{width=500px height=250px}
A random splitting into two halves: left part is training set, right part is validation set. ::: ::: {#fig-validation-auto}{width=600px height=350px}
The validation set approach was used on the `Auto` data set in order to estimate the test error that results from predicting `mpg` using polynomial functions of `horsepower`. Left: Validation error estimates for a single split into training and validation data sets. Right: The validation method was repeated ten times, each time using a different random split of the observations into a training set and a validation set. This illustrates the variability in the estimated test MSE that results from this approach. ::: - Drawbacks of validation set approach - The validation estimate of the test error can be **highly variable**, depending on precisely which observations are included in the training set and which observations are included in the validation set. - In the validation approach, only a subset of the observations - those that are included in the training set rather than in the validation set - are used to fit the model. - This suggests that the validation set error may tend to **overestimate the test error** for the model fit on the entire data set. (Why?) ## $K$-fold cross-validation - **Widely used approach** for estimating test error. - Estimates can be used to select best model, and to give an idea of the test error of the final chosen model. - Idea is to randomly divide the data into $K$ equal-sized parts. We leave out part $k$, fit the model to the other $K-1$ parts (combined), and then obtain predictions for the left-out $k$th part. - This is done in turn for each part $k = 1, 2, \ldots, K$, and then the results are combined. ::: {#fig-10-fold-cv-auto}{width=600px height=350px}
A schematic display of 5-fold CV. A set of $n$ observations is randomly split into five non-overlapping groups. Each of these fifths acts as a validation set (shown in beige), and the remainder as a training set (shown in blue). The test error is estimated by averaging the five resulting MSE estimates. ::: - Let the $K$ parts be $C_1, C_2, \ldots, C_K$, where $C_k$ denotes the indices of the observations in part $k$. There are $n_k$ observations in part $k$. If $N$ is a multiple of $K$, then $n_k = n / K$. - Compute $$ \text{CV}_{(K)} = \sum_{k=1}^K \frac{n_k}{n} \text{MSE}_k, $$ where $$ \text{MSE}_k = \frac{1}{n_k} \sum_{i \in C_k} (y_i - \hat y_i)^2, $$ and $\hat y_i$ is the fit for observation $i$, obtained from the data with part $k$ removed. ### LOOCV - The special case $K=n$ yields $n$-fold or **leave-one out cross-validation (LOOCV)**. - With least-squares linear or polynomial regression, an amazing shortcut makes the cost of LOOCV the same as that of a single model fit! $$ \text{CV}_{(n)} = \frac{1}{n} \sum_{i=1}^n \left( \frac{y_i - \hat y_i}{1 - h_i} \right)^2, $$ where $\hat y_i$ is the $i$th fitted value from the original least squares fit, and $h_i$ is the leverage (diagonal of the "hat" matrix, i.e., $\mathbf{H} = \mathbf{X}(\mathbf{X}^T\mathbf{X})^{–1}\mathbf{X}^T$). This is like the ordinary MSE, except the $i$th residual is divided by $1 - h_i$. - LOOCV sometimes useful, but typically doesn't shake up the data enough. The estimates from each fold are highly correlated and hence their average can have high variance. - A better choice is $K = 5$ or 10. ::: {#fig-10-fold-cv-auto}{width=600px height=350px}
Cross-validation was used on the `Auto` data set in order to estimate the test error that results from predicting `mpg` using polynomial functions of `horsepower`. Left: The LOOCV error curve. Right: 10-fold CV was run nine separate times, each with a different random split of the data into ten parts. The figure shows the nine slightly different CV error curves. ::: ::: {#fig-true-vs-estimated-test-MSE}{width=600px height=350px}
Blue: true test MSE. Black dashed line: LOOCV estimate of test MSE. Orange: 10-fold CV estimate of test MSE. The crosses indicate the minimum of each of the MSE curves. [Left data](https://ucla-biostat-212a.github.io/2026winter/slides/02-statlearn/statlearn.html#fig-tradeoff-truth). [Middle data](https://ucla-biostat-212a.github.io/2026winter/slides/02-statlearn/statlearn.html#fig-tradeoff-smooth-truth). [Right data](https://ucla-biostat-212a.github.io/2026winter/slides/02-statlearn/statlearn.html#fig-tradeoff-wiggly-truth). ::: ## Bias-variance tradeoff for cross-validation - Since each training set is only $(K - 1) / K$ as big as the original training set, the estimates of prediction error will typically be biased upward. (Why?) - This bias is minimized when $K = n$ (LOOCV), but this estimate has high variance, as noted earlier. - $K = 5$ or $10$ provides a good compromise for this bias-variance trade-off. ## Cross-validation for classification problems - We divide the data into $K$ roughly equal-sized parts $C_1, C_2, \ldots, C_K$. $C_k$ denotes the indices of the observations in part $k$. There are $n_k$ observations in part $k$. If $n$ is a multiple of $K$, then $n_k = n / K$. - Compute $$ \text{CV}_k = \sum_{k=1}^K \frac{n_k}{n} \text{Err}_k, $$ where $$ \text{Err}_k = \frac{1}{n_k} \sum_{i \in C_k} I(y_i \ne \hat y_i). $$ - The estimated standard deviation of $\text{CV}_k$ is $$ \hat{\text{SE}}(\text{CV}_k) = \sqrt{\frac{1}{K} \sum_{k=1}^K \frac{(\text{Err}_k - \bar{\text{Err}_k})^2}{K-1}}. $$ This is a useful estimate, but strictly speaking, not quite valid. ## Bootstrap - The bootstrap is a flexible and powerful statistical tool that can be used to quantify the uncertainty associated with a given estimator or learning method. - For example, it can provide an estimate of the standard error of a coefficient, or a confidence interval for that coefficient. - A simple example. - Suppose that we wish to invest a fixed sum of money in two financial assets that yield returns of $X$ and $Y$, respectively, where $X$ and $Y$ are random quantities. - We will invest a fraction $\alpha$ of our money in $X$, and will invest the remaining $1 - \alpha$ in $Y$. - We wish to choose $\alpha$ to minimize the total risk, or variance, of our investment. In other words, we want to minimize $\operatorname{Var}(\alpha X + (1 - \alpha) Y)$. - One can show that the value that minimizes the risk is given by $$ \alpha = \frac{\sigma_{Y}^2 - \sigma_{XY}}{\sigma_X^2 + \sigma_Y^2 - 2 \sigma_{XY}}, $$ where $\sigma_X^2 = \operatorname{Var}(X)$, $\sigma_Y^2 = \operatorname{Var}(Y)$, and $\sigma_{XY} = \operatorname{Cov}(X, Y)$. - But the values of $\sigma_X^2$, $\sigma_Y^2$, and $\sigma_{XY}$ are unknown. - We can compute estimates for these quantities $\hat{\sigma}_X^2$, $\hat{\sigma}_Y^2$, and $\hat{\sigma}_{XY}$, using a data set that contains measurements for $X$ and $Y$. - We can then estimate the value of $\alpha$ that minimizes the variance of our investment using $$ \hat{\alpha} = \frac{\hat{\sigma}_{Y}^2 - \hat{\sigma}_{XY}}{\hat{\sigma}_X^2 + \hat{\sigma}_Y^2 - 2 \hat{\sigma}_{XY}}. $$ ::: {#fig-unrealistic-toy}{width=600px height=600px}
Each panel displays 100 simulated returns for investments $X$ and $Y$. From left to right and top to bottom, the resulting estimates for $\alpha$ are 0.576, 0.532, 0.657, and 0.651. ::: - An unrealistic method: - To estimate the standard deviation of $\hat{\alpha}$, we repeated the process of simulating 100 paired observations of $X$ and $Y$, and estimating $\alpha$ 1,000 times. - We thereby obtained 1,000 estimates for $\alpha$, which we can call $\hat{\alpha}_1, \hat{\alpha}_2, \ldots, \hat{\alpha}_{1000}$. - For these simulations the parameters were set to $\sigma_{X}^2 = 1$, $\sigma_Y^2 = 1.25$, and $\sigma_{XY} = 0.5$, and so we know that the true value of $\alpha$ is 0.6. - The mean over 1,000 estimates for $\alpha$ is $$ \bar{\alpha} = \frac{1}{1000} \sum_{r=1}^{1000} \hat{\alpha}_r = 0.5996, $$ very close to $\alpha = 0.6$, and the standard deviation of the estimates is $$ \sqrt{\frac{1}{1000-1} \sum_{r=1}^{1000} (\hat{\alpha}_r - \bar{\alpha})^2} = 0.083. $$ This gives us a very good idea of the accuracy of $\hat{\alpha}$: $\text{SE}(\hat{\alpha}) \approx 0.083$. So roughly speaking, for a random sample from the population, we would expect $\hat{\alpha}$ to differ from $\alpha$ by approximately 0.08, on average. ::: {#fig-bootstrap-toy}{width=600px height=600px}
A graphical illustration of the bootstrap approach on a small sample containing $n = 3$ observations. Each bootstrap data set contains $n$ observations, sampled with replacement from the original data set. Each bootstrap data set is used to obtain an estimate of $\alpha$. ::: - **Bootstrap** method. - The procedure outlined above cannot be applied, because for real data we cannot generate new samples from the original population. - However, the bootstrap approach allows us to use a computer to mimic the process of obtaining new data sets, so that we can estimate the variability of our estimate without generating additional samples. - Rather than repeatedly obtaining independent data sets from the population, we instead obtain distinct data sets by repeatedly sampling observations from the original data set **with replacement**. - Each of these "bootstrap data sets" is created by sampling with replacement, and is the same size as our original dataset. As a result some observations may appear more than once in a given bootstrap data set and some not at all. - Denoting the first bootstrap data set by $Z^{*1}$, we use $Z^{*1}$ to produce a new bootstrap estimate for $\alpha$, which we call $\hat{\alpha}^{*1}$. - This procedure is repeated $B$ times for some large value of $B$ (say 100 or 1000), in order to produce $B$ different bootstrap data sets $Z^{*1}, Z^{*2}, \ldots, Z^{*B}$, and $B$ corresponding $\alpha$ estimates, $\hat{\alpha}^{*1}, \hat{\alpha}^{*2}, \ldots, \hat{\alpha}^{*B}$. - We estimate the standard error of these bootstrap estimates using the formula $$ \operatorname{SE}_B(\hat{\alpha}) = \sqrt{\frac{1}{B-1} \sum_{r=1}^B (\hat{\alpha}^{*r} - \bar{\hat{\alpha}}^{*})^2}. $$ - This serves as an estimate of the standard error of $\hat{\alpha}$ estimated from the original data set. For this example, $\operatorname{SE}_B(\hat{\alpha}) = 0.087$. ::: {#fig-unrealistic-vs-bootstrap-toy}{width=500px height=350px}
Left: A histogram of the estimates of $\alpha$ obtained by generating 1,000 simulated data sets from the true population. Center: A histogram of the estimates of $\alpha$ obtained from 1,000 bootstrap samples from a single data set. Right: The estimates of $\alpha$ displayed in the left and center panels are shown as boxplots. In each panel, the pink line indicates the true value of $\alpha$. ::: ## Bootstrap in general ::: {#fig-bootstrap-general-pic}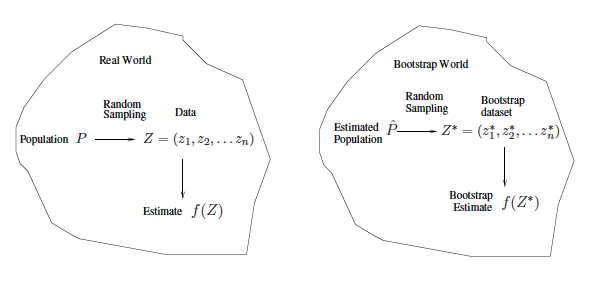{width=500px}
Bootstrap general scheme. ::: - In more complex data situations, figuring out the appropriate way to generate bootstrap samples can require some thought. For example, if the data is a time series (e.g., stock prices), we can't simply sample the observations with replacement (why not?). We can instead create blocks of consecutive observations, and sample those with replacements. Then we paste together sampled blocks to obtain a bootstrap dataset. - Other uses of the bootstrap. - Primarily used to obtain standard errors of an estimate. - Also provides approximate confidence intervals for a population parameter. For example, looking at the histogram in the middle panel of histogram, the 5% and 95% quantiles of the 1000 values is (0.43, 0.72). - This is called a **Bootstrap Percentile confidence interval**. It is the simplest method (among many approaches) for obtaining a confidence interval from the bootstrap. - Can the bootstrap estimate prediction error? - In cross-validation, each of the $K$ validation folds is distinct from the other $K-1$ folds used for training: there is no overlap. This is crucial for its success. - To estimate prediction error using the bootstrap, we could think about using each bootstrap dataset as our training sample, and the original sample as our validation sample. - But each bootstrap sample has significant overlap with the original data. About two-thirds of the original data points appear in each bootstrap sample. - This will cause the bootstrap to seriously underestimate the true prediction error. - The other way around - with original sample = training sample, bootstrap dataset = validation sample - is worse! - Can partly fix this problem by only using predictions for those observations that did not (by chance) occur in the current bootstrap sample. - But the method gets complicated, and in the end, cross-validation provides a simpler, more attractive approach for estimating prediction error. ## Lab ### The validation set approach - Goal: use of the validation set approach to estimate the test error rates that from fitting linear models on the `Auto` data set. - We begin by using the `set.seed()` function in order to set a seed for `R`'s random number generator, so that the reader of this book will obtain the same results as those shown here. - We later (during the weekend) will talk about pipelines that we do all of these automatically. ```{r} library(ISLR2) library(tidyverse) set.seed(1) # sample() function is used to randomly sample the integers from 1 to 392, without replacement train <- sample(392, 196) (train <- sort(train)) ``` - Use the `subset` option in `lm()` to fit a linear regression using only the observations corresponding to the training set. ```{r} ggplot(Auto, aes(x = horsepower, y = mpg)) + geom_point() + geom_smooth() ``` ```{r} lm.fit <- lm(mpg ~ horsepower, data = Auto, subset = train) ``` - Use the `predict()` function Compute the mean squared error of the model on the validation set. ```{r} mean((Auto$mpg[-train] - predict(lm.fit, Auto[-train, ]))^2) #Auto %>% select(mpg, horsepower) %>% slice(-train) %>% # mutate(pred = predict(lm.fit, Auto[-train,])) # %>% # ggplot(aes(x = mpg, y = pred)) + # geom_point() + # geom_smooth(method = "lm") + #geom_abline(intercept = 0, slope = 1, color = "red") + # labs(title = "Validation set approach", x = "Observed mpg", y = "Predicted mpg") ``` Therefore, the estimated test MSE for the linear regression fit is 23.27. - We can use the `poly()` function to estimate the test error for the quadratic and cubic regressions. ```{r} lm.fit2 <- lm(mpg ~ poly(horsepower, 2), data = Auto, subset = train) mean((Auto$mpg - predict(lm.fit2, Auto))[-train]^2) #Auto %>% select(mpg, horsepower) %>% slice(-train) %>% # mutate(pred = predict(lm.fit2, Auto[-train,])) ``` ```{r} lm.fit3 <- lm(mpg ~ poly(horsepower, 3), data = Auto, subset = train) mean((Auto$mpg - predict(lm.fit3, Auto))[-train]^2) ``` - The estimated test MSE for the quadratic and cubic regressions are 18.716 and 18.794, respectively. - These results are not surprising, given that the relationship between `mpg` and `horsepower` appears to be fairly linear. - If we choose a different training set instead, then we will obtain somewhat different errors on the validation set. ```{r} set.seed(2) train <- sample(392, 196) lm.fit <- lm(mpg ~ horsepower, data = Auto, subset = train) mean((Auto$mpg[-train] - predict(lm.fit, Auto[-train,]))^2) lm.fit2 <- lm(mpg ~ poly(horsepower, 2), data = Auto, subset = train) mean((Auto$mpg - predict(lm.fit2, Auto))[-train]^2) lm.fit3 <- lm(mpg ~ poly(horsepower, 3), data = Auto, subset = train) mean((Auto$mpg - predict(lm.fit3, Auto))[-train]^2) ``` - These results are consistent with our previous findings: + a model that predicts mpg using a **quadratic** function of horsepower performs **better** than a model that involves only a **linear** function of horsepower, + there is little evidence in favor of a model that uses a cubic function of horsepower. ### Leave-One-Out Cross-Validation - The LOOCV estimate can be automatically computed for any generalized linear model using the `glm()` and `cv.glm()` functions. - In the lab, we use the `glm()` function to perform **logistic regression** by passing in the `family="binomial"` argument. - If we leave it blank, it will perform linear regression. - We use the `cv.glm()` function in `boot` library to perform **LOOCV** for the previous example ```{r} library(boot) glm.fit <- glm(mpg ~ horsepower, data = Auto) cv.err <- cv.glm(Auto, glm.fit) cv.err$delta ``` - The `cv.glm()` function produces a list with several components. The `delta` component contains LOOCV statistic given in (5.1). - We can repeat this procedure for increasingly complex polynomial fits. ```{r} cv.error <- rep(0, 10) for (i in 1:10) { glm.fit <- glm(mpg ~ poly(horsepower, i), data = Auto) cv.error[i] <- cv.glm(Auto, glm.fit)$delta[1] } cv.error plot(cv.error, type = "b") ``` ### k-Fold Cross-Validation - The `cv.glm()` function can also be used to perform **k-fold CV**. - Below, we use `k=10` to perform 10-fold CV. ```{r} set.seed(17) cv.error.10 <- rep(0, 10) for (i in 1:10) { glm.fit <- glm(mpg ~ poly(horsepower, i), data = Auto) cv.error.10[i] <- cv.glm(Auto, glm.fit, K = 10)$delta[1] } cv.error.10 plot(cv.error.10, type = "b") ``` - We see that the computation time is much shorter than for LOOCV. - Due to the availability of the formula (5.2) for LOOCV; however, unfortunately the `cv.glm()` function does not make use of this formula. - Two numbers associated with delta are the same when LOOCV is performed. When we instead perform k-fold CV, then the two numbers associated with delta differ slightly. The first is the standard k-fold CV estimate, as in equation (5.3). The second is a bias-corrected version. ### The Bootstrap - Performing a bootstrap analysis in R entails only two steps: + First, we must create a function that computes the statistic of interest. + Second, we use the `boot()` function to perform the bootstrap by repeatedly sampling observations from the data set **with replacement**. - The `Portfolio` data set in the ISLR2 package is simulated data of 100 pairs of returns,