---
title: "Automate Creation of CWL Instructions"
author: Le Zhang, Thomas Girke"
date: last-modified
sidebar: tutorials
bibliography: bibtex.bib
---

```{r}
#| include: false
knitr::opts_chunk$set(echo = TRUE)
```
## Introduction
A central concept for designing workflows within the `systemPipeR` environment
is the usage of workflow management containers. For describing analysis
workflows in a generic and flexible manner the [Common Workflow
Language](https://www.commonwl.org/) (CWL) has been adopted throughout the
environment including the workflow management containers [@Amstutz2016-ka].
Using the CWL community standard in `systemPipeR` has many advantages. For
instance, the integration of CWL allows running `systemPipeR` workflows from a
single specification instance either entirely from within R, from various
command line wrappers (e.g., cwl-runner) or from other languages (e.g., Bash or
Python). An important feature of `systemPipeR's` CWL interface is that it
provides two options to run command line tools and workflows based on CWL.
First, one can run CWL in its native way via an R-based wrapper utility for
`cwl-runner` or `cwl-tools` (CWL-based approach). Second, one can run workflows
using CWL's command line and workflow instructions from within R (R-based
approach). In the latter case the same CWL workflow definition files (e.g.
*.cwl* and *.yml*) are used but rendered and executed entirely with R functions
defined by `systemPipeR`, and thus use CWL mainly as a command line and
workflow definition format rather than execution software to run workflows.
Moreover, `systemPipeR` provides several convenience functions that are useful
for designing and debugging workflows, such as a command-line rendering
function to retrieve the exact command-line strings for each data set and
processing step prior to running a command-line.
This tutorial briefly introduces the basics how CWL defines command-line
syntax. Next, it describes how to use CWL within `systemPipeR` for designing,
modifying and running workflows.
## Load package
Recent versions of R (>=4.0.0), Bioconductor (>=3.14) and `systemPipeR` (>=2.0.8)
need to be used to gain access to the functions described in this tutorial.
```{r load_library}
#| include: false
library(systemPipeR)
```
## CWL command line specifications
CWL command line specifications are written in [YAML](http://yaml.org/) format.
In CWL, files with the extension `.cwl` define the parameters of a chosen
command line step or workflow, while files with the extension `.yml` define
the input variables of command line steps.
The following introduces first the basic structure of `.cwl` files.
```{r}
dir_path <- system.file("extdata/cwl/example/", package="systemPipeR")
cwl <- yaml::read_yaml(file.path(dir_path, "example.cwl"))
```
- The `cwlVersion` component specifies the version of CWL that is used here.
- The `class` component declares the usage of a command-line tool.
Note, CWL has another `class` called `Workflow`. The latter defines one
or more command-line tools, while `CommandLineTool` is limited to one.
```{r}
cwl[1:2]
```
- The `baseCommand` component contains the base name of the software to be executed.
```{r}
cwl[3]
```
- The `inputs` component provides the input information required for the command-line software. Important sub-components of this section are:
- `id`: each input has an id assigning a name
- `type`: input type value (e.g. string, int, long, float, double,
File, Directory or Any);
- `inputBinding`: optional component indicating if the input
parameter should appear on the command line. If missing then the
parameter will not appear in the command-line.
```{r}
cwl[4]
```
- The `outputs` component should provide a list of the outputs expected after running a command-line tools.
Important sub-components of this section are:
- `id`: each output has an id assigning a name
- `type`: output type value (e.g. string, int, long, float, double,
File, Directory, Any or `stdout`)
- `outputBinding`: defines how to set the outputs values. The `glob` component will define the name of the output value.
```{r}
cwl[5]
```
- `stdout`: specifies a `filename` for capturing standard output.
Note here we are using a syntax that takes advantage of the inputs section,
using `results_path` parameter and also the `SampleName` to construct the `filename` of the output.
```{r}
cwl[6]
```
Next, the structure and content of the `.yml` files will be introduced. The `.yml` file
provides the parameter values for the `.cwl` components described above.
The following example defines three parameters.
```{r}
yaml::read_yaml(file.path(dir_path, "example_single.yml"))
```
Importantly, if an input component is defined in the corresponding *.cwl* file, then the
required value needs to be provided by the corresponding component of the *.yml* file.
### How to connect CWL description files within `systemPipeR`
A `SYSargsList` container stores several `SYSargs2` instances in a list-like object containing
all instructions required for processing a set of input files with a single or many command-line
steps within a workflow (i.e. several tools of one software or several independent software tools).
A single `SYSargs2` object is created and fully populated with the constructor functions
`loadWF` and `renderWF`.
The following imports a `.cwl` file (here `example.cwl`) for running a simple `echo Hello World`
example where a string `Hello World` will be printed to stdout and redirected to a file named
`M1.txt` located under a subdirectory named `results`.
```{r fromFile}
HW <- loadWF(wf_file="example.cwl", input_file="example_single.yml",
dir_path = dir_path)
HW <- renderWF(HW)
HW
cmdlist(HW)
```
The above example is limited to running only one command-line call, corresponding to one
input file, e.g. representing a single experimental sample. To scale to many command-line
calls, e.g. when processing many input samples, a simple solution offered by `systemPipeR`
is to use `variables`, one for each parameter with many inputs.
The following gives a simple example for defining and processing many inputs.
```{r}
yml <- yaml::read_yaml(file.path(dir_path, "example.yml"))
yml
```
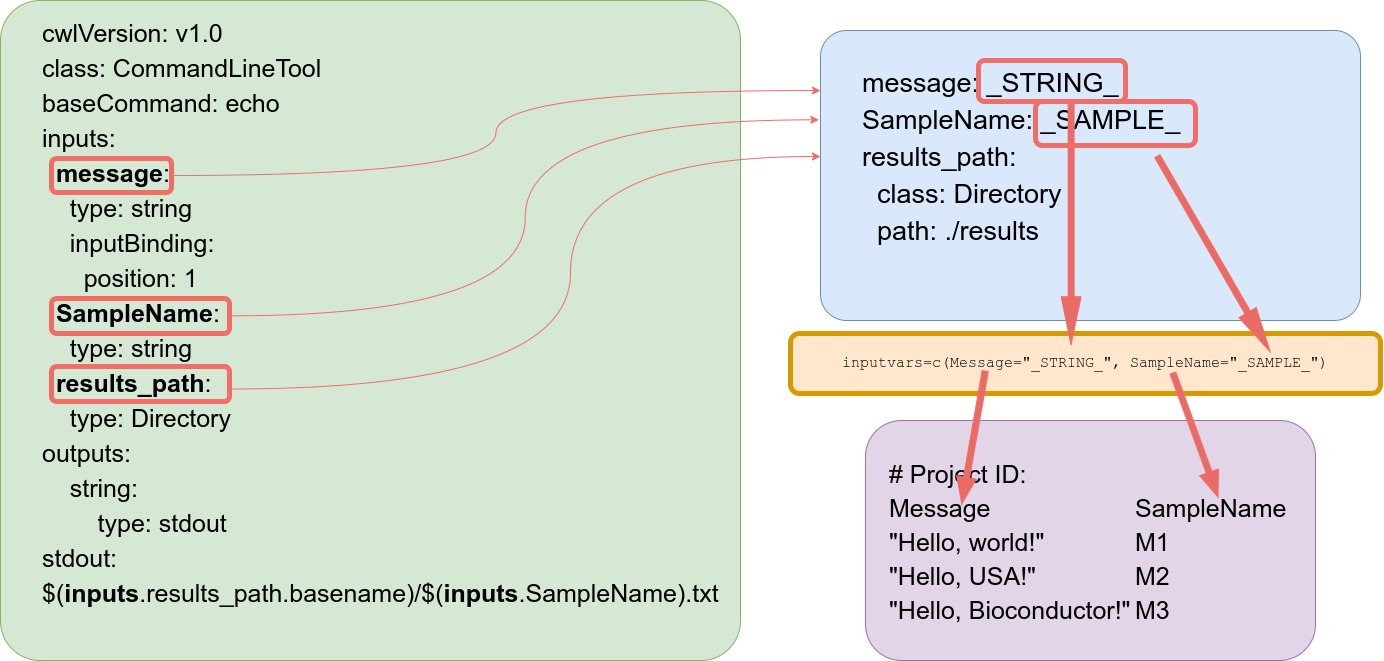
Under the `message` and `SampleName` parameters, variables are used for that will be populated
by values provided by a third file called `targets.`
The following shows the structure of a simple `targets` file.
```{r}
targetspath <- system.file("extdata/cwl/example/targets_example.txt", package="systemPipeR")
read.delim(targetspath, comment.char = "#")
```
With help of a `targets` file, one can define all input files, sample ids and
experimental variables relevant for an analysis workflow. In the above example,
strings defined under the `Message` column will be passed on to the `echo`
command-line tool. In addition, each command-line will be assigned a label or
id specified under `SampleName` column. Any number of additional columns can be
added as needed.
Users should note here, the usage of `targets` files is optional when using
`systemPipeR's` CWL interface. Since targets files are very efficient for
organizing experimental variables, their usage is highly encouraged and well
supported in `systemPipeR`.
#### Connect parameter and targets files
The constructor functions construct an `SYSargs2` instance from three input files:
- `.cwl` file path assigned to `wf_file` argument
- `.yml` file path assigned to `input_file` argument
- `target` file assigned to `targets` argument
As mentioned above, the latter `targets` file is optional. The connection
between input variables (here defined by `input_file` argument) and the
`targets` file are defined under the `inputvars` argument. A named vector is
required, where each element name needs to match the column names in the
`targets` file, and the value must match the names of the *.yml* variables.
This is used to replace the CWL variable and construct the command-lines, usually
one for each input sample.
For consistency the pattern `_XXXX_` is used for variable naming in the `.yml` file, where the
name matches the corresponding column name in the targets file. This pattern is recommended
for easy identification but not enforced.
The following imports a `.cwl` file (same example as above) for running
the `echo` example. However, now several command-line calls are constructed with the
information provided under the `Message` column of the targets file that is passed on to
matching component in the `.yml` file.
```{r fromFile_example}
HW_mul <- loadWorkflow(targets = targetspath, wf_file="example.cwl",
input_file="example.yml", dir_path = dir_path)
HW_mul <- renderWF(HW_mul, inputvars = c(Message = "_STRING_", SampleName = "_SAMPLE_"))
HW_mul
cmdlist(HW_mul)
```
{fig-align="center" width=50%}
## Auto-creation of CWL param files from command-line
Users can define the command-line in a pseudo-bash script format. The following used the
the command-line for `HISAT2` as example.
```{r cmd}
command <- "
hisat2 \
-S \
-x \
-k \
-min-intronlen \
-max-intronlen \
-threads \
-U
"
```
### Define prefix and defaults
- First line is the base command. Each line is an argument with its default value.
- All following lines specify arguments. Lines starting with a `-` or `--` followed
by a non-space delimited letter/word will be interpreted as a prefix, e.g.
`-S` or `--min`. Lines without this prefix will be rendered as non-prefix arguments.
- All default settings are placed inside `<...>`. Omit for arguments without values
such as `--verbose`.
- First argument is the type of the input. `F` for "File", "int" and "string" are unchanged.
- Optional: keyword `out` followed the type. Separation by `,` (comma) indicates
whether this argument is also a CWL output.
- Use `:` to separate keywords and default values. Any non-space separated value after the `:`
will be treated as the default value.
### `createParamFiles` Function
The `createParamFiles` function accepts as input a command-line provided in above `string` syntax.
The function returns a `cwl` with the following components:
- `BaseCommand`: Specifies the program to execute
- `Inputs`: Defines the input parameters of the process
- `Outputs`: Defines the parameters representing the output of the process
The fourth component is the original command-line provided as input.
In interactive mode, the function will verify if everything is correct and
ask the user to proceed. The user can answer "no" and provide more information
at the string input level. Another question is whether to save the generated CWL
results to the corresponding `.cwl` and `.yml` files. When running the function
in non-interactive mode, the results will be returned without asking for confirmation
by the user.
```{r}
cmd <- createParamFiles(command, writeParamFiles = FALSE)
```
If the user chooses not to save the `param` files in the `createParamFiles` call directly,
then the `writeParamFiles` function allows to do this in a separate step.
```{r saving}
writeParamFiles(cmd, overwrite = TRUE)
```
### Accessor functions
#### Print components
Note, the results of `createParamFiles` are stored in a `SYSargs2` container. The individual
components can be accessed as follows.
```{r}
printParam(cmd, position = "baseCommand") ## Print a baseCommand section
printParam(cmd, position = "outputs")
printParam(cmd, position = "inputs", index = 1:2) ## Print by index
printParam(cmd, position = "inputs", index = -1:-2) ## Negative indexing printing to exclude certain indices in a position
cmdlist(cmd)
```
#### Subsetting the command-line
```{r}
cmd2 <- subsetParam(cmd, position = "inputs", index = 1:2, trim = TRUE)
cmdlist(cmd2)
cmd2 <- subsetParam(cmd, position = "inputs", index = c("S", "x"), trim = TRUE)
cmdlist(cmd2)
```
#### Replacing existing argument
```{r}
cmd3 <- replaceParam(cmd, "base", index = 1, replace = list(baseCommand = "bwa"))
cmdlist(cmd3)
```
```{r}
new_inputs <- new_inputs <- list(
"new_input1" = list(type = "File", preF="-b", yml ="myfile"),
"new_input2" = "-L "
)
cmd4 <- replaceParam(cmd, "inputs", index = 1:2, replace = new_inputs)
cmdlist(cmd4)
```
#### Adding new arguments
```{r}
newIn <- new_inputs <- list(
"new_input1" = list(type = "File", preF="-b1", yml ="myfile1"),
"new_input2" = list(type = "File", preF="-b2", yml ="myfile2"),
"new_input3" = "-b3 "
)
cmd5 <- appendParam(cmd, "inputs", index = 1:2, append = new_inputs)
cmdlist(cmd5)
cmd6 <- appendParam(cmd, "inputs", index = 1:2, after=0, append = new_inputs)
cmdlist(cmd6)
```
#### Editing `output` param
```{r}
new_outs <- list(
"sam_out" = ""
)
cmd7 <- replaceParam(cmd, "outputs", index = 1, replace = new_outs)
output(cmd7)
```
## Version information
```{r sessionInfo}
sessionInfo()
```
## Funding
This project is funded by NSF award [ABI-1661152](https://www.nsf.gov/awardsearch/showAward?AWD_ID=1661152).
## References
