Chapter 6: WRF Data Assimilation
Table of Contents
- Introduction
- Installing WRFDA for 3D-Var Run
- Installing WRFPLUS and WRFDA for 4D-Var Run
- Running Observation Preprocessor (OBSPROC)
- Running WRFDA
- Radiance Data Assimilations in WRFDA
- WRFDA Diagnostics
- Updating WRF Boundary Conditions
- Running gen_be
- WRFDA with Multivariate Background Error (MBE) Statistics
- Additional WRFDA Exercises
- Hybrid Data Assimilation
- Description of Namelist Variables
Introduction
Data assimilation is the technique by which observations are combined with a NWP product (the first guess or background forecast) and their respective error statistics to provide an improved estimate (the analysis) of the atmospheric (or oceanic, Jovian, whatever) state. Variational (Var) data assimilation achieves this through the iterative minimization of a prescribed cost (or penalty) function. Differences between the analysis and observations/first guess are penalized (damped) according to their perceived error. The difference between three-dimensional (3D-Var) and four-dimensional (4D-Var) data assimilation is the use of a numerical forecast model in the latter.
The MMM Division of NCAR supports a unified (global/regional, multi-model, 3/4D-Var) model-space data assimilation system (WRFDA) for use by NCAR staff and collaborators, and is also freely available to the general community, together with further documentation, test results, plans etc., from the WRFDA web-page http://www2.mmm.ucar.edu/wrf/users/wrfda/Docs/user_guide_V3.3/users_guide_chap6.htm.
Various components of the WRFDA system are shown in blue in the sketch below, together with their relationship with rest of the WRF system.
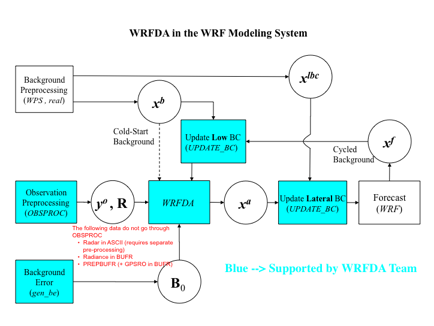
xb: first guess either from a previous WRF forecast or from WPS/REAL output.
xlbc: lateral boundary from WPS/REAL output.
xa: analysis from WRFDA data assimilation system.
xf: WRF forecast output.
yo: observations processed by OBSPROC. (note: PREPBUFR input, Radar and Radiance data donÕt go through OBSPROC)
B0: background error statistics from generic BE data (CV3) or gen_be.
R: observational and representative error statistics.
In this chapter, you will learn how to run the various components of the WRFDA system. For training purposes, you are supplied with a test case including the following input data: a) observation file (in the format prior to OBSPROC), b) WRF netCDF background file (WPS/REAL output used as a first guess of the analysis), and c) Background error statistics (estimate of errors in the background file). You can download the test dataset from http://www2.mmm.ucar.edu/wrf/users/wrfda/download/testdata.html. In your own work, you have to create all these input files yourselves. See the section Running Observation Preprocessor for creating your observation files. See section Running gen_be for generating your background error statistics file if you want to use cv_options=5 or cv_options=6.
Before using your own data, we suggest that you start by running through the WRFDA related programs at least once using the supplied test case. This serves two purposes: First, you can learn how to run the programs with data we have tested ourselves, and second you can test whether your computer is adequate to run the entire modeling system. After you have done the tutorial, you can try running other, more computationally intensive, case studies and experimenting with some of the many namelist variables.
WARNING: It is impossible to test every code upgrade with every permutation of computer, compiler, number of processors, case, namelist option, etc. The ÒnamelistÓ options that are supported are indicated in the ÒWRFDA/var/README.namelistÓ, and these are the default options.
Running with your own domain, hopefully, our test cases will have prepared you for the variety of ways in which you may wish to run WRFDA. Please inform us about your experiences.
As a professional courtesy, we request that you include the following reference in any publications that uses of any component of the community WRFDA system:
Barker, D.M., W. Huang, Y.R. Guo, and Q.N. Xiao., 2004: A
Three-Dimensional (3DVAR) Data Assimilation System For Use With MM5:
Implementation and Initial Results. Mon.
Wea. Rev., 132, 897-914.
Huang, X.Y., Q. Xiao, D.M. Barker, X. Zhang, J. Michalakes,
W. Huang, T. Henderson, J. Bray, Y. Chen, Z. Ma, J. Dudhia, Y. Guo, X. Zhang,
D.J. Won, H.C. Lin, and Y.H. Kuo, 2009: Four-Dimensional Variational Data
Assimilation for WRF: Formulation and Preliminary Results. Mon. Wea. Rev., 137,
299–314.
Running WRFDA requires a Fortran 90 compiler. We have currently tested the WRFDA on the following platforms: IBM (XLF), SGI Altix (INTEL), PC/Linux (PGI, INTEL, GFORTRAN), and Apple (G95/PGI). Please let us know if this does not meet your requirements, and we will attempt to add other machines to our list of supported architectures as resources allow. Although we are interested to hear of your experiences on modifying compile options, we do not yet recommend making changes to the configure file used to compile WRFDA.
Installing WRFDA for 3D-Var Run
a. Obtaining
WRFDA Source Code
Users can download the WRFDA source code
from http://www2.mmm.ucar.edu/wrf/users/wrfda/download/get_source.html.
After the tar file is unzipped (gunzip WRFDAV3.3.TAR.gz) and untarred (untar WRFDAV3.3.TAR), the directory WRFDA should be created; this directory contains the WRFDA source, external libraries, and fixed files. The following is a list of the system components and the content for each directory:
|
Directory Name |
Content |
|
var/da |
WRFDA source code |
|
var/run |
Fixed input files required by WRFDA, such as background error covariance, and radiance related files CRTM coefficients, radiance_info and VARBC.in. |
|
var/external |
Library needed by WRFDA, include crtm, bufr, lapack, blas |
|
var/obsproc |
Obsproc source code , namelist, and observation error file. |
|
var/gen_be |
Source code of generate background error |
|
var/build |
Build all .exe files. |
b. Compile
WRFDA and Libraries
Starting from V3.1.1, some external
libraries (for example, lapack, blas, and NCEP BUFR) are included in the WRFDA
tar file. To compile the WRFDA code, it is necessary to have installed the
netCDF library, which is the only mandatory library if only conventional
observational data from LITTLE_R format file is to be used.
> setenv NETCDF your_netcdf_path
If observational data in PREPBUFR format are to be used, an environmental variable needs to be set like (using the C-shell),
>
setenv BUFR 1
To have NCEP BUFR library compiled and have BUFR-related WRFDA code generated and compiled after configure/compile.
If satellite radiance data are to be used, in addition to NCEP BUFR library, RTM (Radiative Transfer Model) is required. The current RTM versions that WRFDA uses are CRTM V2.0.2 and RTTOV V10. WRFDA can compile with CRTM only, or RTTOV only, or both CRTM and RTTOV together.
Starting from V3.2.1, CRTM V2.0.2 is included in the WRFDA tar file.
> setenv CRTM
1
To have CRTM library compiled and CRTM-related WRFDA code generated and compiled after configure/compile.
If RTTOV v10 is the RTM to be used for radiance data assimilation, for the user should have downloaded and installed the RTTOV library before compiling WRFDA.
RTTOV v10 can be
downloaded from http://research.metoffice.gov.uk/research/interproj/nwpsaf/rtm
After following the RTTOV documentation to compile the RTTOV library, set the RTTOV environment variable to the path where lib/librttov10.1.0_*.a reside.
> setenv RTTOV /usr/local/rttov10/pgi (in this example, there should exist /usr/local/rttov10/pgi/lib/librttov10.1.0_*.a)
Note: Make sure the required libraries were all compiled using the same compiler that will be used to build WRFDA, since the libraries produced by one compiler may not be compatible with code compiled with another.
Assuming all required libraries are available and the WRFDA source code is ready, start to build the WRFDA using the following steps:
To configure WRFDA, enter the WRFDA directory and type
>
./configure wrfda
A list of configuration options for your computer should appear. Each option combines a compiler type and a parallelism option; since the configuration script doesnÕt check which compilers are actually available, be sure to select only among the options for compilers that are available on your system. The parallelism option allows for a single-processor (serial) compilation, shared-memory parallel (smpar) compilation, distributed-memory parallel (dmpar) compilation and distributed-memory with shared-memory parallel (sm+dm) compilation. For example, on a Macintosh computer, the above steps look like:
> ./configure wrfda
checking
for perl5... no
checking
for perl... found /usr/bin/perl (perl)
Will use
NETCDF in dir: /users/noname/work/external/g95/netcdf-3.6.1
PHDF5 not
set in environment. Will configure WRF for use without.
$JASPERLIB
or $JASPERINC not found in environment, configuring to build without grib2
I/O...
------------------------------------------------------------------------
Please
select from among the following supported platforms.
1. Darwin (MACOS) PGI compiler with pgcc (serial)
2. Darwin (MACOS) PGI compiler with pgcc (smpar)
3. Darwin (MACOS) PGI compiler with pgcc (dmpar)
4. Darwin (MACOS) PGI compiler with pgcc (dm+sm)
5. Darwin (MACOS) intel compiler with icc (serial)
6. Darwin (MACOS) intel compiler with icc (smpar)
7. Darwin (MACOS) intel compiler with icc (dmpar)
8. Darwin (MACOS) intel compiler with icc (dm+sm)
9. Darwin (MACOS) intel compiler with cc (serial)
10. Darwin (MACOS) intel compiler with cc (smpar)
11. Darwin (MACOS) intel compiler with cc (dmpar)
12. Darwin (MACOS) intel compiler with cc (dm+sm)
13. Darwin (MACOS) g95 with gcc (serial)
14. Darwin (MACOS) g95 with gcc (dmpar)
15. Darwin (MACOS) xlf (serial)
16. Darwin (MACOS) xlf (dmpar)
Enter
selection [1-10] : 13
------------------------------------------------------------------------
Compile
for nesting? (0=no nesting, 1=basic, 2=preset moves, 3=vortex following)
[default 0]:
Configuration
successful. To build the model type compile .
ÉÉ
After running the configuration script and choosing a compilation option, a configure.wrf file will be created. Because of the variety of ways that a computer can be configured, if the WRFDA build ultimately fails, there is a chance that minor modifications to the configure.wrf file may be needed.
Note: WRF compiles with –r4 option while WRFDA compiles with –r8. For this reason, WRF and WRFDA cannot reside and be compiled under the same directory.
Hint: It is helpful to start with something simple, such as the serial build. If it is successful, move on to build dmpar code. Remember to type Ôclean –aÕ between each build.
To compile the code, type
>
./compile all_wrfvar >&! compile.out
Successful compilation of Ôall_wrfvarÓ will produce 42 executables in the var/build directory which are linked in var/da directory, as well as obsproc.exe in var/obsproc/src directory. You can list these executables by issuing the command (from WRFDA directory)
> ls
-l var/build/*exe var/obsproc/src/obsproc.exe
-rwxr-xr-x 1 users 435816 Mar
9 19:26 var/build/da_advance_time.exe
-rwxr-xr-x 1 users 1195264 Mar 9
19:27 var/build/da_bias_airmass.exe
-rwxr-xr-x 1 users 815088 Mar
9 19:26 var/build/da_bias_scan.exe
-rwxr-xr-x 1 users 780476 Mar
9 19:26 var/build/da_bias_sele.exe
-rwxr-xr-x 1 users 1120408 Mar 9
19:26 var/build/da_bias_verif.exe
-rwxr-xr-x 1 users 1627284 Mar 9
19:26 var/build/da_rad_diags.exe
-rwxr-xr-x 1 users 639940 Mar
9 19:26 var/build/da_tune_obs_desroziers.exe
-rwxr-xr-x 1 users 608912 Mar
9 19:27 var/build/da_tune_obs_hollingsworth1.exe
-rwxr-xr-x 1 users 377748 Mar
9 19:27 var/build/da_tune_obs_hollingsworth2.exe
-rwxr-xr-x 1 users 1600636 Mar 9
19:27 var/build/da_update_bc.exe
-rwxr-xr-x 1 users 1662172 Mar 9
19:27 var/build/da_verif_grid.exe
-rwxr-xr-x 1 users 535916 Mar
9 19:32 var/build/da_verif_obs.exe
-rwxr-xr-x 1 users 29399039 Mar 9 19:32 var/build/da_wrfvar.exe
-rwxr-xr-x 1 users 2014440 Mar 9
19:32 var/build/gen_be_cov2d.exe
-rwxr-xr-x 1 users 2027684 Mar 9
19:27 var/build/gen_be_cov2d3d_contrib.exe
-rwxr-xr-x 1 users 2017952 Mar 9
19:27 var/build/gen_be_cov3d.exe
-rwxr-xr-x 1 users 2027804 Mar 9
19:27 var/build/gen_be_cov3d2d_contrib.exe
-rwxr-xr-x 1 users 2023396 Mar 9
19:27 var/build/gen_be_cov3d3d_bin3d_contrib.exe
-rwxr-xr-x 1 users 2027468 Mar 9
19:27 var/build/gen_be_cov3d3d_contrib.exe
-rwxr-xr-x 1 users 2003888 Mar 9
19:32 var/build/gen_be_diags.exe
-rwxr-xr-x 1 users 2028372 Mar 9
19:32 var/build/gen_be_diags_read.exe
-rwxr-xr-x 1 users 2012816 Mar 9
19:27 var/build/gen_be_ensmean.exe
-rwxr-xr-x 1 users 2045908 Mar 9 19:27
var/build/gen_be_ensrf.exe
-rwxr-xr-x 1 users 2069376 Mar 9
19:32 var/build/gen_be_ep1.exe
-rwxr-xr-x 1 users 2059240 Mar 9
19:32 var/build/gen_be_ep2.exe
-rwxr-xr-x 1 users 2022588 Mar 9
19:32 var/build/gen_be_etkf.exe
-rwxr-xr-x 1 users 2027480 Mar 9
19:27 var/build/gen_be_hist.exe
-rwxr-xr-x 1 users 2093900 Mar 9
19:32 var/build/gen_be_stage0_gsi.exe
-rwxr-xr-x 1 users 2105344 Mar 9
19:32 var/build/gen_be_stage0_wrf.exe
-rwxr-xr-x 1 users 2036928 Mar 9
19:32 var/build/gen_be_stage1.exe
-rwxr-xr-x 1 users 2064784 Mar 9
19:32 var/build/gen_be_stage1_1dvar.exe
-rwxr-xr-x 1 users 2036036 Mar 9
19:32 var/build/gen_be_stage1_gsi.exe
-rwxr-xr-x 1 users 2036024 Mar 9
19:32 var/build/gen_be_stage2.exe
-rwxr-xr-x 1 users 2100760 Mar 9
19:32 var/build/gen_be_stage2_1dvar.exe
-rwxr-xr-x 1 users 566584 Mar
9 19:26 var/build/gen_be_stage2_gsi.exe
-rwxr-xr-x 1 users 2023600 Mar 9
19:32 var/build/gen_be_stage2a.exe
-rwxr-xr-x 1 users 2036060 Mar 9
19:32 var/build/gen_be_stage3.exe
-rwxr-xr-x 1 users 2013852 Mar 9
19:32 var/build/gen_be_stage4_global.exe
-rwxr-xr-x 1 users 2049676 Mar 9
19:27 var/build/gen_be_stage4_regional.exe
-rwxr-xr-x 1 users 2003608 Mar 9
19:32 var/build/gen_be_vertloc.exe
-rwxr-xr-x 1 users 2155760 Mar 9
19:32 var/build/gen_mbe_stage2.exe
-rwxr-xr-x 1 users 1752352 Mar 23 09:29 var/obsproc/src/obsproc.exe
da_wrfvar.exe is the main executable for running WRFDA. Make sure it is created after the compilation. Sometimes (unfortunately) it is possible that other utilities get successfully compiled, while the main da_wrfvar.exe fails; please check the compilation log file carefully to figure out the problem.
The basic gen_be utility for regional model consists of gen_be_stage0_wrf.exe, gen_be_stage1.exe, gen_be_stage2.exe, gen_be_stage2a.exe, gen_be_stage3.exe, gen_be_stage4_regional.exe, and gen_be_diags.exe.
da_updated_bc.exe is used for updating WRF low and lateral boundary condition before and after a new WRFDA analysis is generated.
da_advance_time.exe is a very handy and useful tool for date/time manipulation. Type Òda_advance_time.exeÓ to see its usage instruction.
In addition to the executables
for running WRFDA and gen_be, obsproc.exe
(the executable for preparing conventional data for WRFDA) compilation is also
included in Ò./compile
all_wrfvarÓ. da_advance_time.exe
Go to WRFDA/var/external/bufr and WRFDA/var/external/crtm to check if the libbufr.a and libcrtm.a were generated if you use BUFR and CRTM library.
c.
Clean
Compilation
To remove all object files and executables, type:
clean
To remove all build files, including configure.wrfda, type:
clean -a
The 'clean –a' command is recommended if compilation fails or configuration file is changed.
Installing WRFPLUS and WRFDA for 4D-Var Run
If you intend to run WRF 4D-Var, it is necessary to have WRFPLUS installed. From V3.3, we release a new version of WRFDA and WRFPLUS for 4D-Var. WRFPLUS contains the adjoint and tangent linear models based on a simplified WRF model, which only include dry dynamic processes. We are developing the tangent linear and adjoint codes of several simplified physical packages.
To install WRFPLUS V3.3:
- Get the WRFPLUS zipped tar file from:
http://www2.mmm.ucar.edu/wrf/users/wrfda/download/wrfplus.html
- Unzip and untar the file to WRFPLUS
> gzip -cd
WRFPLUS3.3.TAR.gz | tar -xf -
> cd WRFPLUS
> ./configure
wrfplus
serial means single processor
dmpar wrfplus.exe means Distributed Memory Parallel (MPI)
Note: For Version 3.3 WRFDA 4D-Var, parallel run is still under development, please compile WRFPLUS3.3 with serial mode.
- Compile WRFPLUS
> ./compile em_real
> ls -ls
main/*.exe
You
should see wrf.exe
To install WRFDA for 4D-Var run,
- Before you install WRFDA to run 4D-Var, environment variable should to be set with,
>setenv WRFPLUS_DIR ${your_source_code_dir}/WRFPLUS
- If you intend to use observational data with PREPBUFR format or if you intend to assimilate satellite radiance data, you need set environment variables for BUFR, CRTM, or RTTOV. This procedure is the same as installing WRFDA for 3D-Var run.
>./configure 4dvar
Note: Please compile WRFDA for 4D-Var run with serial mode.
>./compile
all_wrfvar
>ls -ls
var/build/*.exe
You should see da_wrfvar.exe.
Running Observation Preprocessor (OBSPROC)
The OBSPROC program reads observations in LITTLE_R format (a legendary ASCII format, in use since the MM5 era). The LITTLE_R format is also used in OBSGRID program. Please refer to the documentation at http://www2.mmm.ucar.edu/mm5/mm5v3/data/how_to_get_rawdata.html and Chapter 7 of this UserÕs Guide for LITTLE_R format description. For your applications, you will have to prepare your own observation files. Please see http://www2.mmm.ucar.edu/mm5/mm5v3/data/free_data.html for the sources of some freely available observations and the program for converting the observations to LITTLE_R format. Because the raw observation data files could be in any of formats, such as ASCII, BUFR, PREPBUFR, MADIS, HDF, etc. Furthermore, for each of formats, there may be the different versions. To make WRFDA system as general as possible, the LITTLE_R format ASCII file was adopted as an intermediate observation data format for WRFDA system. Some extensions were made in the LITTLE_R format for WRFDA applications. More complete description of LITTLE_R format and conventional observation data sources for WRFDA could be found from the web page: 2010 Summer Tutorial by clicking ÒObservation Pre-processingÓ. The conversion of the user-specific-source data to the LITTLE_R format observation data file is the usersÕ task.
The purposes of OBSPROC are to:
á Remove observations outside the time range and domain (horizontal and top).
á Re-order and merge duplicate (in time and location) data reports.
á Retrieve pressure or height based on observed information using the hydrostatic assumption.
á Check vertical consistency and super adiabatic for multi-level observations.
á Assign observational errors based on a pre-specified error file.
á Write out the observation file to be used by WRFDA in ASCII or BUFR format.
The OBSPROC program—obsproc.exe should be found under the directory WRFDA/var/obsproc/src if Òcompile all_wrfvarÓ was completed successfully.
a. Prepare observational data for 3D-Var
To prepare the observation file, for example, at the analysis time 0h for 3D-Var, all the observations between ±1h (or ±1.5h) will be processed, as illustrated in the following figure, which means that the observations between 23h and 1h are treated as the observations at 0h.
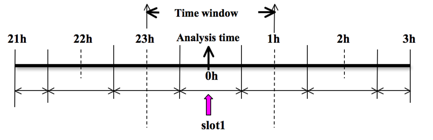
Before running obsproc.exe, create the required namelist file namelist.obsproc (see WRFDA/var/obsproc/README.namelist, or the section Description of Namelist Variables for details.
For your reference, an example file named Ònamelist_obsproc.3dvar.wrfvar-tutÓ has already been created in the var/obsproc directory. Thus, proceed as follows.
> cp
namelist.obsproc.3dvar.wrfvar-tut namelist.obsproc
Next, edit the namelist file namelist.obsproc by changing the following variables to accommodate your experiments.
&record1
obs_gts_filename='obs.2008020512'
&record2
time_window_min
= '2008-02-05_11:00:00',: The earliest time edge as ccyy-mm-dd_hh:mn:ss
time_analysis = '2008-02-05_12:00:00', : The
analysis time as ccyy-mm-dd_hh:mn:ss
time_window_max
= '2008-02-05_13:00:00',: The latest time edge as ccyy-mm-dd_hh:mn:ss
&record6,7,8
Edit all the domain settings to conform to your own experiment. You may pay special attention to NESTIX and NESTJX, which are described in the section Description of Namelist Variables for details.
&record9
use_for =
'3DVAR', ; used for 3D-Var,
default
To run OBSPROC, type
> obsproc.exe >&! obsproc.out
Once obsproc.exe has completed successfully, you will see an observation data file, obs_gts_2008-02-05_12:00:00.3DVAR, in the obsproc directory. This is the input observation file to WRFDA.
obs_gts_2008-02-05_12:00:00.3DVAR is an ASCII file that contains a header section (listed below) followed by observations. The meanings and format of observations in the file are described in the last six lines of the header section.
TOTAL
= 9066, MISS. =-888888.,
SYNOP
= 757, METAR = 2416, SHIP = 145, BUOY
= 250, BOGUS
= 0,
TEMP = 86,
AMDAR
= 19, AIREP
= 205, TAMDAR= 0, PILOT = 85, SATEM = 106, SATOB = 2556,
GPSPW
= 187, GPSZD = 0, GPSRF = 3, GPSEP = 0, SSMT1 = 0, SSMT2 = 0,
TOVS = 0, QSCAT = 2190, PROFL = 61,
AIRSR = 0,
OTHER = 0,
PHIC =
40.00, XLONC = -95.00, TRUE1 =
30.00, TRUE2 = 60.00, XIM11
= 1.00, XJM11 = 1.00,
base_temp=
290.00, base_lapse= 50.00,
PTOP = 1000., base_pres=100000., base_tropo_pres= 20000.,
base_strat_temp= 215.,
IXC = 60, JXC = 90, IPROJ = 1, IDD = 1, MAXNES= 1,
NESTIX= 60,
NESTJX= 90,
NUMC = 1,
DIS = 60.00,
NESTI
= 1,
NESTJ
= 1,
INFO = PLATFORM, DATE, NAME, LEVELS,
LATITUDE, LONGITUDE, ELEVATION, ID.
SRFC = SLP, PW (DATA,QC,ERROR).
EACH = PRES, SPEED, DIR, HEIGHT, TEMP, DEW
PT, HUMID (DATA,QC,ERROR)*LEVELS.
INFO_FMT
= (A12,1X,A19,1X,A40,1X,I6,3(F12.3,11X),6X,A40)
SRFC_FMT
= (F12.3,I4,F7.2,F12.3,I4,F7.3)
EACH_FMT
= (3(F12.3,I4,F7.2),11X,3(F12.3,I4,F7.2),11X,3(F12.3,I4,F7.2))
#------------------------------------------------------------------------------#
ÉÉ
observations ÉÉÉ
Before running WRFDA, you may find it useful to learn more about various types of data that will be processed to WRFDA, e.g., their geographical distribution. This file is in ASCII format and so you can easily view it. For a graphical view about file's content, use the ÒMAP_plotÓ utility to see the data distribution for each type of observations. To use this utility, proceed as follows.
> cd MAP_plot
> make
We
have prepared some configure.user.ibm/linux/mac/É
files for some platforms, when ÒmakeÓ
is typed, the Makefile
will use one of them to determine the compiler and compiler option. Please
modify the Makefile
and configure.user.xxx
to accommodate the complier on your platform. Successful compilation will
produce Map.exe. Note: The successful compilation of Map.exe requires
pre-installed NCARG Graphics libraries under $(NCARG_ROOT)/lib.
Modify the script Map.csh to set the time window and full path of input observation file (obs_gts_2008-02-05_12:00:00.3DVAR). You will need to set the following strings in this script as follows:
Map_plot
= /users/noname/WRFDA/var/obsproc/MAP_plot
TIME_WINDOW_MIN
= Ô2008020511Õ
TIME_ANALYSIS = Ô2008020512Õ
TIME_WINDOW_MAX
= Ô2008020513Õ
OBSDATA = ../obs_gts_2008-02-05_12:00:00.3DVAR
Next, type
>
Map.csh
When the job has completed, you
will have a gmeta file gmeta.{analysis_time} corresponding to analysis_time=2008020512.
This contains plots of data distribution for each type of observations
contained in the OBS data file: obs_gts_2008-02-05_12:00:00.3DVAR.
To view this, type
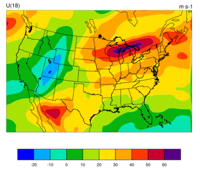
> idt
gmeta.2008020512
It will display (panel by panel) geographical distribution of various types of data. The following graphic shows the geographic distribution of ÒsondeÓ observations for this case.
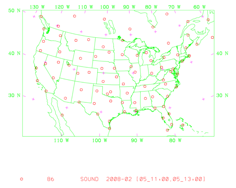
An alternative way to plot the observation is to use ncl script: WRFDA/var/graphics/ncl/plot_ob_ascii_loc.ncl. However, with this method, you need to provide the first guess file to the ncl script, and have ncl installed in your system.
b. Prepare observational data for 4D-Var
To prepare the observation file, for example, at the analysis time 0h for 4D-Var, all observations from 0h to 6h will be processed and grouped in 7 sub-windows from slot1 to slot7, as illustrated in following figure. NOTE: The ÒAnalysis timeÓ in the figure below is not the actual analysis time (0h), it indicates the time_analysis setting in the namelist file and is set to three hours later than the actual analysis time. The actual analysis time is still 0h.
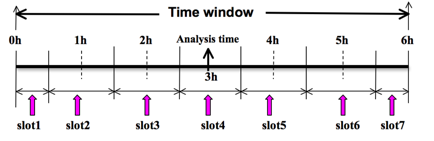
An example file named Ònamelist_obsproc.4dvar.wrfvar-tutÓ has already been created in the var/obsproc directory. Thus, proceed as follows:
> cp
namelist.obsproc.4dvar.wrfvar-tut namelist.obsproc
In the namelist file, you need to change the following variables to accommodate your experiments. In this test case, the actual analysis time is 2008-02-05_12:00:00, but in namelist, the time_analysis should be set to 3 hours later. The different value of time_analysis will make the different number of time slots before and after time_analysis. For example, if you set time_analysis = 2008-02-05_16:00:00, and set the num_slots_past = 4 and time_slots_ahead=2. The final results will be the same as before.
&record1
obs_gts_filename='obs.2008020512'
&record2
time_window_min
= '2008-02-05_12:00:00',: The earliest time edge as ccyy-mm-dd_hh:mn:ss
time_analysis = '2008-02-05_15:00:00', : The
analysis time as ccyy-mm-dd_hh:mn:ss
time_window_max
= '2008-02-05_18:00:00',: The latest time edge as ccyy-mm-dd_hh:mn:ss
&record6,7,8
Edit all the domain settings according to your own experiment. You may pay special attention to NESTIX and NESTJX, which is described in the section Description of Namelist Variables for details.
&record9
use_for =
'4DVAR', ; used for 3D-Var,
default
; num_slots_past and
num_slots_ahead are used ONLY for FGAT and 4DVAR:
num_slots_past = 3, ; the number of time slots
before time_analysis
num_slots_ahead = 3, ; the number of time slots after
time_analysis
To run OBSPROC, type
> obsproc.exe >&! obsproc.out
Once obsproc.exe has completed successfully, you
will see 7 observation data files:
obs_gts_2008-02-05_12:00:00.4DVAR
obs_gts_2008-02-05_13:00:00.4DVAR
obs_gts_2008-02-05_14:00:00.4DVAR
obs_gts_2008-02-05_15:00:00.4DVAR
obs_gts_2008-02-05_16:00:00.4DVAR
obs_gts_2008-02-05_17:00:00.4DVAR
obs_gts_2008-02-05_18:00:00.4DVAR
They are the input observation files to WRF 4D-Var. You can also use ÒMAP_PlotÓ to view the geographic distribution of different observations at different time slots.
Running WRFDA
a. Download Test Data
The WRFDA system requires three input files to run:
a) A WRF first guess and boundary input files output from either WPS/real (cold-start)
or WRF forecast (warm-start)
b) Observations (in ASCII format, PREPBUFR or BUFR for radiance)
c) A background error statistics file (containing background error covariance)
The following table summarizes the above info:
|
Input Data |
Format |
Created By |
|
First Guess |
NETCDF |
WRF Preprocessing System (WPS) and real.exe or WRF |
|
Observations |
ASCII (PREPBUFR also possible) |
Observation Preprocessor (OBSPROC) |
|
Background Error Statistics |
Binary |
/Default CV3 |
In the test case, you will store data in a directory defined by the environment variable $DAT_DIR. This directory can be at any location, and it should have read access. Type
> setenv DAT_DIR your_choice_of_dat_dir
Here, "your_choice_of_dat_dir" is the directory where the WRFDA input data is stored. If it does not exist, create this directory by typing
> cd $DAT_DIR
Download the test data for a ÒTutorialÓ case valid at 12 UTC 5th February 2008 from http://www2.mmm.ucar.edu/wrf/users/wrfda/download/testdata.html
Once you have downloaded ÒWRFDAV3.3-testdata.tar.gzÓ file to $DAT_DIR, extract it by typing
> gunzip WRFDAV3.3-testdata.tar.gz
> tar -xvf
WRFDAV3.3-testdata.tar
Now you should find the following
three sub-directories/files under Ò$DAT_DIRÓ
ob/2008020512/ob.2008020512.gz # Observation data in Òlittle_rÓ format
rc/2008020512/wrfinput_d01 # First guess file
rc/2008020512/wrfbdy_d01 # lateral boundary file
be/be.dat
# Background error file
......
You should first go through the
section ÒRunning Observation Preprocessor (OBSPROC)Ó and have a
WRF-3D-Var-ready observation file (obs_gts_2008-02-05_12:00:00.3DVAR)
generated in your OBSPROC working directory. You could then copy or move obs_gts_2008-02-05_12:00:00.3DVAR
to be in $DAT_DIR/ob/2008020512/ob.ascii.
If you want to try 4D-Var with
Little-R format observations, please go through the section
ÒRunning Observation Preprocessor (OBSPROC)Ó and have the WRF-4D-Var-ready
observation files (obs_gts_2008-02-05_12:00:00.4DVAR,ÉÉ).
You could copy or move the observation files to $DAT_DIR/ob using following commands:
> mv
obs_gts_2008-02-05_12:00:00.4DVAR
$DAT_DIR/ob/2008020512/ob.ascii+
> mv
obs_gts_2008-02-05_13:00:00.4DVAR
$DAT_DIR/ob/2008020513/ob.ascii
> mv
obs_gts_2008-02-05_14:00:00.4DVAR
$DAT_DIR/ob/2008020514/ob.ascii
> mv
obs_gts_2008-02-05_15:00:00.4DVAR
$DAT_DIR/ob/2008020515/ob.ascii
> mv
obs_gts_2008-02-05_16:00:00.4DVAR
$DAT_DIR/ob/2008020516/ob.ascii
> mv
obs_gts_2008-02-05_17:00:00.4DVAR
$DAT_DIR/ob/2008020517/ob.ascii
> mv
obs_gts_2008-02-05_18:00:00.4DVAR
$DAT_DIR/ob/2008020518/ob.ascii-
At this point you have three of
the input files (first guess, observation, and background error statistics
files in directory $DAT_DIR)
required to run WRFDA, and have successfully downloaded and compiled the WRFDA
code. If this is correct, you are ready to learn how to run WRFDA.
b. Run the Case—3D-Var
The data for this case is valid at 12 UTC 5th February 2008. The first guess comes from the NCEP FNL (Final) Operational Global Analysis data, passed through the WRF-WPS and real programs.
To run WRF 3D-Var, first create and cd to a working directory, for example, WRFDA/var/test/tutorial, and then follow the steps below:
> cd
WRFDA/var/test/tutorial
> ln
-sf WRFDA/run/LANDUSE.TBL ./LANDUSE.TBL
> ln
-sf $DAT_DIR/rc/2008020512/wrfinput_d01 ./fg (link first guess file as fg)
> ln
-sf WRFDA/var/obsproc/obs_gts_2008-02-05_12:00:00.3DVAR ./ob.ascii (link
OBSPROC processed observation file as ob.ascii)
> ln
-sf $DAT_DIR/be/be.dat ./be.dat (link background error statistics as be.dat)
> ln
-sf WRFDA/var/da/da_wrfvar.exe ./da_wrfvar.exe (link executable)
If you use PREPBUFR format data, please change the ob_format=1 in &wrfvar3 in namelist.input and link the data as ob.bufr,
> ln
-fs $DATA_DIR/ob/2008020512/gds1.t12.prepbufr.nr ob.bufr
We will begin by editing the file, namelist.input, which is a very basic namelist.input for running the tutorial test case as shown below and provided as WRFDA/var/test/tutorial/namelist.input. Only the time and domain settings need to be specified in this case, if we are using the default settings provided in WRFDA/Registry/Registry.wrfvar)
&wrfvar1
print_detail_grad=false,
/
&wrfvar2
/
&wrfvar3
/
&wrfvar4
/
&wrfvar5
/
&wrfvar6
/
&wrfvar7
/
&wrfvar8
/
&wrfvar9
/
&wrfvar10
/
&wrfvar11
calculate_cg_cost_fn=.false.
/
&wrfvar12
/
&wrfvar13
/
&wrfvar14
/
&wrfvar15
/
&wrfvar16
/
&wrfvar17
/
&wrfvar18
analysis_date="2008-02-05_12:00:00.0000",
/
&wrfvar19
/
&wrfvar20
/
&wrfvar21
time_window_min="2008-02-05_11:00:00.0000",
/
&wrfvar22
time_window_max="2008-02-05_13:00:00.0000",
/
&wrfvar23
/
&time_control
start_year=2008,
start_month=02,
start_day=05,
start_hour=12,
end_year=2008,
end_month=02,
end_day=05,
end_hour=12,
/
&dfi_control
/
&domains
e_we=90,
e_sn=60,
e_vert=41,
dx=60000,
dy=60000,
/
&physics
mp_physics=3,
ra_lw_physics=1,
ra_sw_physics=1,
radt=60,
sf_sfclay_physics=1,
sf_surface_physics=1,
bl_pbl_physics=1,
cu_physics=1,
cudt=5,
num_soil_layers=5, (IMPORTANT: itÕs essential to make sure
the setting here is consistent with the number in your first guess file)
mp_zero_out=2,
co2tf=0,
/
&fdda
/
&dynamics
/
&bdy_control
/
&grib2
/
&namelist_quilt
/
&perturbation
/
>
da_wrfvar.exe >&! wrfda.log
The file wrfda.log (or rsl.out.0000 if run in distributed-memory mode) contains important WRFDA runtime log information. Always check the log after a WRFDA run:
*** VARIATIONAL ANALYSIS ***
DYNAMICS OPTION: Eulerian Mass
Coordinate
alloc_space_field:
domain
1,
606309816 bytes allocat
ed
WRF TILE 1 IS 1 IE 89 JS 1 JE 59
WRF NUMBER OF TILES = 1
Set up
observations (ob)
Using
ASCII format observation input
scan obs ascii
end scan obs ascii
Observation
summary
ob time 1
sound
86 global,
86 local
synop
757 global,
750 local
pilot
85 global,
85 local
satem
106 global,
105 local
geoamv
2556 global,
2499 local
polaramv
0 global,
0 local
airep
224 global,
221 local
gpspw
187 global,
187 local
gpsrf
3 global, 3 local
metar
2416 global,
2408 local
ships
145 global,
140 local
ssmi_rv
0 global,
0 local
ssmi_tb
0 global,
0 local
ssmt1
0 global, 0 local
ssmt2
0 global, 0 local
qscat
2190 global,
2126 local
profiler
61 global,
61 local
buoy
247 global,
247 local
bogus
0 global, 0 local
pseudo
0 global, 0 local
radar
0 global, 0 local
radiance
0 global, 0 local
airs
retrieval 0 global, 0 local
sonde_sfc
86 global,
86 local
mtgirs
0 global, 0 local
tamdar
0 global, 0 local
Set up
background errors for regional application for cv_options = 5
Using the averaged regression
coefficients for unbalanced part
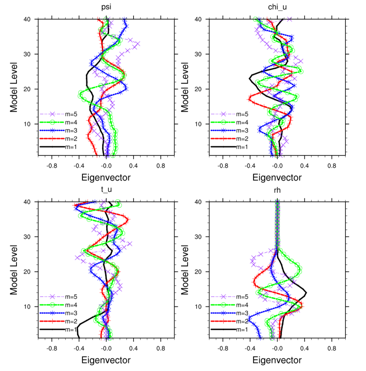
WRF-Var dry control variables
are:psi, chi_u, t_u and ps_u
Humidity control variable is rh
Vertical
truncation for psi = 15( 99.00%)
Vertical
truncation for chi_u = 20( 99.00%)
Vertical
truncation for t_u
= 29( 99.00%)
Vertical
truncation for rh
= 22( 99.00%)
Scaling: var, len, ds: 0.100000E+01 0.100000E+01 0.600000E+05
Scaling: var, len, ds: 0.100000E+01 0.100000E+01 0.600000E+05
Scaling: var, len, ds: 0.100000E+01 0.100000E+01 0.600000E+05
Scaling: var, len, ds: 0.100000E+01 0.100000E+01 0.600000E+05
Scaling: var, len, ds: 0.100000E+01 0.100000E+01 0.600000E+05
Calculate
innovation vector(iv)
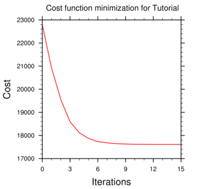
Minimize
cost function using CG method
Starting
outer iteration : 1
Starting
cost function: 2.53214888D+04,
Gradient= 2.90675545D+02
For this
outer iteration gradient target is: 2.90675545D+00
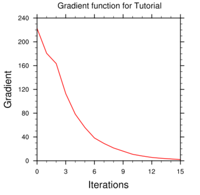
----------------------------------------------------------
Iter Cost Function
Gradient
Step
1 2.32498037D+04
2.55571188D+02
4.90384516D-02
2 2.14988144D+04
2.22354203D+02
5.36154186D-02
3 2.01389088D+04
1.62537907D+02 5.50108123D-02
4 1.93433827D+04
1.26984567D+02
6.02247687D-02
5 1.88877194D+04
9.84565874D+01
5.65160951D-02
6 1.86297777D+04
7.49071361D+01
5.32184146D-02
7 1.84886755D+04
5.41516421D+01 5.02941363D-02
8 1.84118462D+04
4.68329312D+01
5.24003071D-02
9 1.83485166D+04
3.53595537D+01
5.77476335D-02
10 1.83191278D+04
2.64947070D+01
4.70109040D-02
11 1.82984221D+04
2.06996271D+01 5.89930206D-02
12 1.82875693D+04
1.56426527D+01
5.06578447D-02
13 1.82807224D+04
1.15892153D+01
5.59631997D-02
14 1.82773339D+04
8.74778514D+00
5.04582959D-02
15 1.82751663D+04
7.22150257D+00 5.66521675D-02
16 1.82736284D+04
4.81374868D+00
5.89786400D-02
17 1.82728636D+04
3.82286871D+00
6.60104384D-02
18 1.82724306D+04
3.16737517D+00
5.92526480D-02
19 1.82721735D+04
2.23392283D+00 5.12604438D-02
----------------------------------------------------------
Inner
iteration stopped after 19
iterations
Final: 19 iter, J= 1.98187399D+04, g=
2.23392283D+00
----------------------------------------------------------
Diagnostics
Final cost function J = 19818.74
Total number of obs.
= 39800
Final value of J
=
19818.73988
Final value of Jo = 16859.85861
Final value of Jb = 2958.88127
Final value of Jc =
0.00000
Final value of Je =
0.00000
Final value of Jp =
0.00000
Final value of Jl =
0.00000
Final J / total num_obs =
0.49796
Jb factor used(1) =
1.00000 1.00000
1.00000 1.00000
1.00000
1.00000 1.00000
1.00000 1.00000
1.00000
Jb factor used(2) =
1.00000 1.00000 1.00000
1.00000 1.00000
1.00000 1.00000
1.00000 1.00000
1.00000
Jb factor used(3) =
1.00000 1.00000
1.00000
1.00000 1.00000
1.00000 1.00000 1.00000 1.00000
1.00000
Jb factor used(4) =
1.00000 1.00000
1.00000
1.00000 1.00000
1.00000 1.00000
1.00000 1.00000
1.00000
Jb factor used(5)
=
1.00000 1.00000
1.00000 1.00000
1.00000
1.00000 1.00000
1.00000 1.00000
1.00000
Jb factor used
=
1.00000
Je factor used
=
1.00000
VarBC factor used =
1.00000
*** WRF-Var completed successfully ***
A file called namelist.output (which contains the complete namelist settings) will be generated after a successful da_wrfvar.exe run. The settings appearing in namelist.output, but not specified in your namelist.input, are the default values from WRFDA/Registry/Registry.wrfvar.
After successful completion of job, wrfvar_output (the WRFDA analysis file, i.e. the new initial condition for WRF) should appear in the working directory along with a number of diagnostic files. Various text diagnostics output files will be explained in the next section (WRFDA Diagnostics).
To understand the role of various important WRFDA options, try re-running WRFDA by changing different namelist options, for example, making WRFDA convergence criteria more stringent. This is achieved by reducing the value of the convergence criteria ÒEPSÓ to e.g. 0.0001 by adding "EPS=0.0001" in the namelist.input record &wrfvar6. See section (WRFDA additional exercises) for more namelist options.
c. Run the Case—4D-Var
To run WRF 4D-Var, first create and enter into a working directory, for example, WRFDA/var/test/4dvar.
Note: If you want to setup your own directories to run 4D-Var, please make sure you follow the linkages and namelist.input under WRFDA/var/test/4dvar.
Assume the analysis date is 2008020512 and the test data directories are:
>
setenv DATA_DIR /ptmp/$user/DATA
> ls
–lr $DATA_DIR
ob/2008020512
ob/2008020513
ob/2008020514
ob/2008020515
ob/2008020516
ob/2008020517
ob/2008020518
rc/2008020512
be
Note: WRFDA 4D-Var is able to assimilate conventional observation data, satellites radiance BUFR data, radar data, and the input data format can be PREPBUFR format data or observation data processed by OBSPROC.
Assume the working directory is:
>
setenv WORK_DIR $WRFDA_DIR/var/test/4dvar
Then follow the steps below:
1) Link the executables.
> cd
$WORK_DIR
> ln
-fs $WRFDA_DIR/var/da/da_wrfvar.exe .
2) Link the observational data, first guess, BE and LANDUSE.TBL, etc..
> cd
$WORK_DIR
> ln
-fs $DATA_DIR/ob/2008020512/ob.ascii+ ob01.ascii
> ln
-fs $DATA_DIR/ob/2008020513/ob.ascii
ob02.ascii
> ln
-fs $DATA_DIR/ob/2008020514/ob.ascii
ob03.ascii
> ln
-fs $DATA_DIR/ob/2008020515/ob.ascii
ob04.ascii
> ln
-fs $DATA_DIR/ob/2008020516/ob.ascii
ob05.ascii
> ln
-fs $DATA_DIR/ob/2008020517/ob.ascii
ob06.ascii
> ln
-fs $DATA_DIR/ob/2008020518/ob.ascii- ob07.ascii
> ln
-fs $DATA_DIR/rc/2008020512/wrfinput_d01 .
> ln
-fs $DATA_DIR/rc/2008020512/wrfbdy_d01 .
> ln
-fs wrfinput_d01 fg
> ln
-fs $DATA_DIR/be/be.dat .
> ln
-fs WRFDA/run/LANDUSE.TBL ./LANDUSE.TBL
> ln
-fs WRFDA/run/GENPARM.TBL ./GENPARM.TBL
> ln
-fs WRFDA/run/SOILPARM.TBL ./SOILPARM.TBL
> ln
-fs WRFDA/run/VEGPARM.TBL ./VEGPARM.TBL
> ln
–fs WRFDA/run/RRTM_DATA_DBL RRTM_DATA
If you use PREPBUFR format data, please change the ob_format=1 in &wrfvar3 in namelist.input and link the data as ob.bufr,
> ln
-fs $DATA_DIR/ob/2008020512/gds1.t12.prepbufr.nr ob.bufr
If you would like to assimilate
PREPBUFR data at both12hr and 18hr for 4D-Var, you should linked it as follows,
> ln
-fs $DATA_DIR/ob/2008020512/gds1.t12.prepbufr.nr ob01.bufr
> ln
-fs $DATA_DIR/ob/2008020518/gds1.t18.prepbufr.nr ob02.bufr
Note: NCEP BUFR files downloaded from NCEPÕs public ftp server ftp://ftp.ncep.noaa.gov/pub/data/nccf/com/gfs/prod/gdas.${yyyymmddhh} are Fortran-blocked on big-endian machine and can be directly used on big-endian machines (for example, IBM). For most Linux clusters with Intel platforms, users need to download the byte-swapping code ssrc.c (http://www.dtcenter.org/com-GSI/users/support/faqs/index.php, the C code ssrc.c located in the /utils directory of the GSI distribution), and this code will convert a prepbufr file generated on an IBM platofrm to a prepbufr file that can be read on a Linux or Intel Mac platform. Compile ssrc.c with any c compiler (e.g., gcc -o ssrc.exe ssrc.c). To convert an IBM prepbufr file, take the executable (e.g. ssrc.exe), and run it as follows,
ssrc.exe <name of Big Endian prepbufr
file> name of Little Endian prepbufr file
3) Run in single processor mode (serial compilation required for WRF 4D-Var)
Edit $WORK_DIR/namelist.input
to match your experiment settings. The most important namelist variables
related to 4D-Var are listed below, please refer to README.namelist under WRFDA/var directory.
&wrfvar1
var4d=true,
var4d_lbc=true,
var4d_bin=3600,
ÉÉ
/
ÉÉ
&perturbation
trajectory_io=true,
enable_identity=false,
jcdfi_use=true,
jcdfi_diag=1,
jcdfi_penalty=1000.0,
/
> cd
$WORK_DIR
>
./da_wrfvar.exe >&! wrfda.log
Radiance Data Assimilations in WRFDA
This section gives a brief description for various aspects
related to radiance assimilation in WRFDA. Each aspect is described mainly from
the viewpoint of usage rather than more technical and scientific details, which
will appear in separated technical report and scientific paper. Namelist
parameters controlling different aspects of radiance assimilation will be
detailed in the following sections. It should be noted that this section does
not cover general aspects of the WRFDA assimilation. These can be found in
other sections of chapter 6 of this users guide or other WRFDA documentation.
a. Running WRFDA with radiances
In addition to the basic input files (LANDUSE.TBL, fg, ob.ascii, be.dat) mentioned in ÒRunning WRFDAÓ section, the following additional files are required for
radiances: radiance data in NCEP BUFR format, radiance_info files, VARBC.in, RTM (CRTM or RTTOV) coefficient files.
Edit namelist.input (Pay special attention to &wrfvar4, &wrfvar14, &wrfvar21, and
&wrfvar22 for radiance-related options. A
very basic namelist.input for running the radiance test case is provided as WRFDA/var/test/radiance/namelist.input)
> ln
-sf ${DAT_DIR}/gdas1.t00z.1bamua.tm00.bufr_d ./amsua.bufr
> ln
-sf ${DAT_DIR}/gdas1.t00z.1bamub.tm00.bufr_d ./amsub.bufr
> ln
-sf WRFDA/var/run/radiance_info
./radiance_info #
(radiance_info is a directory)
> ln
-sf WRFDA/var/run/VARBC.in
./VARBC.in
(CRTM
only) > ln -sf
WRFDA/var/run/crtm_coeffs ./crtm_coeffs #(crtm_coeffs is a directory)
(RTTOV
only) > ln -sf your_path/rtcoef_rttov10/rttov7pred51L ./rttov_coeffs # (rttov_coeffs is a directory)
See the following sections for more details on each aspect.
b. Radiance Data Ingest
Currently, the ingest interface for NCEP BUFR radiance data
is implemented in WRFDA. The radiance data are available through NCEPÕs public
ftp server ftp://ftp.ncep.noaa.gov/pub/data/nccf/com/gfs/prod/gdas.${yyyymmddhh}
in near real-time (with 6-hour delay) and can meet requirements both for
research purposes and some real-time applications.
So far, WRFDA can read data from the NOAA ATOVS instruments
(HIRS, AMSU-A, AMSU-B and MHS), the EOS Aqua instruments (AIRS, AMSU-A) and
DMSP instruments (SSMIS). Note that NCEP radiance BUFR files are separated by
instrument names (i.e., each file for one type instrument), and each file
contains global radiance (generally converted to brightness temperature) within
6-hour assimilation window from multi-platforms. For running WRFDA, users need
to rename NCEP corresponding BUFR files (table 1) to hirs3.bufr (including HIRS
data from NOAA-15/16/17), hirs4.bufr (including HIRS data from NOAA-18/19, METOP-2), amsua.bufr (including AMSU-A
data from NOAA-15/16/18/19, METOP-2), amsub.bufr (including AMSU-B data from NOAA-15/16/17), mhs.bufr (including MHS data
from NOAA-18/19 and METOP-2), airs.bufr (including AIRS and AMSU-A data from EOS-AQUA) and ssmis.bufr (SSMIS data from
DMSP-16, AFWA provided) for WRFDA filename convention. Note that airs.bufr file
contains not only AIRS data but also AMSU-A, which is collocated with AIRS
pixels (1 AMSU-A pixels collocated with 9 AIRS pixels). Users must place these
files in the working directory where WRFDA executable is run. It should also be
mentioned that WRFDA reads these BUFR radiance files directly without use if
any separate pre-processing program is used. All processing of radiance data,
such as quality control, thinning and bias correction and so on, is carried out
inside WRFDA. This is different from conventional observation assimilation,
which requires a pre-processing package (OBSPROC) to generate WRFDA readable
ASCII files. For reading the radiance BUFR files, WRFDA must be compiled with
the NCEP BUFR library (see http://www.nco.ncep.noaa.gov/sib/decoders/BUFRLIB/).
Table 1: NCEP and WRFDA radiance BUFR file naming convention
|
NCEP BUFR file names |
WRFDA naming convention |
|
gdas1.t00z.1bamua.tm00.bufr_d |
amsua.bufr |
|
gdas1.t00z.1bamub.tm00.bufr_d |
amsub.bufr |
|
gdas1.t00z.1bhrs3.tm00.bufr_d |
hirs3.bufr |
|
gdas1.t00z.1bhrs4.tm00.bufr_d |
hirs4.bufr |
|
gdas1.t00z.1bmhs.tm00.bufr_d |
mhs.bufr |
|
gdas1.t00z.airsev.tm00.bufr_d |
airs.bufr |
Namelist parameters are used to control the reading of
corresponding BUFR files into WRFDA. For instance, USE_AMSUAOBS, USE_AMSUBOBS, USE_HIRS3OBS, USE_HIRS4OBS, USE_MHSOBS, USE_AIRSOBS, USE_EOS_AMSUAOBS and USE_SSMISOBS control whether or not
the respective file is read. These are logical parameters that are assigned to
.FALSE. by default; therefore they must be set to .TRUE. to read the respective observation file. Also note that
these parameters only control whether the data is read, not whether the data
included in the files is to be assimilated. This is controlled by other
namelist parameters explained in the next section.
NCEP BUFR files downloaded from NCEPÕs public ftp server ftp://ftp.ncep.noaa.gov/pub/data/nccf/com/gfs/prod/gdas.${yyyymmddhh}
are Fortran-blocked on big-endian machine and can be directly used on
big-endian machines (for example, IBM). For most Linux clusters with Intel
platforms, users need to download the byte-swapping code ssrc.c (http://www.dtcenter.org/com-GSI/users/support/faqs/index.php, the C code ssrc.c
located in the /utils directory of
the GSI distribution), and this code will convert a prepbufr file generated on
an IBM platofrm to a prepbufr file
that can be read on a Linux or Intel Mac platform. Compile ssrc.c with any c
compiler (e.g., gcc -o ssrc.exe ssrc.c). To convert an IBM prepbufr file, take the executable
(e.g. ssrc.exe),
and run it as follows,
ssrc.exe <name of
Big Endian prepbufr file> name of Little Endian prepbufr file
c. Radiative Transfer Model
The core component for direct radiance assimilation is to
incorporate a radiative transfer model (RTM, should be accurate enough yet
fast) into the WRFDA system as one part of observation operators. Two widely
used RTMs in NWP community, RTTOV (developed by EUMETSAT in Europe), and CRTM
(developed by the Joint Center for Satellite Data Assimilation (JCSDA) in US),
are already implemented in WRFDA system with a flexible and consistent user
interface. Selection a which RTM to use is controlled by a simple namelist
parameter RTM_OPTION
(1 for RTTOV, the default, and 2 for CRTM). WRFDA is designed to be able to
compile with only one of two RTM libraries or without RTM libraries (for those
not interested in radiance assimilation) by the definition of environment
variables ÒCRTMÓ and ÒRTTOVÓ (see Installing WRFDA section).
Both RTMs can calculate radiances for almost all available
instruments aboard various satellite platforms in orbit. An important feature
of WRFDA design is that all data structures related to radiance assimilation
are dynamically allocated during running time according to simple namelist
setup. The instruments to be assimilated are controlled at run time by four
integer namelist parameters: RTMINIT_NSENSOR (the total number of sensors to be assimilated), RTMINIT_PLATFORM (the platforms IDs
array to be assimilated with dimension RTMINIT_NSENSOR, e.g., 1 for NOAA, 9 for
EOS, 10 for METOP and 2 for DMSP), RTMINIT_SATID (satellite IDs array) and RTMINIT_SENSOR (sensor IDs array,
e.g., 0 for HIRS, 3 for AMSU-A, 4 for AMSU-B, 15 for MHS, 10 for SSMIS, 11 for
AIRS). For instance, the configuration for assimilating 12 sensors from 7
satellites (what WRFDA can assimilated currently) will be
RTMINIT_NSENSOR
= 12 14 # 6 AMSUA; 3 AMSUB; 3 MHS; 1 AIRS; 1 SSMIS
RTMINIT_PLATFORM
= 1, 1, 1, 1,9,10,,1, 1, 1, 1, 1,10,
9, 2
RTMINIT_SATID
= 15,16,18,19,2,
2,15,16,17,18,19, 2, 2,16
RTMINIT_SENSOR
= 3, 3, 3, 3,3, 3, 4, 4,
4,15,15,15, 11,10
The instrument triplets (platform, satellite, and sensor ID) in the namelist can be ranked in any order. More detail about the convention of instrument triplet can be found on the web page http://research.metoffice.gov.uk/research/interproj/nwpsaf/rtm/rttov_description.html
or the tables 2 and 3 in the RTTOV v10
Users Guide (http://research.metoffice.gov.uk/research/interproj/nwpsaf/rtm/docs_rttov10/users_guide_10_v1.3.pdf
)
CRTM uses a different instrument naming method. A convert
routine inside WRFDA is already created to make CRTM use the same instrument
triplet as RTTOV such that the user interface remains the same for RTTOV and
CRTM.
When running WRFDA with radiance assimilation switched on
(RTTOV or CRTM), a set of RTM coefficient files need to be loaded. For RTTOV
option, RTTOV coefficient files are to be copied or linked to a sub-directory
Òrttov_coeffsÓ under the working directory; for CRTM option, CRTM coefficient
files are to be copied or linked to a sub-directory Òcrtm_coeffsÓ under the
working directory. Only coefficients listed in namelist are needed. Potentially
WRFDA can assimilate all sensors as long as the corresponding coefficient files
are provided with RTTOV and CRTM. In addition, necessary developments on
corresponding data interface, quality control, and bias correction are also important
to make radiance data assimilated properly. However, a modular design of radiance
relevant routines already facilitates much to add more instruments in WRFDA.
The RTTOV package is not distributed with WRFDA due to
licensing and supporting issues. Users need to follow the instructions
on http://research.metoffice.gov.uk/research/interproj/nwpsaf/rtm to download the
RTTOV source code and supplement coefficient files and emissivity atlas
dataset. Starting from WRFDA V3.3, only RTTOV v10 can be used in WRFDA.
Starting from V3.2.1, the CRTM package is distributed with
WRFDA, which is located in WRFDA/var/external/crtm. The CRTM code in WRFDA is
basically the same as the source code that users can download from the the
following link:
ftp://ftp.emc.ncep.noaa.gov/jcsda/CRTM.
d. Channel Selection
Channel selection in WRFDA is controlled by radiance ÔinfoÕ
files located in the sub-directory Ôradiance_infoÕ under the working directory.
These files are separated by satellites and sensors, e.g., noaa-15-amsua.info,
noaa-16-amsub.info, dmsp-16-ssmis.info and so on. An example for 5 channels
from noaa-15-amsub.info is shown below. The fourth column is used by WRFDA to
control when to use a corresponding channel. Channels with the value Ò-1Ó
indicate that the channel is Ònot assimilatedÓ (channels 1, 2 and 4 in this
case), with the value Ò1Ó means ÒassimilatedÓ (channels 3 and 5). The sixth
column is used by WRFDA to set the observation error for each channel. Other
columns are not used by WRFDA. It should be mentioned that these error values
might not necessarily be optimal for your applications; It is the userÕs
responsibility to obtain the optimal error statistics for your own
applications.
Sensor channel
IR/MW use idum varch polarisation(0:vertical;1:horizontal)
415 1 1
-1 0 0.5500000000E+01 0.0000000000E+00
415 2 1
-1 0 0.3750000000E+01 0.0000000000E+00
415 3 1 1 0
0.3500000000E+01
0.0000000000E+00
415 4 1
-1 0 0.3200000000E+01 0.0000000000E+00
415 5 1 1 0
0.2500000000E+01
0.0000000000E+00
e. Bias Correction
Satellite radiance is generally considered biased with
respect to a reference (e.g., background or analysis field in NWP assimilation)
due to system error of observation itself, reference field, and RTM. Bias
correction is a necessary step prior to assimilating radiance data. There are
two ways of performing bias correction in WRFDA. One is based on the Harris and
Kelly (2001) method and is carried out using a set of coefficient files
pre-calculated with an off-line statistics package, which will apply to a
training dataset for a month-long period. The other is Variational Bias
Correction (VarBC). Only VarBC is
introduced here and recommended for users because of its relative simplicity in
usage.
f. Variational Bias Correction
Getting started with VarBC
To use VarBC, set namelist option USE_VARBC to TRUE and have a
VARBC.in file in the working directory. VARBC.in is a VarBC setup file in ASCII
format. A template is provided with the WRFDA package (WRFDA/var/run/VARBC.in).
Input and Output files
All VarBC input is passed through one single ASCII file
called VARBC.in file. Once WRFDA has run with the VarBC option switched on, it
will produce a VARBC.out file which looks very much like the VARBC.in file you
provided. This output file will then be used as the input file for the next
assimilation cycle.
Coldstart
Coldstarting means starting the VarBC from scratch i.e.
when you do not know the values of the bias parameters.
The Coldstart is a routine in WRFDA. The bias predictor
statistics (mean and standard deviation) are computed automatically and will be
used to normalize the bias parameters. All coldstarted bias parameters are set
to zero, except the first bias parameter (= simple offset), which is set to the
mode (=peak) of the distribution of the (uncorrected) innovations for the given
channel.
A threshold of number of observations can be set through a
namelist option VARBC_NOBSMIN (default = 10), under which it is considered that not
enough observations are present to keep the Coldstart values (i.e. bias
predictor statistics and bias parameter values) for the next cycle. In this
case, the next cycle will do another Coldstart.
Background Constraint for the bias
parameters
The background constraint controls the inertia you want to
impose on the predictors (i.e. the smoothing in the predictor time series). It
corresponds to an extra term in the WRFDA cost function.
It is defined through an integer number in the VARBC.in
file. This number is related to a number of observations: the bigger the
number, the more inertia constraint. If these numbers are set to zero, the
predictors can evolve without any constraint.
Scaling factor
The VarBC uses a specific preconditioning, which can be
scaled through a namelist option VARBC_FACTOR (default = 1.0).
Offline bias correction
The analysis of the VarBC parameters can be performed
"offline", i.e. independently from the main WRFDA analysis. No extra
code is needed, just set the following MAX_VERT_VAR* namelist variables to be 0, which will disable the
standard control variable and only keep the VarBC control variable.
MAX_VERT_VAR1=0.0
MAX_VERT_VAR2=0.0
MAX_VERT_VAR3=0.0
MAX_VERT_VAR4=0.0
MAX_VERT_VAR5=0.0
Freeze VarBC
In certain circumstances, you might want to keep the VarBC
bias parameters constant in time (="frozen"). In this case, the bias
correction is read and applied to the innovations, but it is not updated during
the minimization. This can easily be achieved by setting the namelist options:
USE_VARBC=false
FREEZE_VARBC=true
Passive observations
Some observations are useful for preprocessing (e.g. Quality
Control, Cloud detection) but you might not want to assimilate them. If you
still need to estimate their bias correction, these observations need to go
through the VarBC code in the minimization. For this purpose, the VarBC uses a
separate threshold on the QC values, called "qc_varbc_bad". This
threshold is currently set to the same value as "qc_bad", but can
easily be changed to any ad hoc value.
g. Other namelist variables to control
radiance assimilation
RAD_MONITORING (30)
Integer array of dimension RTMINIT_NSENSER, where 0 for
assimilating mode, 1 for monitoring mode (only calculate innovation).
THINNING
Logical, TRUE will perform thinning on radiance data.
THINNING_MESH (30)
Real array with dimension RTMINIT_NSENSOR, values indicate
thinning mesh (in KM) for different sensors.
QC_RAD
Logical, control if perform quality control, always set to
TRUE.
WRITE_IV_RAD_ASCII
Logical, control if output Observation minus Background
files which are in ASCII format and separated by sensors and processors.
WRITE_OA_RAD_ASCII
Logical, control if output Observation minus Analysis files
(including also O minus B) which are ASCII format and separated by sensors and
processors.
USE_ERROR_FACTOR_RAD
Logical, controls use of a radiance error tuning factor
file Òradiance_error.factorÓ,
which is created with empirical values or generated using variational
tunning method (Desroziers and Ivanov, 2001)
ONLY_SEA_RAD
Logical, controls whether only assimilating radiance over
water.
TIME_WINDOW_MIN
String, e.g., "2007-08-15_03:00:00.0000", start
time of assimilation time window
TIME_WINDOW_MAX
String, e.g., "2007-08-15_09:00:00.0000", end
time of assimilation time window
USE_ANTCORR (30)
Logical array with dimension RTMINIT_NSENSER, control if
performing Antenna Correction in CRTM.
AIRS_WARMEST_FOV
Logical, controls whether using the observation brightness
temperature for AIRS Window channel #914 as criterium for GSI thinning.
USE_CRTM_KMATRIX
Logical, controls whether using CRTM K matrix rather than calling CRTM TL and AD routines for gradient calculation.
USE_RTTOV_KMATRIX
Logical, controls whether using RTTOV K matrix rather than calling RTTOV TL and AD routines for gradient calculation.
RTTOV_EMIS_ATLAS_IR
integer, control the use of IR emissivity atlas.
Emissivity atlas data (should be downloaded separately from the RTTOV web site) need to be copied or linked under the sub-directory emis_data in the working directory if RTTOV_EMIS_ATLAS_IR is set to 1.
RTTOV_EMIS_ATLAS_MW
integer, control the use of MW emissivity atlas.
Emissivity atlas data (should be downloaded separately from the RTTOV web site) need to be copied or linked under the sub-directory emis_data in the working directory if RTTOV_EMIS_ATLAS_MW is set to 1 or 2.
h. Diagnostics and Monitoring
(1) Monitoring capability within WRFDA.
Run WRFDA with the rad_monitoring namelist parameter in
record wrfvar14 in namelist.input.
0 means assimilating mode, innovations (O minus B) are
calculated and data are used in minimization.
1 means monitoring mode: innovations are calculated for
diagnostics and monitoring. Data are not used in minimization.
Number of rad_monitoring should correspond to number
of rtminit_nsensor. If
rad_monitoring is not set, then default value of 0 will be used for all
sensors.
(2) Outputing radiance diagnostics from WRFDA
Run WRFDA with the following namelist variables in record
wrfvar14 in namelist.input.
write_iv_rad_ascii=.true.
to write out (observation-background) and other diagnostics
information in plain-text files with prefix inv followed by instrument name and
processor id. For example, 01_inv_noaa-17-amsub.0000 (01 is outerloop index,
0000 is processor index)
write_oa_rad_ascii=.true.
to write out (observation-background),
(observation-analysis) and other diagnostics information in plain-text files
with prefix oma followed by instrument name and processor id. For example,
01_oma_noaa-18-mhs.0001
Each processor writes out information of one instrument in
one file in the WRFDA working directory.
(3) Radiance diagnostics data processing
A Fortran90 program is used to collect the 01_inv* or
01_oma* files and write out in netCDF format (one instrument in one file with
prefix diags followed by instrument name, analysis date, and suffix .nc)) for
easier data viewing, handling and plotting with netCDF utilities and NCL
scripts.
(4) Radiance diagnostics plotting
NCL scripts (WRFDA/var/graphics/ncl/plot_rad_diags.ncl and
WRFDA/var/graphics/ncl/advance_cymdh.ncl) are used for plotting. The NCL script
can be run from a shell script, or run stand-alone with interactive ncl command
(need to edit the NCL script and set the plot options. Also the path of advance_cymdh.ncl,
a date advancing script loaded in the main NCL plotting script, may need to be
modified).
Step (3) and (4) can be done by running a single ksh script
(WRFDA/var/scripts/da_rad_diags.ksh) with proper settings. In addition to the
settings of directories and what instruments to plot, there are some useful
plotting options, explained below.
|
export OUT_TYPE=ncgm |
ncgm or pdf pdf will be much slower than ncgm and
generate huge output if plots are not split. But pdf has higher resolution
than ncgm. |
|
export
PLOT_STATS_ONLY=false |
true or false true: only statistics of OMB/OMA vs
channels and OMB/OMA vs dates will be plotted. false: data coverage, scatter plots
(before and after bias correction), histograms (before and after bias
correction), and statistics will be plotted. |
|
export
PLOT_OPT=sea_only |
all, sea_only,
land_only |
|
export PLOT_QCED=false |
true or false true: plot only quality-controlled data false: plot all data |
|
export
PLOT_HISTO=false |
true or false: switch
for histogram plots |
|
export PLOT_SCATT=true |
true or false: switch
for scatter plots |
|
export
PLOT_EMISS=false |
true or false: switch
for emissivity plots |
|
export
PLOT_SPLIT=false |
true or false true: one frame in each file false: all frames in one file |
|
export
PLOT_CLOUDY=false |
true or false true: plot cloudy data. Cloudy data to be
plotted are defined by PLOT_CLOUDY_OPT (si or clwp), CLWP_VALUE, SI_VALUE
settings. |
|
export
PLOT_CLOUDY_OPT=si |
si or clwp clwp: cloud liquid water path from model si: scatter index from obs, for amsua,
amsub and mhs only |
|
export CLWP_VALUE=0.2 |
only plot points with clwp >= clwp_value (when clwp_value
> 0) clwp > clwp_value (when clwp_value = 0) |
|
export SI_VALUE=3.0 |
|
(5) evolution of VarBC parameters
NCL scripts (WRFDA/var/graphics/ncl/plot_rad_varbc_param.ncl
and WRFDA/var/graphics/ncl/advance_cymdh.ncl)
are used for plotting evolutions of VarBC parameters.
Updating WRF Boundary Conditions
There are three input files: WRFDA analysis, wrfinput, and wrfbdy files from WPS/real.exe, and a namelist file: param.in for running da_update_bc.exe for domain-1. Before running NWP forecast using the WRF-model with WRFDA analysis, update the values and tendencies for each predicted variable in the first time period in the lateral boundary condition file. For domain-1 (wrfbdy_d01) must be updated to be consistent with the new WRFDA initial condition (analysis). This is absolutely essential. Moreover, in the cycling run mode (warm-start), the low boundary in the WRFDA analysis file also needs to be updated based on the information from the wrfinput file generated by WPS/real.exe at analysis time.
For the nested domains, domain-2, domain-3É, the lateral boundaries are provided by their parent domains, so no lateral boundary update is needed for these domains; but the low boundaries in each of the nested domainsÕ WRFDA analysis files still need to be updated. In these cases, you must set the namelist variable, domain_id > 1 (default is 1 for domain-1), and no wrfbdy_d01file need to be provided to the namelist variable: wrf_bdy_file.
This procedure is performed by the WRFDA utility called da_updated_bc.exe.
Note: Make sure that you have da_update_bc.exe in WRFDA/var/build directory. This executable should be created when you compiled WRFDA code,
To run da_update_bc.exe, follow the steps below:
> cd
WRFDA/var/test/update_bc
> cp
–p $DAT_DIR/rc/2008020512/wrfbdy_d01 ./wrfbdy_d01 (IMPORTANT: make a copy
of wrfbdy_d01 as the wrf_bdy_file will be overwritten by da_update_bc.exe)
> vi
parame.in
&control_param
wrfvar_output_file = './wrfvar_output'
wrf_bdy_file = './wrfbdy_d01'
wrf_input =
'$DAT_DIR/rc/2008020512/wrfinput_d01'
cycling = .false. (set to .true. if
WRFDA first guess comes from a previous WRF forecast.)
debug = .true.
low_bdy_only = .false.
update_lsm = .false.
/
> ln
–sf WRFDA/var/da/da_update_bc.exe ./da_update_bc.exe
>
./da_updatebc.exe
At this stage, you should have the files wrfvar_output and wrfbdy_d01 in your WRFDA working directory. They are the WRFDA updated initial condition and boundary condition for any subsequent WRF model runs. To use, link a copy of wrfvar_output and wrfbdy_d01 to wrfinput_d01 and wrfbdy_d01, respectively, in your WRF working directory.
As of V3.2, some changes were made
to da_update_bc to address some issues that are related to sea-ice and snow
change during cycling runs and radiance data assimilation. The new settings in
parame.in are introduced as follows. (However, for backward compatibility, the
pre-V3.2 parame.in mentioned above still works with V3.2+ da_update_bc)
&control_param
da_file
= '../tutorial/wrfvar_output'
wrf_bdy_file = './wrfbdy_d01'
wrf_input =
'$DAT_DIR/rc/2008020512/wrfinput_d01'
domain_id = 1
debug
= .true.
update_lateral_bdy = .true.
update_low_bdy = .true.
update_lsm = .false.
iswater
= 16
/
update_lateral_bdy is required only for domain 1.
update_low_bdy is needed for all
domains if running in cycling mode.
iswater (water point index) is 16
for USGS LANDUSE and 17 for MODIS LANDUSE.
Running da_update_bc.exe is
recommended,
update_lateral_bdy = .false.
update_low_bdy = .true.
before running WRFDA, if in cycling
mode (especially if you are doing radiance data assimilation and there is SEA
ICE and SNOW in your domain) to get low-bdy updated first guess (da_file will be overwritten by da_update_bc).
Next run da_update_bc.exe with
update_lateral_bdy = .true.
update_low_bdy = .false.
after WRFDA to get updated lateral
boubdary conditions (wrf_bdy_file will be overwritten by da_update_bc).
Running gen_be
Starting with WRFDA version 3.1, users have three choices to define the background error covariance (BE). We call them CV3, cv5, and CV6 respectively. With CV3 and CV5, the background errors are applied to the same set of the control variables, stream function, unbalanced potential velocity, unbalanced temperature, unbalanced surface pressure, and pseudo relative humidity. However, for CV6 the moisture control variable is the unbalanced part of pseudo relative humidity. With CV3, the control variables are in physical space while with CV5 and CV6 the control variables are in eigenvector space. So, the major differences between these two kinds of BE is the vertical covariance. CV3 uses the vertical recursive filter to model the vertical covariance but CV5 and CV6 use empirical orthogonal function (EOF) to represent the vertical covariance. The recursive filters to model the horizontal covariance are also different with these BEs. We have not conducted the systematic comparison of the analyses based on these BEs. However, CV3 (a BE file provided with our WRFDA system) is a global BE and can be used for any regional domains while CV5 and CV6 BEÕs are domain-dependent, which should be generated based in the forecasts data from the same domain. At this time, it is hard to tell which BE is better; the impact on analysis may vary case by case.
CV3 is the NCEP background error covariance, it is estimated in grid space by what has become known as the NMC method (Parrish and Derber 1992) . The statistics are estimated with the differences of 24 and 48-hour GFS forecasts with T170 resolution valid at the same time for 357 cases distributed over a period of one year. Both the amplitudes and the scales of the background error have to be tuned to represent the forecast error in the guess fields. The statistics that project multivariate relations among variables are also derived from the NMC method.
The variance of each variable and the variance of its second derivative are used to estimate its horizontal scales. For example, the horizontal scales of the stream function can be estimated from the variance of the vorticity and stream function.
The vertical scales are estimated with the vertical correlation of each variable. A table is built to cover the range of vertical scales for the variables. The table is then used to find the scales in vertical grid units. The filter profile and the vertical correlation are fitted locally. The scale of the best fit from the table is assigned as the scale of the variable at that vertical level for each latitude. Note that the vertical scales are locally defined so that the negative correlation further away in the vertical direction is not included.
Theoretically, CV3 BE is a generic background error statistics file which, can be used for any case. It is quite straightforward to use CV3 in your own case. To use the CV3 BE file in your case, set cv_options=3 in $wrfvar7 and the be.dat is located in WRFDA/var/run/be.dat.cv3.
To use CV5 or CV6 background error covariance, it is necessary to generate your domain-specific background error statistics with the gen_be utility. The background error statistics file supplied with the tutorial test case can NOT be used for your applications.
The Fortran main programs for gen_be can be found in WRFDA/var/gen_be. The executables of gen_be should be created after you have compiled the WRFDA code (as described earlier). The scripts to run these codes are in WRFDA/var/scripts/gen_be.
The input data for gen_be are WRF forecasts, which are used to generate model perturbations, used as a proxy for estimates of forecast error. For the NMC-method, the model perturbations are differences between forecasts (e.g. T+24 minus T+12 is typical for regional applications, T+48 minus T+24 for global) valid at the same time. Climatological estimates of background error may then be obtained by averaging such forecast differences over a period of time (e.g. one month). Given input from an ensemble prediction system (EPS), the inputs are the ensemble forecasts, and the model perturbations created are the transformed ensemble perturbations. The gen_be code has been designed to work with either forecast difference, or ensemble-based perturbations. The former is illustrated in this tutorial example.
It is important to include forecast differences from at least 00Z and 12Z through the period, to remove the diurnal cycle (i.e. do not run gen_be using just 00Z or 12Z model perturbations alone).
The inputs to gen_be are netCDF WRF forecast output ("wrfout") files at specified forecast ranges. To avoid unnecessary large single data files, it is assumed that all forecast ranges are output to separate files. For example, if we wish to calculate BE statistics using the NMC-method with (T+24)-(T+12) forecast differences (default for regional) then by setting the WRF namelist.input options history_interval=720, and frames_per_outfile=1 we get the necessary output datasets. Then the forecast output files should be arranged as follows: directory name is the forecast initial time, time info in the file name is the forecast valid time. 2008020512/wrfout_d01_2008-02-06_00:00:00 means a 12-hour forecast valid at 2008020600 initialized at 2008020512.
Example dataset for a test case (90 x 60 x 41 gridpoints) can be downloaded from http://www2.mmm.ucar.edu/wrf/users/wrfda/download/testdata.html, untar the gen_be_forecasts_20080205.tar.gz, you will have:
>ls $FC_DIR
-rw-r--r-- 1 users
11556492 2008020512/wrfout_d01_2008-02-06_00:00:00
-rw-r--r-- 1 users
11556492 2008020512/wrfout_d01_2008-02-06_12:00:00
-rw-r--r-- 1 users
11556492 2008020600/wrfout_d01_2008-02-06_12:00:00
-rw-r--r-- 1 users
11556492 2008020600/wrfout_d01_2008-02-07_00:00:00
-rw-r--r-- 1 users
11556492 2008020612/wrfout_d01_2008-02-07_00:00:00
-rw-r--r-- 1 users
11556492 2008020612/wrfout_d01_2008-02-07_12:00:00
In the above example, only 1 day (12Z 05 Feb to 12Z 06 Feb. 2002) of forecasts, every 12 hours are supplied to gen_be_wrapper to estimate forecast error covariance. It is only for demonstration. The minimum number of forecasts required depends on the application, number of grid points, etc. Month-long (or longer) datasets are typical for the NMC-method. Generally, at least a 1-month dataset should be used.
Under WRFDA/var/scripts/gen_be, gen_be_wrapper.ksh is used to generate the BE data, following variables need to be set to fit your case:
export
WRFVAR_DIR=/users/noname/work/code/trunk/phoenix_g95_opt/WRFDA
export
START_DATE=2008020612 # the first
perturbation valid date
export
END_DATE=2008020700 #
the last perturbation valid date
export
NUM_LEVELS=40 # e_vert - 1
export
BIN_TYPE=5
export
FC_DIR=/users/noname/work/exps/friendlies/expt/fc # where wrf forecasts are
export
RUN_DIR=/users/noname/work/exps/friendlies/gen_be${BIN_TYPE}
Note: The START_DATE and END_DATE are perturbation valid dates. As show in the forecast list above, when you have 24-hour and 12-hour forecasts initialized at 2008020512 through 2008020612, the first and final forecast difference valid dates are 2008020612 and 2008020700 respectively.
Note: The forecast dataset should be located in $FC_DIR. Then type:
>
gen_be_wrapper.ksh
Once gen_be_wrapper.ksh run is completed, the be.dat can be found under $RUN_DIR directory.
To get a clear idea about what are included in be.dat, the script gen_be_plot_wrapper.ksh may be used to plot various data in be.dat, for example: