0;31m \u001b[0mtrain_l_sum\u001b[0m \u001b[0;34m+=\u001b[0m \u001b[0ml\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mmean\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0masscalar\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0m\u001b[1;32m 685\u001b[0m \u001b[0mtrain_acc_sum\u001b[0m \u001b[0;34m+=\u001b[0m \u001b[0maccuracy\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0my_hat\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0my\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 686\u001b[0m \u001b[0mtest_acc\u001b[0m \u001b[0;34m=\u001b[0m \u001b[0mevaluate_accuracy\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mtest_iter\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mnet\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mctx\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n",
"\u001b[0;32m~/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/ndarray/ndarray.py\u001b[0m in \u001b[0;36masscalar\u001b[0;34m(self)\u001b[0m\n\u001b[1;32m 1892\u001b[0m \u001b[0;32mif\u001b[0m \u001b[0mself\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mshape\u001b[0m \u001b[0;34m!=\u001b[0m \u001b[0;34m(\u001b[0m\u001b[0;36m1\u001b[0m\u001b[0;34m,\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m:\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1893\u001b[0m \u001b[0;32mraise\u001b[0m \u001b[0mValueError\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m\"The current array is not a scalar\"\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m-> 1894\u001b[0;31m \u001b[0;32mreturn\u001b[0m \u001b[0mself\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0masnumpy\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m[\u001b[0m\u001b[0;36m0\u001b[0m\u001b[0;34m]\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0m\u001b[1;32m 1895\u001b[0m \u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1896\u001b[0m \u001b[0;32mdef\u001b[0m \u001b[0mastype\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mself\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mdtype\u001b[0m\u001b[0;34m,\u001b[0m \u001b[0mcopy\u001b[0m\u001b[0;34m=\u001b[0m\u001b[0;32mTrue\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m:\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n",
"\u001b[0;32m~/anaconda3/envs/gluon/lib/python3.6/site-packages/mxnet/ndarray/ndarray.py\u001b[0m in \u001b[0;36masnumpy\u001b[0;34m(self)\u001b[0m\n\u001b[1;32m 1874\u001b[0m \u001b[0mself\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mhandle\u001b[0m\u001b[0;34m,\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1875\u001b[0m \u001b[0mdata\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mctypes\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mdata_as\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0mctypes\u001b[0m\u001b[0;34m.\u001b[0m\u001b[0mc_void_p\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m,\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[0;32m-> 1876\u001b[0;31m ctypes.c_size_t(data.size)))\n\u001b[0m\u001b[1;32m 1877\u001b[0m \u001b[0;32mreturn\u001b[0m \u001b[0mdata\u001b[0m\u001b[0;34m\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n\u001b[1;32m 1878\u001b[0m \u001b[0;34m\u001b[0m\u001b[0m\n",
"\u001b[0;31mKeyboardInterrupt\u001b[0m: "
]
}
],
"source": [
"lr = 0.1\n",
"num_epochs = 5\n",
"batch_size = 256\n",
"\n",
"ctx = gb.try_gpu()\n",
"\n",
"net.initialize(ctx=ctx, init=init.Xavier(), force_reinit=True)\n",
"\n",
"trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': lr})\n",
"\n",
"train_iter, test_iter = gb.load_data_fashion_mnist(batch_size, resize=96)\n",
"\n",
"gb.train_ch5(net, train_iter, test_iter, batch_size, trainer, ctx, num_epochs)"
]
},
{
"cell_type": "code",
"execution_count": null,
"metadata": {},
"outputs": [],
"source": []
}
],
"metadata": {
"kernelspec": {
"display_name": "Python 3",
"language": "python",
"name": "python3"
},
"language_info": {
"codemirror_mode": {
"name": "ipython",
"version": 3
},
"file_extension": ".py",
"mimetype": "text/x-python",
"name": "python",
"nbconvert_exporter": "python",
"pygments_lexer": "ipython3",
"version": "3.6.7"
}
},
"nbformat": 4,
"nbformat_minor": 2
}